Abstract
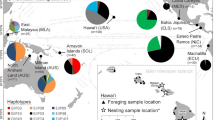
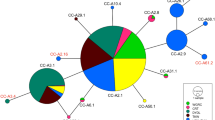
Mitochondrial (mt) DNA control region sequences were analyzed for 249 Atlantic and Mediterranean loggerhead turtles (Carettacaretta Linnaeus, 1758) to elucidate nesting population structure and phylogeographic patterns. Ten haplotypes were resolved among individuals sampled between 1987 and 1993, from ten major loggerhead nesting areas in the region. Two distinct phylogenetic lineages were distinguished, separated by an average of 5.1% sequence divergence. Haplotype frequency comparisons between pairs of populations showed significant differentiation between most regional nesting aggregates and revealed six demographically independent groups, corresponding to nesting beaches from: (1) North Carolina, South Carolina, Georgia and northeast Florida, USA; (2) southern Florida, USA; (3) northwest Florida, USA; (4) Quintana Roo, Mexico; (5) Bahia, Brazil; and (6) Peloponnesus Island, Greece. The distribution of mtDNA haplotypes is consistent with a natal homing scenario, in which nesting colonies separated by a few hundred kilometers represent isolated reproductive aggregates. However, a strong exception to this pattern was observed in the first group defined by mtDNA data (North Carolina to northeast Florida), which included samples from four nesting locations spread across thousands of kilometers of coastline. These locations were characterized by a single haplotype in 104 out of 105 samples, providing inadequate resolution of population divisions. In view of the subdivisions observed elsewhere, we attribute the lack of differentiation between North Carolina and northeast Florida to recent colonization of these warm temperate coastlines (after the Wisconsin glaciation) not to ongoing gene flow among spatially distinct nesting locations. The relationships among observed haplotypes suggest a biogeographic scenario defined by climate, natal homing, and rare dispersal events. The redefined relationships among nesting aggregations in the western Atlantic region (southeastern USA and adjacent Mexico) prompt a reconsideration of management strategies for nesting populations and corresponding habitats in this region.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 28 October 1996 / Accepted: 24 October 1997
Rights and permissions
About this article
Cite this article
Encalada, S., Bjorndal, K., Bolten, A. et al. Population structure of loggerhead turtle (Carettacaretta) nesting colonies in the Atlantic and Mediterranean as inferred from mitochondrial DNA control region sequences. Marine Biology 130, 567–575 (1998). https://doi.org/10.1007/s002270050278
Issue Date:
DOI: https://doi.org/10.1007/s002270050278