Abstract
In the African clawed frog (Xenopus laevis), two deeply divergent allelic lineages of multiple genes of the class I MHC region have been discovered. For the MHC class I UAA locus, functional differences and the molecular basis for lineages maintenance are unknown. Alleles of linked class I region genes also exhibit strong disequilibrium with specific MHC alleles, but the underlying cause is not clear. We use MHC class Ia sequence data to estimate substitution rates and investigate structural differences between allelic lineages from protein models. Results indicate the operation of natural selection, and differences in the steric properties in the F pocket of the peptide-binding region among lineages. Variability in this pocket likely enables allelic lineages to bind very different sets of peptides and to interact differently with MHC chaperones in the endoplasmic reticulum. These results constitute evidence of the molecular evolutionary basis for 1) the maintenance of allelic lineages, 2) functional differences among lineages, and 3) strong linkage disequilibrium of allelic variants of class I region genes in X. laevis.



Similar content being viewed by others
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution 10.1093/bioinformatics/bti263. Bioinformatics 21:2104–2105
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Automat Contr 19:716–723
Andolfatto P, Nordborg M (1998) The effect of gene conversion on intralocus associations. Genetics 148:1397–1399
Anisimova M, Bielawski JP, Yang Z (2002) Accuracy and power of Bayes prediction of amino acid sites under positive selection. Mol Biol Evol 19:950–958
Anisimova M, Nielsen R, Yang Z (2003) Effect of recombination on the accuracy of the likelihood method for detecting positive selection at amino acid sites. Genetics 164:1229–1236
Apanius V, Penn DJ, Slev P, Ruff LR, Potts WK (1997) The nature of selection on the major histocompatibility complex. Crit Rev Immunol 17:179–224
Beiβarth T, Sun J, Kavathas PB, Ortmann B (2000) Increased efficiency of folding and peptide loading of mutant MHC class I molecules. Eur J Immunol 30:1203–1213
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Bos DH, Posada D (2005) Using models of nucleotide evolution to build phylogenetic trees. Dev Comp Immunol 29:211–227
Bos DH, Waldman B (2006) Evolution by recombination and transspecies polymorphism in the MHC class I gene of Xenopus laevis. Mol Biol Evol 23:137–143
Boyington JC, Motyka SA, Schuck P, Brooks AG, Sun PD (2000) Crystal strucuture of an NK cell immunolglobulin-like receptor in complex with its class I MHC ligand. Nature 405:537–543
Burnham KP, Anderson DR (2002) Model selection and multimodel inference: a practical information-theoretic approach. Springer, Berlin Heidelberg New York
Carrington M, O’Brien SJ (2003) The influence of HLA genotype on AIDS. Annu Rev Med 54:535–551
Crandall KA, Kelsey CR, Imamichi H, Lane HC, Salzman NP (1999) Parallel evolution of drug resistance in HIV: failure of non-synonymous/synonymous substitution rate ratio to detect selection. Mol Biol Evol 16:372–382
Doherty P, Zinkernagel RM (1975) Enhanced immunological surveillance in mice heterozygous at the H-2 gene complex. Nature 256:50–52
Elliott T, Williams A (2005) The optimization of peptide cargo bound to MHC class I molecules by the peptide-loading complex. Immunol Rev 207:89–99
Ferrara GB, Bacigalupo A, Lamparelli T, Lanino E, Delfino L, Morabito A, Parodi AM, Pera C, Pozzi S, Sormani MP, Bruzzi P, Bordo D, Bolognesi M, Bandini G, Bontadini A, Barbanti M, Frumento G (2001) Bone marrow transplantation from unrelated donors: the impact of mismatches with substitutions at position 116 of the human leukocyte antigen class I heavy chain. Blood 98:3150–3155
Flajnik MF, Kaufman J, Riegert P, Du Pasquier L (1984) Identification of class I major histocompatibility complex encoded molecules in the Amphibian Xenopus. Immunogenetics 20:134–143
Flajnik MF, Ohta Y, Greenberg AS, Salter-Cid L, Carrizosa A, Du Pasquier L, Kasahara M (1999) Two ancient allelic lineages at the single classical class I locus in the Xenopus MHC. J Immunol 163:3826–3833
Gao GF, Tormo J, Gerth UC, Wyer JR, McMichael AJ, Stuart DI, Bell JI, Jones EY, Jakobsen BK (1997) Crystal structure of the complex between human CD8aa and HLA-A2. Nature 387:630–634
Garbi N, Tanaka S, van den Broek M, Momburg F, Hammerling GJ (2005) Accessory molecules in the assembly of major histocompatibility complex class I/peptide complexes: how essential are they for CD8+ T-cell immune responses? Immunol Rev 207:77–88
Garboczi DN, Ghosh P (1996) Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 384:134–141
Grandea AG III, Van Kaer L (2001) Tapasin: an ER chaperone that controls MHC class I assembly with peptide. Trends Immunol 22:194–199
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-Pdbviewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Guex N, Diemand A, Peitsch MC (1999) Protein modeling for all. Trends Biochem Sci 29:364–367
Guindon S, Gascuel O (2003) A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Syst Biol 52:696–704
Harris MR, Yu YYL, Kindle CS, Hansen TH, Solheim JC (1998) Calreticulin and calnexin interact with different protein and glycan determinants during the assembly of MHC class I. J Immunol 160:5404–5409
Hashimoto K, Okamura K, Yamaguchi H, Ototake M, Nakanishi T, Kurosawa Y (1999) Conservation and diversification of MHC class I and its related molecules in vertebrates. Immunol Rev 167:81–100
Hildebrand WH, Turnquist HR, Prilliman KR, Hickman HD, Schenk EL, McIlhaney MM, Solheim JC (2002) HLA class I polymorphism has a dual impact on ligand binding and chaperone interaction. Hum Immunol 63:248–255
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335:167–170
Hulsmeyer M, Hillig RC, Volz A, Ruhl M, Schroder W, Saenger W, Ziegler A, Uchanska-Ziegler B (2002) HLA-B27 subtypes differentially associated with disease exhibit subtle structural alterations. J Biol Chem 277:47844–47853
Ina Y (1995) New methods for estimating the numbers of synonymous and non-synonymous substitutions. J Mol Evol 40:190–226
Joly E, Le Rolle AF, Gonzalez AL, Mehling B, Stevens J, Coadwell WJ, Hunig T, Howard JC, Butcher GW (1998) Co-evolution of rat TAP transporters and MHC class I RT1-A molecules. Curr Biol 8:169–172
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8:275–282
Kaufman J (1999) Co-evolving genes in MHC haplotypes: the “rule” for nonmammalian vertebrates? Immunogenetics 50:228–236
Kaufman J, Anderson R, Avila D, Engberg J, Lambris J, Salomonsen J, Welinder K, Skjodt K (1992) Different features of the MHC class I heterodimer have evolved at different rates. J Immunol 142:1532–1546
Kaufman J, Salomonsen J, Flajnik MF (1994) Evolutionary conservation of MHC class I and class II molecules—different yet the same. Semin Immunol 6:411–424
Maruyama T, Nei M (1981) Genetic variability maintained by mutation and overdominant selection in finite populations. Genetics 98:441–459
Muse SV (1996) Estimating synonymous and non-synonymous substitution rates. Mol Biol Evol 13:105–114
Namikawa C, Salter-Cid L, Flajnik MF, Kato Y, Nonaka M, Sasaki M (1995) Isolation of Xenopus LMP-7 homologues: striking allelic diversity and linkage to MHC. J Immunol 155:1964–1971
Neisig A, Wubbolts R, Zang X, Melief C, Neefjes J (1996) Allele-specific differences in the interaction of MHC class I molecules with transporters associated with antigen processing. J Immunol 156:3196–3206
Nonaka M, Namikawa C, Kato Y, Sasaki M, Salter-Cid L, Flajnik MF (1997a) Major histocompatibility complex gene mapping in the amphibian Xenopus implies a primordial organization. Proc Natl Acad Sci U S A 94:5789–5791
Nonaka M, Namikawa-Yamada C, Sasaki M, Salter-Cid L, Flajnik MF (1997b) Evolution of proteasome subunits d and LMP2. J Immunol 159:734–740
Ohta Y, Powis SJ, Coadwell WJ, Haliniewski DE, Liu Y, Li H, Flajnik MF (1999) Identification and mapping of Xenopus TAP2 genes. Immunogenetics 49:171–182
Ohta Y, McKinney EC, Criscitiello MF, Flajnik MF (2002) Proteosome, transporter associated with antigen processing, and class I genes in the nurse shark Ginglymostoma cirratum: evidence for a stable class I region and MHC haplotype lineages. J Immunol 168:771–781
Ohta Y, Powis SJ, Lohr RL, Nonaka M, Du Pasquier L, Flajnik MF (2003) Two highly divergent ancient allelic lineages of the transporter associated with antigen processing (TAP) gene in Xenopus: further evidence for co-evolution among MHC class I region genes. Eur J Immunol 33:3017–3027
Paquet M-E, Williams DB (2002) Mutant MHC class I molecules define interaction between components of the peptide-loading complex. Int Immunol 14:347–358
Parham P, Ohta T (1996) Population biology of antigen presentation by MHC class I molecules. Science 272:67–74
Parham P, Lomen CE, Lawlor DA, Ways JP, Holmes N, Coppin HL, Salter RD, Wan AM, Ennis PD (1988) Nature of polymorphism in HLA-A, -B, -C molecules. Proc Natl Acad Sci U S A 85:4005–4009
Rozas J, Rozas R (1999) DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15:174–175
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor, Cold Spring Harbor, New York
Saper MA, Bjorkman PJ, Wiley DC (1991) Refined structure of the human histocompatibility antigen HLA-A2 at 2.6 A resolution. J Mol Biol 219:277–319
Schwede T, Kopp J, Guex N, Peitsch MC (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385
Sharp PM (1997) In search of molecular Darwinism. Nature 385:111–112
Shum BP, Guethlein LA, Flodin LR, Adkinson MA, Hedrick RP, Nehring RB, Stet RJM, Secombes C, Parham P (2001) Modes of salmon MHC class I and II evolution differ from the primate paradigm. J Immunol 166:3297–3308
Sobolev V, Sorokine A, Prilusky J, Abola E, Edelman M (1999) Automated analysis of interatomic contacts in protiens. Bioinformatics 15:327–332
Suzuki Y, Gojobori T (1999) A method for detecting positive selection at single amino acid sites. Mol Biol Evol 16:1315–1328
Suzuki Y, Nei M (2004) False-positive selection identified by ml-based methods: examples from the Sig1 gene of the diatom Thalassiosira weissflogii and the tax gene of a human T-cell lymphotropic virus. Mol Biol Evol 21:914–921
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Turnquist HR, Thomas HJ, Prilliman KR, Lutz CT, Hildebrand WH, Solheim JC (2000) HLA-B polymorphism affects interactions with multiple endoplasmic reticulum proteins. Eur J Immunol 30:3021–3028
Williams AP, Au Peh C, Purcell AW, McClusky J, Elliot T (2002) Optimization of the MHC class I peptide cargo is dependent on tapasin. Immunity 16:509–520
Wong WSW, Yang Z, Goldman N, Nielsen R (2004) Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics 168:1041–1051
Wu TT, Kabat EA (1970) An analysis of the sequences of the variable regions of Bence Jones proteins and myeloma light chains and their implications for antibody complementarity. J Exp Med 132:211–250
Yang Z (2000) Phylogenetic analysis by maximum likelihood (PAML). University College London, London
Yang Z (2001) Adaptive molecular evolution. In: Balding DJ, Cannings C, Bishop M (eds) Handbook of statistical genetics. Wiley, New York, pp 327–350
Yang Z, Bielawski JP (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15:496–502
Yang Z, Nielsen R, Goldman N, Pedersen A-MK (2000) Codon substitution models for heterogeneous selection pressure and amino acid sites. Genetics 155:431–449
Yu YYL, Turnquist HR, Myers NB, Balendiran GK, Hansen TH, Solheim JC (1999) An extensive region of an MHC class I a2 domain loop influences interaction with assembly complex. J Immunol 163:4427–4433
Zernich D, Purcell AW, Macdonald WA, Kjer-Nielsen L, Ely LK, Laham N, Crockford T, Mifsud NA, Bharadwaj M, Chang L, Tait BD, Holdsworth R, Brooks AG, Bottomley SP, Beddoe T, Peh CA, Rossjohn J, McCluskey J (2004) Natural HLA class i polymorphism controls the pathway of antigen presentation and susceptibility to viral evasion. J Exp Med 200:13–24
Zhang C, Anderson A, DeLisi C (1998) Structural principles that govern the peptide-binding motifs of class I MHC molecules. J Mol Biol 281:929–947
Acknowledgements
Thanks to Martin Flajnik, Neil Gemmell, the DeWoody lab group, and anonymous peer reviewers for suggestions on earlier versions of this manuscript. Funding is provided by the Marsden Fund (Royal Society of New Zealand) and a Ph.D. scholarship from the University of Canterbury.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the supplementary material.
Supplement 1
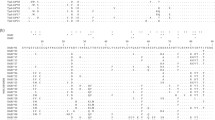
Phylogenetic trees of amino acid sequence from the MHC class Ia of Xenopus laevis. Trees reconstructed under the maximum likelihood criterion using the JTT model of amino acid changes with a gamma distribution of rate variation among sites. The model was selected as the best-fit model using Akaike’s Information Criterion in the program PROTTEST. Numbers indicate bootstrap support for nodes, and nodes with less than 50% bootstrap support are collapsed into unresolved polytomies. These topologies feature a general lack of resolution due to the short length of the sequences and ambiguous phylogenetic signal (PDF 32 kb)
Supplement 2
(PDF 62 kb)
Rights and permissions
About this article
Cite this article
Bos, D.H., Waldman, B. Polymorphism, natural selection, and structural modeling of class Ia MHC in the African clawed frog (Xenopus laevis). Immunogenetics 58, 433–442 (2006). https://doi.org/10.1007/s00251-006-0114-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-006-0114-5