Abstract
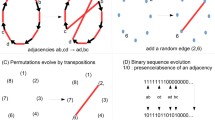
In representing the evolutionary history of a set of binary DNA sequences by a connected graph, a set theoretical approach is introduced for studying recombination events. We show that set theoretical constraints have direct implications on the number of recombination events. We define a new lower bound on the number of recombination events and demonstrate the usefulness of our new approach through several explicit examples.
Similar content being viewed by others
References
Hein, J.: A Heuristic Method to Reconstruct the History of Sequences Subject to Recombination. J. Mol. Evol. 20, 402–411 (1993)
Hudson, R.R., Kaplan, N.L.: Statistical Properties of the Number of Recombination Events in the History of a Sample of DNA Sequences. Genetics 11, 147–164 (1985)
Kreitman, M.: Nucleotide Polymorphism at the Alcohol Dehydrogenase Locus of Drosophila Melanogaster. Nature 304, 412–417 (1983)
Myers, S.R., Griffiths, R.C.: Bounds on the Minimum Number of Recombination Events in a Sample History. Genetics 163, 375–394 (2003)
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding author: Y.S. Song Preferred postal address: Department of Statistics, University of Oxford, 1 South Parks Road, Oxford, OX1 3TG, UK.
Rights and permissions
About this article
Cite this article
Song, Y., Hein, J. On the minimum number of recombination events in the evolutionary history of DNA sequences. J. Math. Biol. 48, 160–186 (2004). https://doi.org/10.1007/s00285-003-0227-5
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-003-0227-5