Abstract
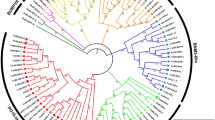
Increasing evidence suggests that a gene family encoding proteins containing BURP domains have diverse functions in plants, but systematic characterization of this gene family have not been reported. In this study, 17 BURP family genes (OsBURP01–17) were identified and analyzed in rice (Oryza sativa L.). These genes have diverse exon–intron structures and distinct organization of putative motifs. Based on the phylogenetic analysis of BURP protein sequences from rice and other plant species, the BURP family was classified into seven subfamilies, including two subfamilies (BURP V and BURP VI) with members from rice only and one subfamily (BURP VII) with members from monocotyledons only. Two BURP gene clusters, belonging to BURP V and BURP VI, were located in the duplicated region on chromosome 5 and 6 of rice, respectively. Transcript level analysis of BURP genes of rice in various tissues and organs revealed different tempo-spatial expression patterns, suggesting that these genes may function at different stages of plant growth and development. Interestingly, all the genes of the BURP VII subfamily were predominantly expressed in flower organs. We also investigated the expression patterns of BURP genes of rice under different stress conditions. The results suggested that, except for two genes (OsBURP01 and OsBURP13), all other members were induced by at least one of the stresses including drought, salt, cold, and abscisic acid treatment. Two genes (OsBURP05 and OsBURP16) were responsive to all the stress treatments and most of the OsBURP genes were responsive to salt stress. Promoter sequence analysis revealed an over-abundance of stress-related cis-elements in the stress-responsive genes. The data presented here provide important clues for elucidating the functions of genes of this family.







Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- ABRE:
-
ABA responsive element
- BURP:
-
BNM2, USP, RD22, and PG1β
- DRE:
-
Dehydration-responsive element
- LTRE:
-
Low temperature-responsive element
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15:63–78
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373
Banzai T, Sumiya K, Hanagata N, Dubinsky Z, Karube I (2002) Molecular cloning and characterization of genes encoding BURP domain-containing protein in the mangrove, Bruguiera gymnorrhiza. Trees 16:87–93
Bassüner R, Bäumlein H, Huth A, Jung R, Wobus U, Rapoport TA, Saalbach G, Müntz K (1988) Abundant embryonic mRNA in field bean (Vicia faba L.) codes for a new class of seed proteins: cDNA cloning and characterization of the primary translation product. Plant Mol Biol 11:321–334
Batchelor AK, Boutilier K, Miller SS, Hattori J, Bowman LA, Hu M, Lantin S, Johnson DA, Miki BL (2002) SCB1, a BURP-domain protein gene, from developing soybean seed coats. Planta 215:523–532
Baumlein H, Boerjan W, Nagy I, Bassuner R, Van Montagu M, Inze D, Wobus U (1991) A novel seed protein gene from Vicia faba is developmentally regulated in transgenic tobacco and Arabidopsis plants. Mol Gen Genet 225:459–467
Boutilier KA, Gines MJ, DeMoor JM, Huang B, Baszczynski CL, Iyer VN, Miki BL (1994) Expression of the BnmNAP subfamily of napin genes coincides with the induction of Brassica microspore embryogenesis. Plant Mol Biol 26:1711–1723
Chen L, Guan L, Seo M, Hoffmann F, Adachi T (2005) Developmental expression of ASG-1 during gametogenesis in apomictic guinea grass (Panicum maximum). J Plant Physiol 162:1141–1148
Datta N, LaFayette PR, Kroner PA, Nagao RT, Key JL (1993) Isolation and characterization of three families of auxin down-regulated cDNA clones. Plant Mol Biol 21:859–869
Finn RD, Mistry J, Schuster-Bockler B, Griffiths-Jones S, Hollich V, Lassmann T, Moxon S, Marshall M, Khanna A, Durbin R, Eddy SR, Sonnhammer EL, Bateman A (2006) Pfam: clans, web tools and services. Nucleic Acids Res 34:D247–D251
Garcia-Hernandez M, Berardini TZ, Chen G, Crist D, Doyle A, Huala E, Knee E, Lambrecht M, Miller N, Mueller LA, Mundodi S, Reiser L, Rhee SY, Scholl R, Tacklind J, Weems DC, Wu Y, Xu I, Yoo D, Yoon J, Zhang P (2002) TAIR: a resource for integrated Arabidopsis data. Funct Integr Genomics 2:239–253
Granger C, Coryell V, Khanna A, Keim P, Vodkin L, Shoemaker RC (2002) Identification, structure, and differential expression of members of a BURP domain containing protein family in soybean. Genome 45:693–701
Hattori J, Boutilier KA, van Lookeren Campagne MM, Miki BL (1998) A conserved BURP domain defines a novel group of plant proteins with unusual primary structures. Mol Gen Genet 259:424–428
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300
Jeon JS, Chung YY, Lee S, Yi GH, Oh BG, An G (1999) Isolation and characterization of an anther-specific gene, RA8, from rice (Oryza sativa L.). Plant Mol Biol 39:35–44
Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Letunic I, Copley RR, Schmidt S, Ciccarelli FD, Doerks T, Schultz J, Ponting CP, Bork P (2004) SMART 4.0: towards genomic data integration. Nucleic Acids Res 32:D142–D144
Liang D, Wu C, Li C, Xu C, Zhang J, Kilian A, Li X, Zhang Q, Xiong L (2006) Establishment of a patterned GAL4-VP16 transactivation system for discovering gene function in rice. Plant J 46:1059–1072
Mulder NJ, Apweiler R (2008) The InterPro database and tools for protein domain analysis. In: Current Protocols Bioinformatics, chap 2, Unit 2.7. Wiley, New York
Narusaka Y, Nakashima K, Shinwari ZK, Sakuma Y, Furihata T, Abe H, Narusaka M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J 34:137–148
Ouyang S, Zhu W, Hamilton J, Lin H, Campbell M, Childs K, Thibaud-Nissen F, Malek RL, Lee Y, Zheng L, Orvis J, Haas B, Wortman J, Buell CR (2007) The TIGR rice genome annotation resource: improvements and new features. Nucleic Acids Res 35:D883–D887
Pla M, Vilardell J, Guiltinan MJ, Marcotte WR, Niogret MF, Quatrano RS, Pages M (1993) The cis-regulatory element CCACGTGG is involved in ABA and water-stress responses of the maize gene rab28. Plant Mol Biol 21:259–266
Ragland M, Soliman KM (1997) Sali5-4a and Sali3-2, two genes induced by aluminum in soybean roots. Plant Physiol 114:395
Ramakers C, Ruijter JM, Deprez RH, Moorman AF (2003) Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339:62–66
Rensing SA, Lang D, Zimmer AD, Terry A, Salamov A, Shapiro H, Nishiyama T, Perroud PF, Lindquist EA, Kamisugi Y, Tanahashi T, Sakakibara K, Fujita T, Oishi K, Shin IT, Kuroki Y, Toyoda A, Suzuki Y, Hashimoto S, Yamaguchi K, Sugano S, Kohara Y, Fujiyama A, Anterola A, Aoki S, Ashton N, Barbazuk WB, Barker E, Bennetzen JL, Blankenship R, Cho SH, Dutcher SK, Estelle M, Fawcett JA, Gundlach H, Hanada K, Heyl A, Hicks KA, Hughes J, Lohr M, Mayer K, Melkozernov A, Murata T, Nelson DR, Pils B, Prigge M, Reiss B, Renner T, Rombauts S, Rushton PJ, Sanderfoot A, Schween G, Shiu SH, Stueber K, Theodoulou FL, Tu H, Van de Peer Y, Verrier PJ, Waters E, Wood A, Yang L, Cove D, Cuming AC, Hasebe M, Lucas S, Mishler BD, Reski R, Grigoriev IV, Quatrano RS, Boore JL (2008) The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319:64–69
Retief JD (2000) Phylogenetic analysis using PHYLIP. Methods Mol Biol 132:243–258
Tang Y-L, Li X-J, Zhong Y-T, Zhang Y-Z (2007) Functional analysis of soybean SALI3-2 in yeast. J Shenzhen Univ Sci Eng 24:324–330
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Treacy BK, Hattori J, Prud’homme I, Barbour E, Boutilier K, Baszczynski CL, Huang B, Johnson DA, Miki BL (1997) Bnm1, a Brassica pollen-specific gene. Plant Mol Biol 34:603–611
Wang A, Xia Q, Xie W, Datla R, Selvaraj G (2003) The classical Ubisch bodies carry a sporophytically produced structural protein (RAFTIN) that is essential for pollen development. Proc Natl Acad Sci USA 100:14487–14492
Ware D (2007) Gramene: a resource for comparative grass genomics. Methods Mol Biol 406:315–330
Watson CF, Zheng L, DellaPenna D (1994) Reduction of tomato polygalacturonase beta subunit expression affects pectin solubilization and degradation during fruit ripening. Plant Cell 6:1623–1634
Xiong L, Yang Y (2003) Disease resistance and abiotic stress tolerance in rice are inversely modulated by an abscisic acid-inducible mitogen-activated protein kinase. Plant Cell 15:745–759
Yamaguchi-Shinozaki K, Shinozaki K (1993) The plant hormone abscisic acid mediates the drought-induced expression but not the seed-specific expression of rd22, a gene responsive to dehydration stress in Arabidopsis thaliana. Mol Gen Genet 238:17–25
Yamaguchi-Shinozaki K, Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 6:251–264
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10:88–94
Yu S, Zhang L, Zuo K, Li Z, Tang K (2004) Isolation and characterization of a BURP domain-containing gene BnBDC1 from Brassica napus involved in abiotic and biotic stress. Physiol Plant 122:210–218
Zheng L, Heupel RC, DellaPenna D (1992) The beta subunit of tomato fruit polygalacturonase isoenzyme 1: isolation, characterization, and identification of unique structural features. Plant Cell 4:1147–1156
Acknowledgments
This work was supported by the grants from the National Program on the Development of Basic Research, the National Program on High Technology Development, the National Natural Science Foundation, and the Ministry of Education of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
X. Ding and X. Hou contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ding, X., Hou, X., Xie, K. et al. Genome-wide identification of BURP domain-containing genes in rice reveals a gene family with diverse structures and responses to abiotic stresses. Planta 230, 149–163 (2009). https://doi.org/10.1007/s00425-009-0929-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-009-0929-z