Abstract
Enamelin (ENAM) plays an important role in the mineralization of the forming enamel matrix. We have performed an evolutionary analysis of mammalian ENAM to identify highly conserved residues or regions that could have important function (selective pressure), to predict mutations that could be associated with amelogenesis imperfecta in humans, and to identify possible adaptive evolution of ENAM during 200 million years ago of mammalian evolution. In order to fulfil these objectives, we obtained 36-ENAM sequences that are representative of the mammalian lineages. Our results show a remarkably high conservation pattern in the region of the 32-kDa fragment of ENAM, especially its phosphorylation, glycosylation, and proteolytic sites. In primates and rodents we also identified several sites under positive selection, which could indicate recent evolutionary changes in ENAM function. Furthermore, the analysis of the unusual signal peptide provided new insights on the possible regulation of ENAM secretion, a hypothesis that should be tested in the near future. Taken together, these findings improve our understanding of ENAM evolution and provide new information that would be useful for further investigation of ENAM function as well as for the validation of mutations leading to amelogenesis imperfecta.










Similar content being viewed by others
References
Aoba T, Moreno EC (1987) The enamel fluid in the early secretory stage of porcine amelogenesis: chemical composition and saturation with respect to enamel mineral. Calcif Tissue Int 41:86–94
Brookes SJ, Lyngstadaas SP, Robinson C, Shore RC, Wood SR, Kirkham J (2002) Enamelin compartmentalization in developing porcine enamel. Connect Tissue Res 43:477–481
Chen Y-C, Peng G-S, Wang M-F, Tsao T-P, Yin S-J (2009) Polymorphism of ethanol-metabolism genes and alcoholism: correlation of allelic variations with the pharmacokinetic and pharmacodynamic consequences. Chem Biol Int 178:2–7
Davis MJ, Hanson KA, Clark F, Fink JL, Zhang F, Kasukawa T, Kai C, Kawai J, Carninci P, Hayashizaki Y, Teasdale RD (2006) Differential use of signal peptides and membrane domains is a common occurrence in the protein output of transcriptional units. PLoS Genet 2:e46
Davit-Béal T, Tucker T, Sire JY (2009) Loss of teeth and enamel in tetrapods: fossil record, genetic data and morphological adaptations. J Anat 214:277–501
Dayhoff MO, Schwartz R, Orcutt BC (1978) A model of evolutionary change in proteins, matrixes for detecting distant relationships. In: Dayhoff MO (ed) Atlas of protein sequence and structure, vol 5. National Biomedical Research Foundation, Washington, DC, pp 345–358
Delgado S, Casane D, Bonnaud L, Laurin M, Sire JY, Girondot M (2001) Molecular evidence for Precambrian origin of amelogenin, the major protein of vertebrate enamel. Mol Biol Evol 18:2146–2153
Delgado S, Girondot M, Sire JY (2005) Molecular evolution of amelogenin in mammals. J Mol Evol 60:12–30
Delgado S, Couble ML, Magloire H, Sire JY (2006) Cloning, sequencing, and expression of the amelogenin gene in two scincid lizards. J Dent Res 85:138–143
Delgado S, Ishiyama M, Sire JY (2007) Validation of amelogenesis imperfecta inferred from amelogenin evolution. J Dent Res 86:326–330
Deméré TA, McGowen MR, Berta A, Gatesy J (2008) Morphological and molecular evidence for a stepwise evolutionary transition from teeth to baleen in mysticete whales. Syst Biol 57:15–37
Deutsch D (1989) Structure and function of enamel gene products. Anat Rec 224:189–210
Dong J, Gu TT, Simmons D, MacDougall M (2000) Enamelin maps to human chromosome 4q21 within the autosomal dominant amelogenesis imperfecta locus. Eur J Oral Sci 108:353–358
Doron-Faigenboim A, Stern A, Mayrose I, Bacharach E, Pupko T (2005) Selecton: a server for detecting evolutionary forces at a single amino-acid site. Bioinformatics 21:2101–2103
Dunker AK, Brown CJ, Lawson JD, Iakoucheva LM, Obradovic Z (2002) Intrinsic disorder and protein function. Biochemistry 41:6573–6582
Endo T, Ikeo K, Gojobori T (1996) Large-scale search for genes on which positive selection may operate. Mol Biol Evol 13:685–690
Fan D, Lakshminarayanan R, Moradian-Oldak J (2008) The 32 kDa enamelin undergoes conformational transitions upon calcium binding. J Struct Biol 163:109–115
Fisher LW, Fedarko NS (2003) Six genes expressed in bones and teeth encode the current members of the SIBLING family of proteins. Connect Tissue Res 44(Suppl 1):33–40
Fukae M, Tanabe T (1985) Separation of non-amelogenin component from purified amelogenin preparation of immature porcine enamel. Jpn J Oral Biol 27:1249–1251
Fukae M, Tanabe T (1987) Nonamelogenin components of porcine enamel in the protein fraction free from the enamel crystals. Calcif Tissue Int 40:286–293
Fukae M, Tanabe T, Murakami C, Dohi N, Uchida T, Shimizu M (1996) Primary structure of the porcine 89-kDa enamelin. Adv Dent Res 10:111–118
Gomis-Ruth FX, Bayes A, Sotiropoulou G, Pampalakis G, Tsetsenis T, Villegas V, Aviles FX, Coll M (2002) The structure of human prokallikrein 6 reveals a novel activation mechanism for the kallikrein family. J Biol Chem 277:27273–27281
Gutierrez SJ, Chaves M, Torres DM, Briceno I (2007) Identification of a novel mutation in the enamelin gene in a family with autosomal-dominant amelogenesis imperfecta. Arch Oral Biol 52:503–506
Hart PS, Michalec MD, Seow WK, Hart TC, Wright JT (2003a) Identification of the enamelin (g.8344delG) mutation in a new kindred and presentation of a standardized ENAM nomenclature. Arch Oral Biol 48:589–596
Hart TC, Hart PS, Gorry MC, Michalec MD, Ryu OH, Uygur C, Ozdemir D, Firatli S, Aren G, Firatli E (2003b) Novel ENAM mutation responsible for autosomal recessive amelogenesis imperfecta and localised enamel defects. J Med Genet 40:900–906
Hasegawa M, Kishino H, Yano T (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Higgins DG, Thompson JD, Gibson TJ (1996) Using CLUSTAL for multiple sequence alignments. Methods Enzymol 266:383–402
Hiss JA, Resch E, Schreiner A, Meissner M, Starzinski-Powitz A, Schneider G (2008) Domain organization of long signal peptides of single-pass integral membrane proteins reveals multiple functional capacity. PLoS ONE 3:e2767
Hu JC, Yamakoshi Y (2003) Enamelin and autosomal-dominant amelogenesis imperfecta. Crit Rev Oral Biol Med 14:387–398
Hu C-C, Fukae M, Uchida T, Qian Q, Zhang CH, Ryu OH, Tanabe T, Yamakoshi Y, Murakami C, Dohi N, Shimizu M, Simmer JP (1997a) Cloning and characterization of porcine enamelin mRNAs. J Dent Res 76:1720–1729
Hu CC, Fukae M, Uchida T, Qian Q, Zhang CH, Ryu OH, Tanabe T, Yamakoshi Y, Murakami C, Dohi N, Shimizu M, Simmer JP (1997b) Sheathlin: cloning, cDNA/polypeptide sequences, and immunolocalization of porcine enamel sheath proteins. J Dent Res 76:648–657
Hu CC, Simmer JP, Bartlett JD, Qian Q, Zhang C, Ryu OH, Xue J, Fukae M, Uchida T, MacDougall M (1998) Murine enamelin: cDNA and derived protein sequences. Connect Tissue Res 39:47–61
Hu CC, Hart TC, Dupont BR, Chen JJ, Sun X, Qian Q, Zhang CH, Jiang H, Mattern VL, Wright JT, Simmer JP (2000) Cloning human enamelin cDNA, chromosomal localization, and analysis of expression during tooth development. J Dent Res 79:912–919
Hu JC, Zhang CH, Yang Y, Karrman-Mardh C, Forsman-Semb K, Simmer JP (2001) Cloning and characterization of the mouse and human enamelin genes. J Dent Res 80:898–902
Hu JC, Yamakoshi Y, Yamakoshi F, Krebsbach PH, Simmer JP (2005) Proteomics and genetics of dental enamel. Cells Tissues Organs 181:219–231
Hu JC, Hu Y, Smith CE, McKee MD, Wright JT, Yamakoshi Y, Papagerakis P, Hunter GK, Feng JQ, Yamakoshi F, Simmer JP (2008) Enamel defects and ameloblast-specific expression in Enam knock-out/lacz knock-in mice. J Biol Chem 283:10858–10871
Kang HY, Seymen F, Lee SK, Yildirim M, Tuna EB, Patir A, Lee KE, Kim JW (2009) Candidate gene strategy reveals ENAM mutations. J Dent Res 88:266–269
Kawasaki K, Weiss KM (2003) Mineralized tissue and vertebrate evolution: the secretory calcium-binding phosphoprotein gene cluster. Proc Natl Acad Sci USA 100:4060–4065
Kawasaki K, Weiss KM (2008) SCPP gene evolution and the dental mineralization continuum. J Dent Res 87:520–531
Kelley JL, Swanson WJ (2008) Dietary change and adaptive evolution of enamelin in humans and among Primates. Genetics 178:1595–1603
Kim JW, Seymen F, Lin BP, Kiziltan B, Gencay K, Simmer JP, Hu JC (2005a) ENAM mutations in autosomal-dominant amelogenesis imperfecta. J Dent Res 84:278–282
Kim JW, Hu JCC, Lee JI, Moon SK, Kim YJ, Jang KT, Lee SH, Kim CC, Hahn SH, Simmer JP (2005b) Mutational hot spot in the DSPP gene causing dentinogenesis imperfecta type II. Hum Genet 116:186–191
Pond SLK, Frost SD (2005a) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Pond SLK, Frost SD (2005b) Datamonkey: rapid detection of selective pressure on individual sites of codon alignments. Bioinformatics 21:2531–2533
Pond SLK, Frost SD, Muse SV (2005) HyPhy: hypothesis testing using phylogenies. Bioinformatics 21:676–679
Kozak M (1984) Compilation and analysis of sequences upstream from the translational start site in eukaryotic mRNAs. Nucleic Acids Res 12:857–872
Kozak M (1991) A short leader sequence impairs the fidelity of initiation by eukaryotic ribosomes. Gene Expr 1:111–115
Kurys G, Tagaya Y, Bamford R, Hanover JA, Waldmann TA (2000) The long signal peptide isoform and its alternative processing direct the intracellular trafficking of interleukin-15. J Biol Chem 275:30653–30659
Lu Y, Papagerakis P, Yamakoshi Y, Hu JC, Bartlett JD, Simmer JP (2008) Functions of KLK4 and MMP-20 in dental enamel formation. Biol Chem 389:695–700
Mårdh CK, Bäckman B, Holmgren G, Hu JC, Simmer JP, Forsman-Semb K (2002) A nonsense mutation in the enamelin gene causes local hypoplastic autosomal dominant amelogenesis imperfecta (AIH2). Hum Mol Genet 11(9):1069–1074
Martoglio B, Dobberstein B (1998) Signal sequences: more than just greasy peptides. Trends Cell Biol 8:410–415
Masuya H, Shimizu K, Sezutsu H, Sakuraba Y, Nagano J, Shimizu A, Fujimoto N, Kawai A, Miura I, Kaneda H, Kobayashi K, Ishijima J, Maeda T, Gondo Y, Noda T, Wakana S, Shiroishi T (2005) Enamelin (Enam) is essential for amelogenesis: ENU-induced mouse mutants as models for different clinical subtypes of human amelogenesis imperfecta (AI). Hum Mol Genet 14:575–583
Ozdemir D, Hart PS, Firatli E, Aren G, Ryu OH, Hart TC (2005) Phenotype of ENAM mutations is dosage-dependent. J Dent Res 84:1036–1041
Patthy L (1999) Genome evolution and the evolution of exon-shuffling—a review. Gene 238:103–114
Rajpar MH, Harley K, Laing C, Davies RM, Dixon MJ (2001) Mutation of the gene encoding the enamel-specific protein, enamelin, causes autosomal-dominant amelogenesis imperfecta. Hum Mol Genet 10:1673–1677
Rambaut A, Bromham L (1998) Estimating divergence dates from molecular sequences. Mol Biol Evol 15:442–448
Ravindranath RM, Moradian-Oldak J, Fincham AG (1999) Tyrosyl motif in amelogenins binds N-acetyl-D-glucosamine. J Biol Chem 274:2464–2471
Ravindranath HH, Chen LS, Zeichner-David M, Ishima R, Ravindranath RM (2004) Interaction between the enamel matrix proteins amelogenin and ameloblastin. Biochem Biophys Res Commun 323:1075–1083
Reuter M, Engelstadter J, Fontanillas P, Hurst LD (2008) A test of the null model for 5′ UTR evolution based on GC content. Mol Biol Evol 25:801–804
Ryu OH, Fincham AG, Hu CC, Zhang C, Qian Q, Bartlett JD, Simmer JP (1999) Characterization of recombinant pig enamelysin activity and cleavage of recombinant pig and mouse amelogenins. J Dent Res 78:743–750
Sansom IJ, Smith MP, Armstrong HA, Smith MM (1992) Presence of the earliest vertebrate hard tissue in conodonts. Science 256:1308–1311
Schmid K, Yang Z (2008) The trouble with sliding windows and the selective pressure in BRCA1. PLoS ONE 3(11):e3746
Seedorf H, Klaften M, Eke F, Fuchs H, Seedorf U, Hrabe de Angelis M (2007) A mutation in the enamelin gene in a mouse model. J Dent Res 86:764–768
Sire JY, Delgado S, Fromentin D, Girondot M (2005) Amelogenin: lessons from evolution. Arch Oral Biol 50:205–212
Sire JY, Delgado S, Girondot M (2006) The amelogenin story: origin and evolution. Eur J Oral Sci 114(Suppl 1):64–77
Sire JY, Davit-Béal T, Delgado S, Gu X (2007) The origin and evolution of enamel mineralization genes. Cells Tissues Organs 186:25–48
Sire JY, Delgado S, Girondot M (2008) Hen’s teeth with enamel cap: from dream to impossibility. BMC Evol Biol 8:e246
Springer MS, Murphy WJ (2007) Mammalian evolution and biomedicine: new views from phylogeny. Biol Rev Camb Philos Soc 82:375–392
Stern A, Doron-Faigenboim A, Erez E, Martz E, Bacharach E, Pupko T (2007) Selecton 2007: advanced models for detecting positive and purifying selection using a Bayesian inference approach. Nucleic Acids Res 35:W506–W511
Subramanian S, Kumar S (2006) Evolutionary anatomies of positions and types of disease-associated and neutral amino acid mutations in the human genome. BMC Genomics 7:e306
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Tanabe T, Aoba T, Moreno EC, Fukae M, Shimuzu M (1990) Properties of phosphorylated 32 kd nonamelogenin proteins isolated from porcine secretory enamel. Calcif Tissue Int 46:205–215
Tanabe T, Fukae M, Shimizu M (1994) Degradation of enamelins by proteinases found in porcine secretory enamel in vitro. Arch Oral Biol 39:277–281
Termine JD, Belcourt AB, Christner PJ, Conn KM, Nylen MU (1980) Properties of dissociatively extracted fetal tooth matrix proteins. I. Principal molecular species in developing bovine enamel. J Biol Chem 255:9760–9768
Tsunoyama K, Gojobori T (1998) Evolution of nicotinic acetylcholine receptor subunits. Mol Biol Evol 15:518–527
Uchida T, Tanabe T, Fukae M, Shimizu M, Yamada M, Miake K, Kobayashi S (1991a) Immunochemical and immunohistochemical studies, using antisera against porcine 25 kDa amelogenin, 89 kDa enamelin and the 13–17 kDa nonamelogenins, on immature enamel of the pig and rat. Histochemistry 96:129–138
Uchida T, Tanabe T, Fukae M, Shimizu M (1991b) Immunocytochemical and immunochemical detection of a 32 kDa nonamelogenin and related proteins in porcine tooth germs. Arch Histol Cytol 54:527–538
van Rheede T, Bastiaans T, Boone DN, Hedges SB, de Jong WW, Madsen O (2006) The platypus is in its place: nuclear genes and indels confirm the sister group relation of monotremes and therians. Mol Biol Evol 23:587–597
von Heijne G (1985) Signal sequences: the limits of variation. J Mol Biol 184:99–105
Warren WC, Hillier LW, Marshall Graves JA, Birney E, Ponting CP, Grutzner F, Belov K, Miller W, Clarke L, Chinwalla AT, Yang SP, Heger A, Locke DP, Miethke P, Waters PD, Veyrunes F, Fulton L, Fulton B, Graves T, Wallis J, Puente XS, Lopez-Otin C, Ordonez GR, Eichler EE, Chen L, Cheng Z, Deakin JE, Alsop A, Thompson K, Kirby P, Papenfuss AT, Wakefield MJ, Olender T, Lancet D, Huttley GA, Smit AF, Pask A, Temple-Smith P, Batzer MA, Walker JA, Konkel MK, Harris RS, Whittington CM, Wong ES, Gemmell NJ, Buschiazzo E, Vargas Jentzsch IM, Merkel A, Schmitz J, Zemann A, Churakov G, Kriegs JO, Brosius J, Murchison EP, Sachidanandam R, Smith C, Hannon GJ, Tsend-Ayush E, McMillan D, Attenborough R, Rens W, Ferguson-Smith M, Lefevre CM, Sharp JA, Nicholas KR, Ray DA, Kube M, Reinhardt R, Pringle TH, Taylor J, Jones RC, Nixon B, Dacheux JL, Niwa H, Sekita Y, Huang X, Stark A, Kheradpour P, Kellis M, Flicek P, Chen Y, Webber C, Hardison R, Nelson J, Hallsworth-Pepin K, Delehaunty K, Markovic C, Minx P, Feng Y, Kremitzki C, Mitreva M, Glasscock J, Wylie T, Wohldmann P, Thiru P, Nhan MN, Pohl CS, Smith SM, Hou S, Renfree MB (2008) Genome analysis of the platypus reveals unique signatures of evolution. Nature 453:175–183
Weiner S (1986) Organization of extracellularly mineralized tissues: a comparative study of biological crystal growth. CRC Crit Rev Biochem 20:365–408
Yamakoshi Y (1995) Carbohydrate moieties of porcine 32 kDa enamelin. Calcif Tissue Int 56:323–330
Yamakoshi Y, Pinheiro FH, Tanabe T, Fukae M, Shimizu M (1998) Sites of asparagine-linked oligosaccharides in porcine 32 kDa enamelin. Connect Tissue Res 39:39–46
Yamakoshi Y, Hu JC, Liu S, Zhang C, Oida S, Fukae M, Simmer JP (2003) Characterization of porcine dentin sialoprotein (DSP) and dentin sialophosphoprotein (DSPP) cDNA clones. Eur J Oral Sci 111:60–67
Yamakoshi Y, Hu JC, Fukae M, Yamakoshi F, Simmer JP (2006) How do enamelysin and kallikrein 4 process the 32-kDa enamelin? Eur J Oral Sci 114(Suppl 1):45–51
Yoon H, Laxmikanthan G, Lee J, Blaber SI, Rodriguez A, Kogot JM, Scarisbrick IA, Blaber M (2007) Activation profiles anad regulatory cascades of the human kallikrein-related peptidases. J Biol Chem 282:31852–31864
Acknowledgments
We thank Mehboob Chilwan (Erasmus, University of Keele, UK) for English corrections. This work was supported by CNRS and UPMC (UMR 7138) Grants.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
239_2009_9302_MOESM1_ESM.doc
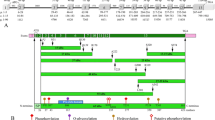
Amino acid alignment of the 36 mammalian ENAMs analyzed. The sequences were aligned against the human sequence and are ordered following the mammalian relationships shown in Fig. 3. The 24 complete sequences are indicated in bold characters. This alignment leads to 1,550 positions (including all gaps). The signal peptide (exons 3 and 4) is underlined. Remarkably conserved residues and motifs are on gray background.][: exon limits; (.): residue identical to human ENAM residue; (−): indel; (=): unknown amino acid; (#): the 77 residues identical in all sequences; (+): the 10 site-specific positive selection in human ENAM (see also Figs. 5 and 7).(DOC 120 kb)
239_2009_9302_MOESM2_ESM.doc
Comparison of the 20 nucleotides located at the 5′ and 3′ UTR, and at both splice sites of introns 3–9 in ten ENAM sequences of species representative of the main mammalian lineages. Conserved positions are indicated in bold characters. Three mutations at the splicing sites (boxed bp) are known to lead to AIH2 (see Fig. 1).(DOC 78 kb)
Rights and permissions
About this article
Cite this article
Al-Hashimi, N., Sire, JY. & Delgado, S. Evolutionary Analysis of Mammalian Enamelin, The Largest Enamel Protein, Supports a Crucial Role for the 32-kDa Peptide and Reveals Selective Adaptation in Rodents and Primates. J Mol Evol 69, 635–656 (2009). https://doi.org/10.1007/s00239-009-9302-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9302-x