Abstract
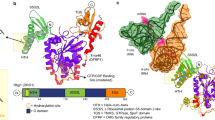
The testis-specific gene Jingwei (jgw) is a newly evolved short-chain dehydrogenase/reductase in Drosophila. Preliminary substrate screening indicated that JGW prefers long-chain primary alcohols as substrates, including several exotic alcohols such as farnesol and geraniol. Using steady-state kinetics analyses and molecular docking, we not only confirmed JGW’s substrate specificity, but also demonstrated that the new enzymatic activities of JGW evolved extensively after exon-shuffling from a preexisting enzyme. Analysis of JGW orthologs in sister species shows that subsequent evolutionary changes following the birth of JGW altered substrate specificities and enzyme stabilities. Our results lend support to a general mechanism for the evolution of a new enzyme, in which catalytic chemistry evolves first followed by diversification of substrate utilization.




Similar content being viewed by others
Abbreviations
- JGW:
-
Jingwei
- SDR:
-
Short-chain dehydrogenase/reductase
- ADH:
-
Alcohol dehydrogenase
References
Albery WJ, Knowles JR (1976) Evolution of enzyme function and the development of catalytic efficiency. Biochemistry 15:5631–5640
Babbitt PC, Gerlt JA (1997) Understanding enzyme superfamilies. Chemistry as the fundamental determinant in the evolution of new catalytic activities. J Biol Chem 272:30591–30594
Babbitt PC, Mrachko GT, Hasson MS, Huisman GW, Kolter R, Ringe D, Petsko GA, Kenyon GL, Gerlt JA (1995) A functionally diverse enzyme superfamily that abstracts the alpha protons of carboxylic acids. Science 267:1159–1161
Benach J, Atrian S, Gonzalez-Duarte R, Ladenstein R (1998) The refined crystal structure of Drosophila lebanonensis alcohol dehydrogenase at 1.9 A resolution. J Mol Biol 282:383–399
Benach J, Atrian S, Gonzalez-Duarte R, Ladenstein R (1999) The catalytic reaction and inhibition mechanism of Drosophila alcohol dehydrogenase: observation of an enzyme-bound NAD-ketone adduct at 1.4 A resolution by X-ray crystallography. J Mol Biol 289:335–355
Benach J, Atrian S, Ladenstein R, Gonzalez-Duarte R (2001) Genesis of Drosophila ADH: the shaping of the enzymatic activity from a SDR ancestor. Chem Biol Interact 130–132:405–415
Benach J, Winberg JO, Svendsen JS, Atrian S, Gonzalez-Duarte R, Ladenstein R (2005) Drosophila alcohol dehydrogenase: acetate-enzyme interactions and novel insights into the effects of electrostatics on catalysis. J Mol Biol 345:579–598
Bustamante CD, Townsend JP, Hartl DL (2000) Solvent accessibility and purifying selection within proteins of Escherichia coli and Salmonella enterica. Mol Biol Evol 17:301–308
Chambers GK (1991) Gene expression, adaptation and evolution in higher organisms. Evidence from studies of Drosophila alcohol dehydrogenases. Comp Biochem Physiol B 99:723–730
Chen Z, Lu L, Shirley M, Lee WR, Chang SH (1990) Site-directed mutagenesis of glycine-14 and two “critical” cysteinyl residues in Drosophila alcohol dehydrogenase. Biochemistry 29:1112–1118
Chenevert SW, Fossett NG, Chang SH, Tsigelny I, Baker ME, Lee WR (1995) Amino acids important in enzyme activity and dimer stability for Drosophila alcohol dehydrogenase. Biochem J 308(Pt 2):419–423
Dean AM, Neuhauser C, Grenier E, Golding GB (2002) The pattern of amino acid replacements in alpha/beta-barrels. Mol Biol Evol 19:1846–1864
Golding GB, Dean AM (1998) The structural basis of molecular adaptation. Mol Biol Evol 15:355–369
Heinstra PW, Geer BW, Seykens D, Langevin M (1989) The metabolism of ethanol-derived acetaldehyde by alcohol dehydrogenase (EC 1.1.1.1) and aldehyde dehydrogenase (EC 1.2.1.3) in Drosophila melanogaster larvae. Biochem J 259:791–797
Henehan GT, Chang SH, Oppenheimer NJ (1995) Aldehyde dehydrogenase activity of Drosophila melanogaster alcohol dehydrogenase: burst kinetics at high pH and aldehyde dismutase activity at physiological pH. Biochemistry 34:12294–12301
Horowitz NH (1945) On the Evolution of Biochemical Syntheses. Proc Natl Acad Sci USA 31:153–157
Hsu CC, Hong Z, Wada M, Franke D, Wong CH (2005) Directed evolution of d-sialic acid aldolase to l-3-deoxy-manno-2-octulosonic acid (l-KDO) aldolase. Proc Natl Acad Sci USA 102:9122–9126
Jornvall H, Persson B, Krook M, Atrian S, Gonzalez-Duarte R, Jeffery J, Ghosh D (1995) Short-chain dehydrogenases/reductases (SDR). Biochemistry 34:6003–6013
Juan E, Gonzalez-Duarte R (1981) Determination of some biochemical and structural features of alcohol dehydrogenases from Drosophila simulans and Drosophila virilis. Comparison of their properties with the Drosophila melanogaster Adhs enzyme. Biochem J 195:61–69
Kaessmann H, Vinckenbosch N, Long M (2009) RNA-based gene duplication: mechanistic and evolutionary insights. Nat Rev Genet 10:19–31
Ladenstein R, Winberg JO, Benach J (2008) Medium- and short-chain dehydrogenase/reductase gene and protein families: Structure-function relationships in short-chain alcohol dehydrogenases. Cell Mol Life Sci 65:3918–3935
Long M, Langley CH (1993) Natural selection and the origin of jingwei, a chimeric processed functional gene in Drosophila. Science 260:91–95
Long M, Wang W, Zhang J (1999) Origin of new genes and source for N-terminal domain of the chimerical gene, jingwei, in Drosophila. Gene 238:135–141
Lunzer M, Miller SP, Felsheim R, Dean AM (2005) The biochemical architecture of an ancient adaptive landscape. Science 310:499–501
Oppermann U, Filling C, Hult M, Shafqat N, Wu X, Lindh M, Shafqat J, Nordling E, Kallberg Y, Persson B, Jornvall H (2003) Short-chain dehydrogenases/reductases (SDR): the 2002 update. Chem Biol Interact 143–144:247–253
Oue S, Okamoto A, Yano T, Kagamiyama H (1999) Redesigning the substrate specificity of an enzyme by cumulative effects of the mutations of non-active site residues. J Biol Chem 274:2344–2349
Pace CN (1990) Measuring and increasing protein stability. Trends Biotechnol 8:93–98
Perona JJ, Craik CS (1997) Evolutionary divergence of substrate specificity within the chymotrypsin-like serine protease fold. J Biol Chem 272:29987–29990
Petsko GA, Kenyon GL, Gerlt JA, Ringe D, Kozarich JW (1993) On the origin of enzymatic species. Trends Biochem Sci 18:372–376
Rothman SC, Voorhies M, Kirsch JF (2004) Directed evolution relieves product inhibition and confers in vivo function to a rationally designed tyrosine aminotransferase. Protein Sci 13:763–772
Scopes RK (1996) Protein purification principles and practice. Springer, New York
Wang W, Zhang J, Alvarez C, Llopart A, Long M (2000) The origin of the Jingwei gene and the complex modular structure of its parental gene, yellow emperor, in Drosophila melanogaster. Mol Biol Evol 17:1294–1301
Weinreich DM, Delaney NF, Depristo MA, Hartl DL (2006) Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312:111–114
Winberg JO, McKinley-McKee JS (1988) Drosophila melanogaster alcohol dehydrogenase. Biochemical properties of the NAD+-plus-acetone-induced isoenzyme conversion. Biochem J 251:223–227
Winberg JO, McKinley-McKee JS (1992) Kinetic interpretations of active site topologies and residue exchanges in Drosophila alcohol dehydrogenases. Int J Biochem 24:169–181
Winberg JO, McKinley-McKee JS (1998) Drosophila melanogaster alcohol dehydrogenase: mechanism of aldehyde oxidation and dismutation. Biochem J 329(Pt 3):561–570
Winberg JO, Thatcher DR, McKinley-McKee JS (1982) Alcohol dehydrogenase from the fruitfly Drosophila melanogaster. Substrate specificity of the alleloenzymes AdhS and AdhUF. Biochim Biophys Acta 704:7–16
Winberg JO, Hovik R, McKinley-McKee JS, Juan E, Gonzalez-Duarte R (1986) Biochemical properties of alcohol dehydrogenase from Drosophila lebanonensis. Biochem J 235:481–490
Winberg JO, Brendskag MK, Sylte I, Lindstad RI, McKinley-McKee JS (1999) The catalytic triad in Drosophila alcohol dehydrogenase: pH, temperature and molecular modelling studies. J Mol Biol 294:601–616
Zhang J, Dean AM, Brunet F, Long M (2004) Evolving protein functional diversity in new genes of Drosophila. Proc Natl Acad Sci USA 101:16246–16250
Zhang J, Long M, Li L (2005) Translational effects of differential codon usage among intragenic domains of new genes in Drosophila. Biochim Biophys Acta 1728(3):135–142
Acknowledgments
We thank all the members in the Dean laboratory at the University of Minnesota, Li laboratory at Northwestern University, and Long laboratory at the University of Chicago for experimental assistance and helpful discussions. This study was supported by grants to A.M.D. from the National Institutes of Health, L.L from National Institute of Health (R01NS056086), and M.L. from the National Science Foundation and the National Institutes of Health, a National Science Foundation CAREER award, and a Packard Fellowship in Science and Engineering. Hinds fund from the University of Chicago to J.Z.
Author information
Authors and Affiliations
Corresponding authors
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
239_2010_9384_MOESM1_ESM.doc
Supplementary material: Supplementary material includes data of structural based sequence alignment and Ramachandran Plot of JGW protein: D. teissieri and D. yakuba JGW. (DOC 329 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Yang, H., Long, M. et al. Evolution of Enzymatic Activities of Testis-Specific Short-Chain Dehydrogenase/Reductase in Drosophila . J Mol Evol 71, 241–249 (2010). https://doi.org/10.1007/s00239-010-9384-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-010-9384-5