Abstract
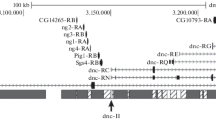
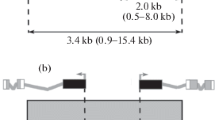
Using electron microscopic (EM) data on the formation of a novel band from theP-element material after its insertion in the interband and the procedure of P-target rescue, DNA interband regions 3A5/A6, 3C5-6/C7, and 60E8-9/E10 of Drosophila melanogasterpolytene chromosomes were cloned and sequenced. EM analysis of the 3C region have shown that the formation of the full-size 3C5-6/C7 interband requires a 880-bp DNA sequence removed by deletion Df(1)fa swb. A comparison of DNA sequences of six bands, two of which were obtained in the present work and four were described earlier, demonstrated the uniqueness of each of them in the Drosophilagenome and heterogeneity of their molecular organization. Interband 60E8-9/E10 contains gene rpl19transcribed throughout the development, in particular in salivary glands. In the other interbands examined 5" and 3" nontranslated gene regions are located. These results suggest that Drosophilainterbands may contain both housekeeping genes and regulatory sequences of currently inactive genes from adjacent bands.
Similar content being viewed by others
REFERENCES
Beermann, W., Chromomeres and Genes, Results and Problems in Cell Differentiation, Beermann, W., Ed., Berlin: Springer-Verlag, 1972, vol. 4, pp. 1-33.
Zhimulev, I.F., Belyaeva, E.S., and Semeshin, V.F., Informational Content of Polytene Chromosome Bands and Puffs, CRC-Crit. Rev. Biochem., 1981, vol. 11, no. 4, pp. 303-340.
Zhimulev, I.F., Khromomernaya organizatsiya politennykh khromosom (The Chromomere Organization of Polytene Chromosomes), Novosibirsk: Nauka, 1994.
Crick, F., General Model for the Chromosomes of Higher Organisms, Nature, 1971, vol. 234, pp. 25-27.
Paul, J., General Theory of Chromosomes Structure and Gene Activation in Eukaryotes, Nature, 1972, vol. 238, pp. 444-446.
Sorsa, V., Electron Microscopic Mapping and Ultrastructure of Drosophila Polytene Chromosomes, Insect Ultrastructure, King, R.C. and Akai, H., Eds., New York: Plenum, 1984, vol. 2, pp. 75-107.
Semeshin, V.F., Belyaeva, E.S., Zhimulev, I.F., et al., Electron Microscopic Analysis of Drosophila Polytene Chromosomes: IV. Mapping of Morphological Structures Appearing as a Result of Transformation of DNA Sequences into Chromosomes, Chromosoma, 1986, vol. 93, pp. 461-468.
Semeshin, V.F., Demakov, S.A., Perez Alonso, M., et al., Electron Microscopic Analysis of Drosophila Polytene Chromosomes: V. Characteristics of Structures Formed by Transposed DNA Segments of Mobile Elements, Chromosoma, 1989, vol. 97, pp. 396-412.
Demakov, S.A., Semeshin, V.F., and Zhimulev, I.F., Cloning and Molecular Genetic Analysis of Drosophila melanogaster Interband DNA, Mol. Gen. Genet., 1993, vol. 238, pp. 437-443.
Shvarts, Yu.B., Demakov, S.A., and Zhimulev, I.F., Cloning and Analysis of DNA from 85D9/D10 and 86B4/B6 Interband Regions of Drosophila melanogaster Polytene Chromosomes, Genetika (Moscow), 1998, vol. 34, no. 8, pp. 1081-1089.
Schwartz, Yu.B., Demakov, S.A., and Zhimulev, I.F., Polytene Chromosome Interband DNA Is Organized into Nucleosomes, Mol. Gen. Genet., 2001, vol. 265, pp. 311-315.
Schwartz, Yu.B., Ioudinkova, E.S., Demakov, S.A., et al., Interbands of Drosophila melanogaster Polytene Chromosomes Contain Matrix Association Regions, J. Cell Biochem., 1999, vol. 72, pp. 368-372.
Ramos, R.G.P., Grimwade, B.G., Wharton, K.A., et al., Physical and Functional Definition of the Drosophila Notch Locus by P-Element Transformation, Genetics, 1989, vol. 123, pp. 337-348.
Maniatis, T., Fritsch, E.F., and Sambrook, J., Molecular Cloning: A Laboratory Manual, Cold Spring Harbor, New York: Cold Spring Harbor Lab., 1982.
Altschul, S.F., Gish, W., Miller, W., et al., Basic Local Alignment Search Tool, J. Mol. Biol., 1990, vol. 215, pp. 403-410.
FlyBase, The FlyBase Database of the Drosophila Genome Projects and Community Literature, Nucleic Acids Res., 1999, vol. 27, pp. 85-88; http://flybase.bio.indiana.edu/.
Singh, G.A., Kramer, J.A., and Krawetz, S.A., Mathematical Model to Predict Regions of Chromatin Attachment to the Nuclear Matrix, Nucleic Acids Res., 1997, vol. 25, no. 7, pp. 1419-1425; http://www.futuresoft.org/MAR–Wiz.
Frisch, M., http://genomatix.gsf.de/cgi-bin/smartest_pd/ smartest.pl.
Morgenstern, B., Frech, K., Dress, A., and Werner, T., DIALIGN: Finding Local Similarities by Multiple Sequence Alignment, Bioinformatics, 1998, vol. 14, pp. 290-294; http://genomatix.gsf.de/cgi-bin/dialgn/dialign/pl.
Quandt, K., Frech, K., Karas, H., et al., MatInd and MatInspector: New Fast and Versatile Tools for Detection of Consensus Matches in Nucleotide Sequence Data, Nucleic Acids Res., 1995, vol. 23, pp. 4878-4884; http://genomatix. gsf.de/cgi-bin/matinspector/matinspector.pl/.
Semeshin, V.F., Artero, R., Perez-Alonso, M., and Shloma, V.V., Electron Microscopic In Situ Hybridization of Digoxigenin-dUTP-Labeled DNA Probes with Drosophila melanogaster Polytene Chromosomes, Chromosome Res., 1998, vol. 6, pp. 405-410.
Hart, K., Klein, T., and Wilcox, M.A., Minute Encoding a Ribosomal Protein Enhances Wing Morphogenesis Mutants, Mech. Dev., 1993, vol. 43, nos. 2-3, pp. 101-110.
Korge, G., Heide, I., Sehnert, M., and Hofmann, A., Promoter Is an Important Determinant of Developmentally Regulated Puffing at the Sgs-4 Locus of Drosophila melanogaster, Dev. Biol., 1980, vol. 138, pp. 324-337.
O'Kane, C.J. and Gehring, W.J., Detection In Situ of Genomic Regulatory Elements in Drosophila, Proc. Natl. Acad. Sci. USA, 1987, vol. 84, pp. 9123-9127.
Adams, M.D., Celniker, S.E., Holt, R.A., et al., The Genome Sequence of Drosophila melanogaster, Science, 2000, vol. 287, no. 5461, pp. 2185-2195.
Rubin, G.M., Hong, L., Brokstein, P., et al., A Drosophila Complementary DNA Resource, Science, 2000, vol. 287, no. 5461, pp. 2222-2224.
Goode, S., Melnick, M., Chou, T.B., and Perrimon, N., The Neurogenic Genes egghead and brainiac Define a Novel Signaling Pathway Essential for Epithelial Morphogenesis during Drosophila Oogenesis, Development (Cambridge, UK), 1996, vol. 122, no. 12, pp. 3863-3879.
Hollmann, M., Drosophila melanogaster WDS (wds) and egghead (egh) Genes, Complete cds, GenBank/ EMBL/DDBJ 2000.2.9. AF233288.
Keppy, D.O. and Welshons, W.J., The Cytogenetics of a Recessive Visible Mutant Associated with a Deficiency Adjacent to the Notch Locus in Drosophila melanogaster, Genetics, 1977, vol. 85, no. 3, pp. 497-506.
Rykowski, M.C., Parmelee, S.J., Agard, D.A., and Sedat, J.W., Precise Determination of the Molecular Limits of a Polytene Chromosome Band: Regulatory Sequences for the Notch Gene Are in the Interband, Cell (Cambridge, Mass.), 1988, vol. 54, pp. 461-472.
Kidd, S., Kelley, M.R., and Young, M.W., Sequence of the Notch Locus of Drosophila melanogaster: Relationship of the Encoded Protein to Mammalian Clotting and Growth Factors, Mol. Cell. Biol., 1986, vol. 6, pp. 3094-3108.
Vasquez, J. and Schedl, P., Deletion of an Insulator Element by the Mutation facet-strawberry in Drosophila melanogaster, Genetics, 2000, vol. 155, pp. 1297-1311.
Kelley, M.R., Kidd, S., Berg, R.L., and Young, M.W., Restriction of P-Element Insertions at the Notch Locus of Drosophila melanogaster, Mol. Cell. Biol., 1987, vol. 7, pp. 1545-1548.
Whitmore, T., Localization of a “Housekeeping” Gene in Chironomus thummi thummi, Cytobios, 1986, vol. 46, pp. 193-200.
Zhimulev, I.F. and Belyaeva, E.S., Detailed Description of Puffing Patterns in the Salivary Gland Chromosomes of Normally Developing Larvae and Prepupae of Drosophila melanogaster, Dros. Inf. Serv., 1999, vol. 82, pp. 9-20.
Aalfs, J.D. and Kingston, R.E., What Does “Chromatin Remodeling” Mean?, Trends Biochem. Sci., 2000, vol. 25, pp. 548-555.
Mirkovitch, J., Gasser, S.M., and Laemmli, U.K., Genes and Loops in 320,000 Base Pairs of the Drosophila melanogaster Chromosome, J. Mol. Biol., 1986, vol. 190, pp. 255-258.
Iarovaia, O., Hancock, R., Lagarkova, M., et al., Mapping of Genomic DNA Loop Organization in a 500-Kilobase Region of the Drosophila X Chromosome by the Topoisomerase II-Mediated DNA Loop Excision Protocol, Mol. Cell. Biol., 1996, vol. 16, pp. 302-308.
Zhao, K., Hart, C.M., and Laemmli, U.K., Visualization of Chromosomal Domains with Boundary Element-Associated Factor BEAF-32, Cell (Cambridge, Mass.), 1995, vol. 81, pp. 879-889.
Cai, H.N. and Levine, M., The gypsy Insulator Can Function as a Promoter-Specific Silencer in the Drosophila Embryo, EMBO. J., 1997, vol. 16, no. 7, pp. 1732-1741.
Scott, K.C., Taubman, A.D., and Geyer, P.K., Enhancer Blocking by the Drosophila gypsy Insulator Depends upon Insulator Anatomy and Enhancer Strength, Genetics, 1999, vol. 153, pp. 787-798.
Parnell, T.J. and Geyer, P.K., Differences in Insulator Properties Revealed by Enhancer Blocking Assays on Episomes, EMBO J., 2000, vol. 19, no. 21, pp. 5864-5874.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Demakov, S.A., Gorchakov, A.A., Schwartz, Y.B. et al. Analysis of DNA Interband Regions 3A5/A6, 3C5-6/C7, and 60E8-9/E10 in Polytene Chromosomes of Drosophila melanogaster. Russian Journal of Genetics 37, 1247–1256 (2001). https://doi.org/10.1023/A:1012548807155
Issue Date:
DOI: https://doi.org/10.1023/A:1012548807155