Abstract
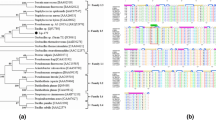
A novel lipolytic enzyme-producing endophytic strain PC2 was successfully isolated from the seeds of an ideal bioenergy plant Pistacia chinensis Bunge. Based on the analysis of morphology and 16S rRNA sequence, bacterial strain PC2 was identified as a subspecies of Pseudomonas putida, therefore named as P. putida PC2. Whole-genome sequencing showed PC2 contained a 1224-nucleotide lipase gene (named lip-PC2) predicted to encode a 407-amino-acid protein. Purified lipases from both the original PC2 strain and heterologously expressed Escherichia coli were nearly 50 kD with specific activity of 9.48 U/mL. LIP-PC2 displayed the maximal activity at 50°C or pH 8.0, and maintained above 80% relative activity in the range of from 40 to 60°C or pH in the range of from 6.0 to 8.0, indicating thermostable and alkaline properties. Enzyme activity was enhanced by Mg2+, Na+ and Mn2+, but strongly inhibited by Cu2+, Zn2+ Co2+, EDTA as well as organic solvents and surfactants. Additionally, the analysis of amino acid sequence and structure indicated that LIP-PC2 was a novel member belonging to family I.3 of bacterial lipolytic enzymes and its catalytic triad was consisted of Ser-200, Asp-342 and His-374.
Similar content being viewed by others
References
Li, H.L., Zhang, Z.X., Lin, S.Z., and Li, X.X., Forestry Studies in China, 2007, vol. 9, no. 2, pp. 164–168.
Xiong, E., Wu, X., Shi, J., Wang, X., and Wang, W., PLoS One, 2013, vol. 8, no. 5, p. e64276.
Piromyou, P., Songwattana, P., Greetatorn, T., Okubo, T., Kakizaki, K.C., Prakamhang, J., et al., Microbes Environ., 2015, vol. 30, no. 4, pp. 291–300.
Carrell, A.A. and Frank, A.C., Front. Microbiol., 2014, vol. 5, no. 333, p. 10.3389.
Sahai, A.S. and Manocha, M.S., FEMS Microbiol. Rev., 1993, vol. 11, no. 4, pp. 317–338.
Zhang, H.W., Song, Y.C., and Tan, R.X., Nat. Prod. Rep., 2006, vol. 23, no. 5, pp. 753–771.
Song, C.Z. and Chen, D.H., Res. J. Biotech., 2015, vol. 10, no. 8, pp. 105–112.
Olson, G.J., Woese, C.R., and Overbeek, R., J. Bacteriol., 1994, vol. 176, pp. 1–6.
Jaeger, K.E., Ransac, S., Dijkstra, B.W., Colson, C., van Heuve., M., and Misset, O., FEMS Microbiol. Rev., 1994, vol. 15, no. 1, pp. 29–63.
Jaeger, K.E. and Eggert, T., Curr. Opin. Biotech., 2002, vol. 13, no. 4, pp. 390–397.
Ollis, D.L., Cheah, E., Cygler, M., Dijkstra, B.W., Frolow, F., Franken, S.M., et al., Protein Eng., 1992, vol. 5, pp. 197–211.
Arpigny, J.L. and Jaeger, K.E., Biochem. J., 1999, vol. 343, no. 1, pp. 177–183.
Tanaka, D., Yoneda, S., Yamashiro, Y., Sakatoku, A., Kayashima, T., Yamakawa, K., et al., Appl. Biochem. Biotech., 2012, vol. 168, no. 2, pp. 327–338.
Charbonneau, D.M. and Beauregard, M., PLoS One, 2013, vol. 8, no. 10, p. e76675.
Kim, H.J., Jeong, Y.S., Jung, W.K., Kim, S.K., Lee, H.W., Kahng, H.Y., et al., Mol. Biotech., 2015, vol. 57, no. 9, pp. 781–792.
Winkler, U.K. and Stuckmann, M., J. Bacteriol., 1979, vol. 138, no. 3, pp. 663–670.
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., Higgins, D.G., and Nucleic Acids Res., 1997, vol. 25, pp. 4876–4882.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., and Kumar, S., Mol. Biol. Evol., 2013, vol. 30, pp. 2725–2729.
Petersen, T.N., Brunak, S., von Heijn., G., and Nielsen, H., Nat. Methods, 2011, vol. 8, no. 10, pp. 785–786.
Bjellqvist, B., Basse, B., Olsen, E., and Celis, J.E., Electrophoresis, 1994, vol. 15, no. 1, pp. 529–539.
Bjellqvist, B., Hughes, G.J., Pasquali, C., Paquet, N., Ravier, F., Sanchez, J.C., et al., Electrophoresis, 1993, vol. 14, no. 1, 1023–1033.
Gasteiger, E., Hoogland, C., Gattiker, A., Duvaud, S.E., Wilkins, M.R., Appel, R.D., et al., Humana Press, 2005.
Robert, X. and Gouet, P., Nucleic Acids Res., 2, vol. 42, no. W1, pp. W320–W324.
Ericsson, D.J., Kasrayan, A., Johansson, P., Bergfors, T., Sandstrom, A.G., Bäckvall, J.E., et al., J. Mol. Biol., 2008, vol. 376, no. 1, pp. 109–119.
Angkawidjaja, C., Kanaya, S., Cell Mol. Life Sci., 2006, vol. 63, pp. 2804–2817.
Rashid, N., Shimada, Y., Ezaki, S., Atomi, H., and Imanaka, T., Appl. Environ. Microbiol., 2001, vol. 67, no. 9, pp. 4064–4069.
Yamashiro, Y., Sakatoku, A., Tanaka, D., and Nakamura, S., Appl. Biochem. Biotechnol., 2013, vol. 171, pp. 989–1000.
Amada, K., Haruki, M., Imanaka, T., Morikawa, M., and Kanaya, S., Biochem. Biophys. Acta—Protein Struct. M., 2000, vol. 1478, no. 2, pp. 201–210.
Luo, Y., Zheng, Y., Jiang, Z., Ma, Y., and Wei, D., Appl. Microbiol. Biotech., 2006, vol. 73, no. 2, pp. 349–355.
Madan, B. and Mishra, P., Appl. Microbiol. Biotech., 2010, vol. 85, no. 3, pp. 597–604.
Panizza, P., Syfantou, N., Pastor, F.I.J., Rodriguez, S., and Diaz, P., J. Appl. Microbiol., 2013, vol. 114, no. 3, pp. 722–732.
Angkawidjaja, C., You, D.J., Matsumura, H., Kuwahara, K., Koga, Y., Takano, K., et al., FEBS Lett., 2007, vol. 581, no. 26, pp. 5060–5064.
Kim, J., Jang, S.H., and Lee, C.W., Biosci. Biotech. Biochem., 2013, vol. 77, no. 2, pp. 320–323.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
Rights and permissions
About this article
Cite this article
Song, C., Liu, Z., Xie, Q. et al. Characterization of a novel thermo-stable lipase from endophyte Pseudomonas putida in Pistacia chinensis Bunge. Appl Biochem Microbiol 53, 524–532 (2017). https://doi.org/10.1134/S0003683817050143
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683817050143