Abstract
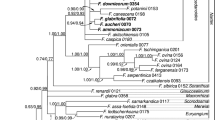
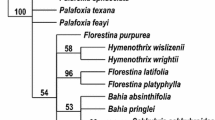
Pyroleae (Ericaceae) consist of four genera, all of which are distributed widely in temperate coniferous or sometimes deciduous forests of the Northern Hemisphere. To investigate the phylogenetic relationships among these genera and to explore the evolution of the characteristics of the subfamily, we conducted maximum parsimony and Bayesian analyses with nrDNA ITS and three cpDNA intergenic spacers (atpB-rbcL, trnS-trnG and trnL-trnF). The results from cpDNA and combined cpDNA + ITS data sets strongly support the monophyly of Pyroleae as well as a sister relationship between Pyrola and Moneses–Chimaphila, with Orthilia as the basal lineage. The sister-group relationship between Moneses and Chimaphila is supported by a set of synapomorphies, e.g., single flower, colpate pollen, five bundles in the style, straight fruiting pedicel orientation, complete capsule dehiscence, and the basic chromosome number, x = 13. The Moneses–Chimaphila–Pyrola clade is supported by at least one homologous character of pollen in tetrads. Conflicts associated with the phylogenetic position of Orthilia may imply a hybrid origin for it, and therefore further study is needed.






Similar content being viewed by others
References
Anderberg AA (1993) Cladistic interrelationships and major clades of the Ericales. Plant Syst Evol 184:207–231
Anderberg AA (1994) Cladistic analysis of Enkianthus with notes on the early diversification of the Ericaceae. Nord J Bot 14:385–401
Andres H (1914) Piroleen-Studien. Beiträge zur Kenntnis der Morphologie, Phytogeographie und allgemeinen Systematik der Pirolaceae. Verh Bot Vereins Prov Brandenburg 56:1–76
Björkman E (1960) Monotropa hypopithys L.—an epiparasite on tree roots. Physiol Plant 13:399–401
Böcher TW (1961) Studies in Pyrolaceae-two interesting wintergreen from West Greenland. Bot Tidsskr 57:28–37
Bull JJ, Huelsenbeck JP, Cunningham CW, Swofford DL, Waddell PJ (1993) Partitioning and combining data in phylogenetic analysis. Syst Biol 42:384
Chippindale PT, Wiens JJ (1994) Weighting, partitioning, and combining characters in phylogenetic analysis. Syst Biol 43:278
Copeland HF (1941) Further studies on Monotropoideae. Madroño 6:97–119
Copeland HF (1947) Observations on the structure and classification of the Pyroleae. Madroño 9:65–102
Cronquist A (1981) An integrated system of classification of flowering plants. Columbia University Press, New York
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Drude O (1889) Pirolaceae. In: Engler A, Prantl K (eds) Die natürlichen Pflanzenfamilien, vol 4. Engelmann, Leipzig, pp 3–11
Erdtman G (1952) Pollen morphology and plant taxonomy. Angiosperms. Almqvist & Wiksell, Stockholm
Farris JS, Källersjö M, Kluge AG, Bult C (1995) Constructing a significance test for incongruence. Syst Biol 44:570
Freudenstein JV (1990) A phylogenetic study of the Pyroloideae (Ericaceae). Am J Bot 77:132
Freudenstein JV (1999) Relationships and character transformation in Pyroloideae (Ericaceae) based on ITS sequences, morphology, and development. Syst Bot 24:398–408
Haber E, Cruise JE (1974) Generic limits in the Pyroloideae (Ericaceae). Can J Bot 52:877–883
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hamilton MB (1999) Four primer pairs for the amplification of chloroplast intergenic regions with intraspecific variation. Mol Ecol 8:521–523
Henderson MW (1919) A comparative study of the structure and saprophytism of the Pyrolaceae and Monotropaceae with reference to their derivation from the Ericaceae. Contr Bot Lab Univ Pennsylvania 5:42–109
House HD (1921) Nomenclatorial notes on certain American plants. I. Am Midl Nat 7:126–135
Huelsenbeck J, Ronquist F (2001) MrBayes: Bayesian inference of phylogeny, version 3.1. 2. Bioinformatics 17:754–755
Jensen LCW (1961) Pollination studies with native Minnesota Pyrola and Moneses species. Proc Minn Acad Sci 29:210–218
Judd WS, Kron KA (1993) Circumscription of Ericaceae (Ericales) as determined by preliminary cladistic analyses based on morphological, anatomical, and embryological features. Brittonia 45:99–114
Kelchner SA (2000) The evolution of non-coding chloroplast DNA and its application in plant systematics. Ann Mo Bot Gard 87:482–498
Knaben G, Engelskjøn T (1968) Studies in Pyrolaceae, especially in the Pyrola rotundifolia complex. Arb Univ Bergen Mat Naturvitensk Ser 1967:1–71
Knudsen JT, Olesen JM (1993) Buzz-pollination and patterns in sexual traits in North European Pyrolaceae. Am J Bot 80:900–913
Křísa B (1971) Bertrag zur taxonomie und chorologie der gattung Pyrola L. Bot Jahrb Syst 90:476–508
Kron KA (1996) Phylogenetic relationships of Empetraceae, Epacridaceae, Ericaceae, Monotropaceae, and Pyrolaceae: evidence from nuclear ribosomal 18 s sequence data. Ann Bot 77:293–303
Kron KA, Judd WS, Stevens PF, Crayn DM, Anderberg AA, Gadek PA, Quinn CJ, Luteyn JL (2002) Phylogenetic classification of Ericaceae: molecular and morphological evidence. Bot Rev 68:335–423
Landhäusser SM, Stadt KJ, Lieffers VJ (1997) Photosynthetic strategies of summergreen and evergreen understory herbs of the boreal mixedwood forest. Oecologia 112:173–178
Maddison WP, Maddison DR (2005) Mesquite: a modular system for evolutionary analysis. Version 1.06. http://mesquiteproject.org. Accessed 22 Sept 2006
Manen JF, Natali A, Ehrendorfer F (1994) Phylogeny of Rubiaceae-Rubieae inferred from the sequence of a cpDNA intergene region. Plant Syst Evol 190:195–211
Mason-Gamer RJ, Kellogg EA (1996) Testing for phylogenetic conflict among molecular data sets in the tribe Triticeae (Gramineae). Syst Biol 45:524
Müller KF (2005) SeqState—primer design and sequence statistics for phylogenetic DNA data sets. Appl Bioinformatics 4:65–69
Nowicke JW (1966) Pollen morphology and classification of the Pyrolaceae and Monotropaceae. Ann Mo Bot Gard 53:213–219
Nylander JAA (2004) MrModeltest (version 2.2). http://www.abc.se/~nylander/
Nylander JAA, Ronquist F, Huelsenbeck JP, Nieves-Aldrey JL (2004) Bayesian phylogenetic analysis of combined data. Syst Biol 53:47–67
Posada D, Buckley TR (2004) Model selection and model averaging in phylogenetics: advantages of akaike information criterion and Bayesian approaches over likelihood ratio tests. Syst Biol 53:793–808
Qin HN, Stevens PF (2005) Ericaceae. In: Fang MY, Fang RZ, He MY, Hu LZ, Yang HB, Qin HN, Min TL, Chamberlain DF, Stevens PF, Wallace GD, Anderberg AA (eds) Flora of China, vol 14. Missouri Botanical Garden, St. Louis, pp 245–255
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381
Singh P, Carew GC (1990) Inland spruce cone rust of black spruce: Effect on cone and seed yield, and seed quality. Eur J Forest Pathol 20:397–404
Stevens PF (1971) A classification of the Ericaceae: subfamilies and tribes. Bot J Linn Soc 64:1–53
Swofford DL (2003) PAUP*: Phylogenetic analysis using parsimony (* and other methods), vers. 4.0b10. Sinauer, Sunderland
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Takahashi H (1988) Pollen morphology and systematics in two subfamilies of the Ericaceae: Pyroloideae and Monotropoideae. J Korean Plant Taxon 18:9–17
Takhtajan AL (1980) Outline of the classification of flowering plants (Magnoliophyta). Bot Rev 46:225–359
Tedersoo L, Pellet P, Kõljalg U, Selosse MA (2007) Parallel evolutionary paths to mycoheterotrophy in understorey Ericaceae and Orchidaceae: ecological evidence for mixotrophy in Pyroleae. Oecologia 151:206–217
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Thorne RF (1983) Proposed new realignments in the angiosperms. Nord J Bot 3:85–117
Thorne RF (1992) Classification and geography of the flowering plants. Bot Rev 58:225–327
Wallace GD (1975) Interrelationships of the subfamilies of the Ericaceae and derivation of the Monotropoideae. Bot Not 128:286–298
Warner BG, Chinnappa CC (1986) Taxonomic implications and evolutionary trends in pollen of Canadian Ericales. Can J Bot 64:3113–3126
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications, vol 18. Academic Press, San Diego, pp 315–322
Wood CE Jr (1961) The genera of Ericaceae in the southeastern United States. J Arnold Arbor 42:10–80
Acknowledgments
This study was part of a PhD project by Zhen-wen Liu and was supported by the National Natural Science Foundation of China (Grant 30900075). The authors are grateful to John V. Freudenstein, Hidie Takahashima and Shu-dong Zhang for allowing us to use DNA samples and leaf material. We thank Xun Gong for support during the laboratory work. We appreciate Sylvia Phillips, Julian Harber and David Boufford for polishing our English language. We are greatly indebted to two anonymous reviewers, whose comments were of great help in improving the quality of this paper.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, Zw., Wang, Zh., Zhou, J. et al. Phylogeny of Pyroleae (Ericaceae): implications for character evolution. J Plant Res 124, 325–337 (2011). https://doi.org/10.1007/s10265-010-0376-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-010-0376-8