Abstract
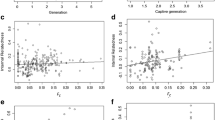
Inbreeding depression is frequently a concern of managers interested in restoring endangered species. Decisions to reduce the potential for inbreeding depression by balancing genotypic contributions to reintroduced populations may exact a cost on long-term demographic performance of the population if those decisions result in reduced numbers of animals released and/or restriction of particularly successful genotypes (i.e., heritable traits of particular family lines). As part of an effort to restore a migratory flock of Whooping Cranes (Grus americana) to eastern North America using the offspring of captive breeders, we obtained a unique dataset which includes post-release mark–recapture data, as well as the pedigree of each released individual. We developed a Bayesian formulation of a multi-state model to analyze radio-telemetry, band-resight, and dead recovery data on reintroduced individuals, in order to track survival and breeding state transitions. We used studbook-based individual covariates to examine the comparative evidence for and degree of effects of inbreeding, genotype, and genotype quality on post-release survival of reintroduced individuals. We demonstrate implementation of the Bayesian multi-state model, which allows for the integration of imperfect detection, multiple data types, random effects, and individual- and time-dependent covariates. Our results provide only weak evidence for an effect of the quality of an individual’s genotype in captivity on post-release survival as well as for an effect of inbreeding on post-release survival. We plan to integrate our results into a decision-analytic modeling framework that can explicitly examine tradeoffs between the effects of inbreeding and the effects of genotype and demographic stochasticity on population establishment.






Similar content being viewed by others
References
Barker RJ, White GC, McDougal M (2005) Movement of paradise shelduck between molt sites: a joint multistate-dead recovery mark recapture model. J Wildl Manag 69:1194–1201
Blouin MS, Parsons M, LaCaille V, Lotz S (1996) Use of microsatellite loci to classify individuals by relatedness. Mol Ecol 5:393–401
Canadian Wildlife Service, and U.S. Fish and Wildlife Service (2005) International recovery plan for the whooping crane. Ottawa, Canada: Recovery of Nationally Endangered Wildlife (RENEW), and U.S. Fish and Wildlife Service, New Mexico
Clemen RT (1996) Making hard decisions: an introduction to decision analysis. Duxbury, Belmont
Crnokrak P, Roff DA (1999) Inbreeding depression in the wild. Heredity 83:260–270
Fischer J, Lindenmayer DB (2000) An assessment of the published results of animal relocations. Biol Conserv 96:1–11
Frankham R (1995) Conservation genetics. Annu Rev Genet 29:305–327
Frankham R (2007) Genetic adaptation to captivity in species conservation programs. Mol Ecol 2007:1–5
Frankham R, Ballou JD, Briscoe DA (2002) Introductiono to conservation genetics. Cambridge University Press, Cambridge
Gelman A, Carlin JB, Stern HS, Rubin DB (2004) Bayesian data analysis, 2nd edn. Chapman & Hall/CRC, Boca Raton
Gilks W, Thomas A, Spiegelhalter D (1996) A language and program for complex Bayesian modelling. Statistician 43:169–178
Glenn TC, Stephan W, Braun MJ (1999) Effects if a population bottleneck on whooping crane mitochondrial DNA variation. Conserv Biol 13:1097–1107
Green MJ, Medley GF, Browne WJ (2009) Use of posterior predictive assessments to evaluate model fit in multilevel logistic regression. Vet Res 40:30
Jones KL, Glenn TC, Lacy RC, Pierce JR, Unruh N, Mirande CM, Chavez-Ramirez F (2002) Refining the whooping crane studbook by incorporating microsatellite DNA and leg-banding analyses. Conserv Biol 16:789–799
Kendall WL, Conn PB, Hines JE (2006) Combining multi-state capture-recapture data with tag recoveries to estimate demographic parameters. Ecology 87:169–177
Lande R (1988) Genetics and demography in biological conservation. Science 241:1455–1460
Lebreton J-D, Almeras T, Pradel R (1999) Competing events, mixtures of information and multistratum recapture models. Bird Study Suppl 46:S39–S46
Link WA, Barker RJ (2006) Model weights and the foundations of multimodel inference. Ecology 87:2626–2635
Link WA, Royle JA, Hatfield JS (2003) Demographic analysis from summaries of an age-structured population. Biometrics 59:778–785
Millar R (2009) Comparison of hierarchical Bayesian models for over-dispersed count data using DIC and Bayes factors. Biometrics 65:962–969
Mirande CM (1995) The genealogy of the whooping crane (Grus americana). International Crane Foundation, Baraboo
Moore CT, Converse SJ, Folk M, Boughton R, Brooks B, French JB, O’Meara TE, Putnam M, Rodgers J, Spalding M (2008) Releases of whooping cranes to the Florida nonmigratory flock: a structured decision-making approach. Report to the International Whooping Crane Recovery Team. Florida Fish and Wildlife Research Institute In-House Report IHR2008-009. Available at: http://research.myfwc.com/publications/publication_info.asp?id=58528
Moore CT, Converse SJ, Folk MJ, Runge MC, Nesbitt SA (2011) Evaluating release alternatives for a long-lived bird species under uncertainty about long-term demographic rates. J Ornithol (in press)
Queller DC, Goodnight KF (1989) Estimating relatedness using genetic markers. Evolution 43:258–275
R Development Core Team (2004) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Sturtz S, Ligges U, Gelman A (2005) R2WinBUGS: a package for running WinBUGS from R. J Stat Softw 12:1–16
Urbanek RP, Fondow LEA, Satyshur CD, Lacy AE, Zimorski SE, Wellington M (2005) First cohort of migratory whooping cranes reintroduced to eastern North America: the first year after release. Proc North Am Crane Workshop 9:213–223
Urbanek RP, Fondow LEA, Zimorski SE, Wellington MA, Nipper MA (2009) Winter release and management of reintroduced migratory whooping cranes Grus americana. Bird Conser Int 19:1–12
Wolf CM, Griffith B, Reed C, Temple SA (1996) Avian and mammalian translocations: update and reanalysis of 1987 survey data. Conserv Biol 10:1142–1154
Acknowledgments
We thank Ken Jones who provided the inbreeding coefficients. We thank the field and breeding center staff who were involved in collecting and providing data, especially J. Chandler, A. Fasoli, B. Hartup, M. Wellington, and S. Zimorski. We extend our appreciation to the Whooping Crane Eastern Partnership for facilitating our collaboration and to the National Fish and Wildlife Foundation for funding this work. D. Diefenbach, K. Jones, E. Zipkin, and one anonymous reviewer provided helpful reviews of the draft manuscript. Use of trade, product, or firm names does not imply endorsement by the U.S. Government.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by W. L. Kendall.
Rights and permissions
About this article
Cite this article
Converse, S.J., Royle, J.A. & Urbanek, R.P. Bayesian analysis of multi-state data with individual covariates for estimating genetic effects on demography. J Ornithol 152 (Suppl 2), 561–572 (2012). https://doi.org/10.1007/s10336-011-0695-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10336-011-0695-0