Abstract
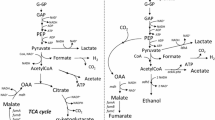
To utilize fermentative bacteria for producing the alternative fuel hydrogen, we performed successive rounds of P1 transduction from the Keio Escherichia coli K-12 library to introduce multiple, stable mutations into a single bacterium to direct the metabolic flux toward hydrogen production. E. coli cells convert glucose to various organic acids (such as succinate, pyruvate, lactate, formate, and acetate) to synthesize energy and hydrogen from formate by the formate hydrogen-lyase (FHL) system that consists of hydrogenase 3 and formate dehydrogenase-H. We altered the regulation of FHL by inactivating the repressor encoded by hycA and by overexpressing the activator encoded by fhlA, removed hydrogen uptake activity by deleting hyaB (hydrogenase 1) and hybC (hydrogenase 2), redirected glucose metabolism to formate by using the fdnG, fdoG, narG, focA, focB, poxB, and aceE mutations, and inactivated the succinate and lactate synthesis pathways by deleting frdC and ldhA, respectively. The best of the metabolically engineered strains, BW25113 hyaB hybC hycA fdoG frdC ldhA aceE, increased hydrogen production 4.6-fold from glucose and increased the hydrogen yield twofold from 0.65 to 1.3 mol H2/mol glucose (maximum, 2 mol H2/mol glucose).

Similar content being viewed by others
References
Abdel-Hamid AM, Attwood MM, Guest JR (2001) Pyruvate oxidase contributes to the aerobic growth efficiency of Escherichia coli. Microbiology 147:1483–1498
Andrews SC, Berks BC, McClay J, Ambler A, Quail MA, Golby P, Guest JR (1997) A 12-cistron Escherichia coli operon (hyf) encoding a putative proton-translocating formate hydrogenlyase system. Microbiology 143:3633–3647
Angelides KJ, Akiyama SK, Hammes GG (1979) Subunit stoichiometry and molecular weight of the pyruvate dehydrogenase multienzyme complex from Escherichia coli. Proc Natl Acad Sci U S A 76:3279–3283
Archana A, Sasikala C, Ramana Ch V (2003) Augmentation of H2 photoproduction in Rhodopseudomonas palustris by N-heterocyclic aromatic compounds. Biotechnol Lett 25:79–82
Axley MJ, Grahame DA, Stadtman TC (1990) Escherichia coli formate-hydrogen lyase. Purification and properties of the selenium-dependent formate dehydrogenase component. J Biol Chem 265:18213–18218
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:2006–0008
Bagramyan K, Trchounian A (2003) Structural and functional features of formate hydrogen lyase, an enzyme of mixed-acid fermentation from Escherichia coli. Biochemistry (Mosc) 68: 1159–1170
Bertero MG, Rothery RA, Palak M, Hou C, Lim D, Blasco F, Weiner JH, Strynadka NC (2003) Insights into the respiratory electron transfer pathway from the structure of nitrate reductase A. Nat Struct Biol 10:681–687
Blattner FR, Plunkett G 3rd, Bloch CA, Perna NT, Burland V, Riley M, Collado-Vides J, Glasner JD, Rode CK, Mayhew GF, Gregor J, Davis NW, Kirkpatrick HA, Goeden MA, Rose DJ, Mau B, Shao Y (1997) The complete genome sequence of Escherichia coli K-12. Science 277:1453–1474
Böck A, Sawers G (1996) Cellular and Molecular Biology. In: Neidhardt FC, Curtiss JR II, Ingraham JL, Lin ECC, Low KB, Magasanik B, Reznikoff WS, Riley M, Schaechter M, Umbarger HE (eds) Escherichia coli and Salmonella. 2nd edn. ASM Press, Washington, pp 262–282
Cherepanov PP, Wackernagel W (1995) Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 158:9–14
Das D, Veziroğlu TN (2001) Hydrogen production by biological processes: a survey of literature. Int J Hydrogen Energy 26:13–28
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Dunn S (2002) Hydrogen futures: toward a sustainable energy system. Int J Hydrogen Energy 27:235–264
Evans DJ, Pickett CJ (2003) Chemistry and the hydrogenases. Chem Soc Rev 32:268–275
Forzi L, Sawers RG (2007) Maturation of [NiFe]-hydrogenases in Escherichia coli. Biometals 20:565–578
Glick BR, Wang PY, Schneider H, Martin WG (1980) Identification and partial characterization of an Escherichia coli mutant with altered hydrogenase activity. Can J Biochem 58:361–367
Hansel A, Lindblad P (1998) Toward optimization of cyanobacteria as biotechnologically relevant producers of molecular hydrogen, a clean and renewable energy source. Appl Microbiol Biotechnol 50:153–160
Iverson TM, Luna-Chavez C, Cecchini G, Rees DC (1999) Structure of the Escherichia coli fumarate reductase respiratory complex. Science 284:1961–1966
Jung GY, Jung HO, Kim JR, Ahn Y, Park S (1999) Isolation and characterization of Rhodopseudomonas palustris P4 which utilizes CO with the production of H2. Biotechnol Lett 21:525–529
Kitagawa M, Ara T, Arifuzzaman M, Ioka-Nakamichi T, Inamoto E, Toyonaga H, Mori H (2005) Complete set of ORF clones of Escherichia coli ASKA library (A complete set of E. coli K-12 ORF archive): unique resources for biological research. DNA Res 12:291–299
Klibanov AM, Alberti BN, Zale SE (1982) Enzymatic synthesis of formic acid from H2 and CO2 and production of hydrogen from formic acid. Biotechnol Bioeng 24:25–36
Kraemer JT, Bagley DM (2007) Improving the yield from fermentative hydrogen production. Biotechnol Lett 29:685–695
Lee SJ, Lee DY, Kim TY, Kim BH, Lee J, Lee SY (2005) Metabolic engineering of Escherichia coli for enhanced production of succinic acid, based on genome comparison and in silico gene knockout simulation. Appl Environ Microbiol 71:7880–7887
Maeda T, Sanchez-Torres V, Wood TK (2007a) Escherichia coli hydrogenase 3 is a reversible enzyme possessing hydrogen uptake and synthesis activities. Appl Microbiol Biotechnol 76:1035–1042
Maeda T, Sanchez-Torres V, Wood TK (2007b) Metabolic engineering to enhance bacterial hydrogen production. Microb Biotechnol. doi:https://doi.org/10.1111/j.1751-7915.2007.00003.x
Maeda T, Vardar G, Self WT, Wood TK (2007c) Inhibition of hydrogen uptake in Escherichia coli by expressing the hydrogenase from the cyanobacterium Synechocystis sp. PCC 6803. BMC Biotechnol 7:25
Mat-Jan F, Alam KY, Clark DP (1989) Mutants of Escherichia coli deficient in the fermentative lactate dehydrogenase. J Bacteriol 171:342–348
Minnan L, Jinli H, Xiaobin W, Huijuan X, Jinzao C, Chuannan L, Fengzhang Z, Liangshu X (2005) Isolation and characterization of a high H2-producing strain Klebsiella oxytoca HP1 from a hot spring. Res Microbiol 156:76–81
Oh Y-K, Seol E-H, Kim JR, Park S (2003) Fermentative biohydrogen by a new chemoheterotrophic bacterium Citrobacter sp. Y19. Int J Hydrogen Energy 28:1353–1359
Patnaik R, Louie S, Gavrilovic V, Perry K, Stemmer WP, Ryan CM, del Cardayré S (2002) Genome shuffling of Lactobacillus for improved acid tolerance. Nat Biotechnol 20:707–712
Penfold DW, Forster CF, Macaskie LE (2003) Increased hydrogen production by Escherichia coli strain HD701 in comparison with the wild-type parent strain MC4100. Enzyme Microb Technol 33:185–189
Penfold DW, Sargent F, Macaskie LE (2006) Inactivation of the Escherichia coli K-12 twin-arginine translocation system promotes increased hydrogen production. FEMS Microbiol Lett 262:135–137
Rachman MA, Furutani Y, Nakashimada Y, Kakizono T, Nishio N (1997) Enhanced hydrogen production in altered mixed acid fermentation of glucose by Enterobacter aerogenes. J Ferment Bioeng 83:358–363
Rossmann R, Sawers G, Böck A (1991) Mechanism of regulation of the formate-hydrogenlyase pathway by oxygen, nitrate, and pH: definition of the formate regulon. Mol Microbiol 5:2807–2814
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Schlensog V, Lutz S, Böck A (1994) Purification and DNA-binding properties of FHLA, the transcriptional activator of the formate hydrogenlyase system from Escherichia coli. J Biol Chem 269:19590–19596
Silhavy TJ, Berman ML, Enquist LW (1984) Experiments with gene fusions. Cold Spring Harbor Laboratories, Cold Spring Habor, NY
Sode K, Watanabe M, Makimoto H, Tomiyama M (1999) Construction and characterization of fermentative lactate dehydrogenase Escherichia coli mutant and its potential for bacterial hydrogen production. Appl Biochem Biotech 77–79:317–323
Suppmann B, Sawers G (1994) Isolation and characterization of hypophosphite-resistant mutants of Escherichia coli: identification of the FocA protein, encoded by the pfl operon, as a putative formate transporter. Mol Microbiol 11:965–982
Wang H, Gunsalus RP (2003) Coordinate regulation of the Escherichia coli formate dehydrogenase fdnGHI and fdhF genes in response to nitrate, nitrite, and formate: roles for NarL and NarP. J Bacteriol 185:5076–5085
Woods DD (1936) Hydrogenlyases: the synthesis of formic acid by bacteria. Biochem J 30:515–527
Yi KB, Harrison DP (2005) Low-pressure sorption-enhanced hydrogen production. Ind Eng Chem Res 44:1665–1669
Yoshida A, Nishimura T, Kawaguchi H, Inui M, Yukawa H (2005) Enhanced hydrogen production from formic acid by formate hydrogen lyase-overexpressing Escherichia coli strains. Appl Environ Microbiol 71:6762–6768
Yoshida A, Nishimura T, Kawaguchi H, Inui M, Yukawa H (2006) Enhanced hydrogen production from glucose using ldh- and frd-inactivated Escherichia coli strains. Appl Microbiol Biotechnol 73:67–72
Acknowledgment
The authors thank the National of Institute of Genetics, Japan, for sending the Keio and ASKA clones. This research was supported by DARPA (HR0011–06–1–0001).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maeda, T., Sanchez-Torres, V. & Wood, T.K. Enhanced hydrogen production from glucose by metabolically engineered Escherichia coli . Appl Microbiol Biotechnol 77, 879–890 (2007). https://doi.org/10.1007/s00253-007-1217-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-1217-0