Abstract
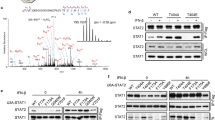
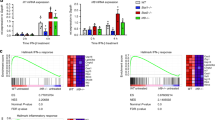
The Interferon Regulatory Factors (IRFS) play an important role in the transcriptional control of growth regulatory and immunoregulatory genes. The inducibility and availability of IRF-1 and IRF-2 are influenced by external stimuli, such as virus infection or interferon treatment. In the present study, we sought to examine the potential modulatory role of phosphorylation on IRF-1 transcriptional activity. During the purification of IRF recombinant proteins, a kinase activity copurified with IRF- 1 (and IRF-2) from baculovirus infected Sf9 insect cell extracts, but not from E. coli extracts. The kinase activity was also identified in Jurkat T cells, specifically interacted with IRF proteins in GST affinity chromatography, and phosphorylated IRF-1 with high specificity in vitro. Using an in gel kinase assay with recombinant IRF-1 as substrate, two molecular weight forms of the kinase (43 and 38 kDa) were identified. Biochemical criteria identified the kinase activity as the alpha catalytic subunit of casein kinase II (CKII). Furthermore, far western analysis of protein-protein interactions demonstrated that casein kinase II directly interacted with IRF-1 protein. Deletion mutation analysis of IRF-1 revealed that IRF-1 was phosphorylated at two clustered sites, one located between amino acids 138-150, the other in the C-terminal acidic activation domain between amino acids 219-231. Cotransfection studies comparing wild type and point mutated forms of IRF-1 demonstrated that mutations of the four phosphoaceptor residues in the C-terminal transactivation domain, significantly decreased transactivation by IRF-1, indicating that casein kinase II may be involved in the regulation of IRF-1 function. Strikingly, the casein kinase II clusters in IRF-1 resemble the sites identified in the C-terminal PEST domain of I<κBα [29]. The present experiments, together with previously published studies with I<κBα, c-Jun and other proteins, indicate a broad role for casein kinase II phosphorylation in the regulation of transcription factor activity.
Similar content being viewed by others
References
Hiscott J, Nguyen H, Lin R: Molecular mechanisms of interferon beta gene induction. SeminVirol 6: 161–173,1995
Fujita T, Sakakibara J, Sudo Y, Miyamoto M, Kimura Y, Taniguchi T: Evidence for a nuclear factor(s), IRF-1, mediating induction and silencing properties to human IFN-β gene regulatory elements. EMBO J 7: 3397–3405, 1988
Miyamoto M, Fujita T, Kimura Y, Maruyama M, Harada H, Sudo Y, Miyata T, Taniguchi T: Regulated expression of a gene encoding a nuclear factor, IRF-1, that specifically binds to the IFN-β gene regulatory elements. Cell 54: 903–913, 1988
Fujita T, Kimura Y, Miyamoto M, Barsoumian EL, Taniguchi T: Induction of endogenous IFN-α and IFN-β genes by a regulatory transcription factor IRF-1. Nature 337: 270–272, 1989
Harada H, Fujita T, Miyamoto M, Kimura Y, Maruyama M, Furia A, Miyata T, Taniguchi T: Structurally similar but functionally distinct factors, IRF-1 and IRF-2, bind to the same regulatory elements of IFN and IFN-inducible genes. Cell 58: 729–739, 1989
Nguyen H, Hiscott J, Pitha PM: The growing family of IRF transcription factors. Cyt Growth Fact Rev 8: 293–312, 1997
Reis LFL, Harada H, Wolchok JD, Taniguchi T, Vilcek J: Critical role of a common transcription factor, IRF-1, in the regulation of IFN-β and IFN-inducible genes. EMBO J 11: 185–193, 1992
Uegaki K, Shirakawa M, Fujita T, Taniguchi T, Kyogoku Y: Characterization of the DNA binding domain of the mouse IRF-2 protein. Protein Eng 6: 195–200, 1993
Lin R, Mustafa A, Nguyen H, Hiscott J: Mutational analysis of interferon (IFN) regulatory factors 1 and 2: Effects on the induction of IFN-β gene expression. J Biol Chem 269: 17542–17549, 1994
Yamamoto H, Lamphier MS, Fujita T, Taniguchi T, Harada H: The oncogenic transcription factor IRF-2 possesses a transcriptional repression and a latent activation domain. Oncogene 9: 1423–1428, 1994
Harada H, Kitagawa M, Tanaka N, Yamamoto H, Harada K, Ishihara M, Taniguchi T: Anti-oncogenic and oncogenic potentials of interferon regulatory factors-1 and-2. Science 259: 971–974, 1993
Willman CL, Sever CE, Pallavicini MG, Harada H, Tanaka N, Slovak ML, Yamamoto H, Harada K, Meeker TC, List AF, Taniguchi T: Deletion of IRF-1, mapping to chromosome 5q31.1, in human leukemia and preleukemic myelodysplasias. Science 259: 968–971, 1993
Harada H, Kondo T, Ogawa S, Tamura T, Kitagawa M, Tanaka N, Lamphier MS, Hisamaru H, Taniguchi T: Accelerated exon skipping of IRF-1 mRNA in human myelodysplasia/leukemia; a possible mechanism of tumor suppressor inactivation. Oncogene 9: 3313–3320, 1994
Kondo T, Minamino N, Nagamura-Inoue T, Matsumoto M, Taniguchi T, Tanaka N: Identification and characterization of nucleophosmin/B23/numatrin which binds the anti-oncogenic transcription factor IRF-1 and manifests oncogenic activity. Oncogene 15: 1275–1281, 1997
Kirchoff S, Koromilas AE, Schaper F, Grashoff M, Sonenberg N, Hauser H: IRF-1 induced cell growth inhibition and interferon induction requires the activity of the protein kinase PKR. Oncogene 11: 439–445, 1995
Yang Y-L, Reis LFL, Pavlovic J, Aguzzi A, Schaefer R, Kumar A, Williams BRG, Aguet M, Weissman C: Deficient signalling in mice devoid of double-stranded RNAdependent protein kinase. EMBO J 14: 6095–6106, 1995
Kumar A, Yang Y-L, Flati V, Der S, Kadereit S, Deb A, Haque J, Reis L, Weissman C, Williams BRG: Deficient cytokine signalling in mouse embryo fibroblasts with a targeted deletion in the PKR gene: Role of IRF-1 and NF-κB. EMBO J 16: 406–416, 1997 179
Tamura T, Ishihara M, Lamphier MS, Tanaka N, Oishi I, Aizawa S, Matsuyama T, Mak TW, Taki S, Taniguchi T: An IRF-1-dependent pathway of DNA damage-induced apoptosis in mitogen-activated T-lymphocytes. Nature 376: 596–599, 1995
Ozawa H, Matsuyama T, Mak TW, Aizawa S, Tokino T, Oren M, Taniguchi T: Cooperation of the tumour suppressors IRF-1 and p53 in response to DNA damage. Nature 382: 816–818, 1996
Nguyen H, Lin R, Hiscott J: Activation of multiple growth regulatory genes following inducible expression of IRF-1 or IRF/RelA fusion proteins. Oncogene 15: 1425–1435, 1997
Tan RS, Taniguchi T, Harada H: Identification of the lysyl oxidase gene as target of the antioncogenic transcription factor, IRF-1, and its possible role in tumor suppression. Cancer Res 56: 2417–2421, 1996
Matsuyama T, Kimura T, Kitagawa M, Watanabe N, Kundig T, Amakawa R, Kishihara K, Wakeham A, Potter J, Furlonger C, Narendran A, Suzuki H, Ohashi P, Paige C, Taniguchi T, Mak T: Targeted disruption of IRF-1 or IRF-2 results in abnormal type I IFN induction and aberrant lymphocyte development. Cell 75: 83–97, 1993
Taki S, Sato T, Ogasawara K, Fukuda T, Sato M, Hida S, Suzuki G, Mitsuyama M, Furuta T, Kojima S, Taniguchi T, Asano Y: Multistage regulation of Th1-type immune responses by the transcription factor IRF-1. Immunity 6: 1–20, 1997
Duncan GS, Mittrucker HW, Kagi D, Matsuyama T, Mak TW: The transcription factor interferon regulatory factor-1 is essential for natural killer cell function in vivo. J Exp Med 184: 2043–2048, 1996
Kamijo R, Harada H, Matsuyama T, Bosland M, Gerecitano J, Shapiro D, Le J, Koh SI, Kimura T, Green SJ, Mak TW, Taniguchi T, Vilcek J: Requirement for transcription factor IRF-1 in NO synthase induction in macrophages. Science 263: 1612–1615, 1994
Martin E, Nathan C, Xie Q: Role of interferon regulatory factor 1 in induction of nitric oxide synthase. J Exp Med 180: 977–984, 1994
Pinna LA: Casein kinase II: An ‘Eminence grise’ in cellular regulation? Biochim Biophys Acta 1054: 267–284, 1990
Lin R, Gewert D, Hiscott J: Differential transcriptional activation in vitro by NF-κB/Rel proteins. J Biol Chem 270: 3123–3131, 1995
Lin R, Beauparlant P, Makris C, Meloche S, Hiscott J: Phosphorylation of IκBα in the C-terminal PEST domain by casein kinase II affects intrinsic protein stability. Mol Cell Biol 16: 1401–1409, 1996
Hibi M, Lin A, Minden M, Karin M: Identification of an oncoproteinand UV-responsive protein kinase that binds and potentiates the c-Jun activation domain. Genes Dev 7: 2135–2148, 1993
Garoufalis E, Kwan I, Lin R, Mustafa A, Pepin N, Roulston A, Lacoste J, Hiscott J: Viral induction of the human interferon beta promoter: Modulation of transcription by NF-κB/rel proteins and interferon regulatory factors. J Virol 68: 4707–4715, 1994
Thanos D, Maniatis T: Identification of the rel family members required for virus induction of the human beta interferon gene. Mol Cell Biol 15: 152–164, 1995
Leblanc J-F, Cohen L, Rodrigues M, Hiscott J: Synergism between distinct enhanson domains in viral induction of the human beta interferon gene. Mol Cell Biol 10: 3987–3993, 1990
Meisner H, Czech MP: Phosphorylation of transcriptional factors and cell-cycle-dependent proteins by casein kinase Il. Curr Opin Cell Biol 3: 474–483, 1991
Krek W, Maridor G, Nigg EA: Casein kinase II is a predominantly nuclear enzyme. J Cell Biol 116: 43–55, 1992
Yu IJ, Spector DL, Bac Y-S, Marshak DR: Immunocytochemical localization of casein kinase II during interphase and mitosis. J Cell Biol 114: 1217–1232, 1991
Lin A, Frost J, Deng T, Smeal T, al-Alawi N, Kikkawa U, Hunter T, Brenner D, Karin M: Casein kinase II is a negative regulator of c-Jun DNA binding and AP-1 activity. Cell 70: 777–789, 1992
McElhinny JA, Trushin SA, Bren GD, Chester N, Paya CV: Casein kinase II phosphorylates IκBα at S-283, S-289,S-293, and T-291 and is required for its degradation. Mol Cell Biol 16: 899–906, 1996
Barroga CF, Stevenson JK, Schwarz EM, Verma IM: Constitutive phosphorylation of IκBα by casein kinase II. Proc Natl Acad Sci USA 92: 7637–7641, 1995
Roulston A, Lin R, Beauparlant P, Wainberg MA, Hiscott J: Regulation of HIV-1 and cytokine gene expression in myeloid cells by NF-κB/Rel transcription factors. Microbiol Rev 59: 481–505, 1995
Thanos D, Maniatis T: NF-κB: A lesson in family values. Cell 80: 529–532, 1995
Verma IM, Stevenson JK, Schwarz EM, Antwerp DV, Miyamoto S: Rel/NFκB/IκB family: Intimate tales of association and dissociation. Genes Dev 9: 2723–2735, 1995
Baeuerle PA, Henkel T: Function and activation of NF-κB in the immune system. Annu Rev Immunol 12: 141–179, 1994
Baldwin AS Jr: The NF-κB and IκB proteins: New discoveries and insights. Annu Rev Immunol 14: 649–681, 1996
Brown K, Gerstberger S, Carlson L, Franzoso G, Siebenlist U: Control of IκB-α proteolysis by site-specific, signal-induced phosphorylation. Science 267: 1485–1488, 1995
Brockman JA, Scherer DC, McKinsey TA, Hall SM, Qi X, Lee WY, Ballard DW: Coupling of a signal response domain in IκBα to multiple pathways for NF-κB activation. Mol Cell Biol 15: 2809–2818, 1995
Traenckner EBM, Pahl HL, Henkel T, Schmidt KN, Wilk S, Baeuerle PA: Phosphorylation of human IκBα on serines 32 and 36 controls IκBα proteolysis and NF-αB activation in response to diverse stimuli. EMBO J 14: 2876–2883, 1995
Chen Z, Hagler J, Palombella VJ, Melandri F, Scherer D, Ballard D, Maniatis T: Signal-induced site-specific phosphorylation targets IκBα to the ubiquitin-proteasome pathway. Genes Dev 9: 1586–1597, 1995
Scherer DC, Brockman JA, Chen Z, Maniatis T, Ballard DW: Signal-induced degradation of IκBα requires site-specific ubiquitination. Proc Natl Acad Sci USA 92: 11259–11263, 1995
Rodriguez MS, Wright J, Thompson J, Thomas D, Baleux F, Virelizier JL, Hay RT, Arenzana-Seisdedos F: Identification of lysine residues for signal-induced ubiquitination and degradation of IκBα in vivo. Oncogene 12: 2425–2435, 1996
Régnier C, Song HY, Gao X, Goeddel DV, Cao Z, Rothe M: Identification and characterization of an IκB kinase. Cell 90: 373–383, 1997
DiDonato JA, Hayakawa M, Rothwarf DM, Zandi E, Karin M: A cytokine-responsive IκB kinase that activates the transcription factor NF-κB. Nature 388: 548–554, 1997
Rodriguez MS, Michalopoulos I, Arenzana-Seisdedos F, Hay RT: Inducible degradation of IκBα in vitro and in vivo requires the acidic C-terminal domain of the protein. Mol Cell Biol 15: 2413–2419, 1995
Beauparlant P, Lin R, Hiscott J: The role of the C-terminal domain of IκBα in protein degradation and stability. J Biol Chem 271: 10690–10696, 1996
Ernst MK, Dunn LL, Rice NR: The PEST-like sequence of IκBα is responsible for inhibition of DNA binding but not for cytoplasmic retention of c-Rel or RelA homodimers. Mol Cell Biol 15: 872–882, 1995
Jaffray E, Wood KM, Hay RT: Domain organization of IκBα and the sites of interaction with NF-αB p65. Mol Cell Biol 15: 2166–2172, 1995
Brown K, Franzoso G, Baldi L, Carlson L, Mills L, Lin Y-C, Gerstberger S, Siebenlist U: The signal response of IκBα is regulated by transferable N-and C-terminal domains. Mol Cell Biol 17: 3021–3027, 1997
Beauparlant P, Kwon H, Clarke M, Lin R, Sonenberg N, Wainberg M, Hiscott J: Transdominant mutants of IκBα block Tat-TNF synergistic activation of HIV-1 expression and virus multiplication. J Virol 70: 5777–5785, 1996 180
Wu B-Y, Woffendin C, Duckett CS, Ohno T, Nabel GJ: Regulation of human retroviral latency by the NF-κB/IαB family: Inhibition of human immunodeficiency virus replication by licb through a Rev-dependent mechanism. Proc Natl Acad Sci USA 92: 1480–1484, 1995
Fritz CC, Zapp ML, Green MR: A human nucleoporin-like protein that specifically interacts with HIV Rev. Nature 376: 530–533, 1995
Bogerd H, Flidell RA, Madore S, Cullen BR: Identification of a novel cellular cofactor for the Rev/Rex class of retroviral regulatory proteins. Cell 82: 485–494, 1995
Stutz F, Neville M, Rosbash M: Identification of a novel poreassociated protein as a functional target of the HIV-1 Rev protein in yeast. Cell 82: 495–506, 1995
Fritz CC, Green MR: HIV Rev uses a conserved cellular protein export pathway for the nucleocytoplasmic transport of viral RNAS. Curr Biol 6: 848–854, 1996
Arenzana-Seisdedos F, Thompson J, Rodriguez MS, Bachelerie F, Thomas D, Hay RT: Inducible nuclear expression of newly synthesized IκBα negatively regulates DNA-binding and the transcriptional activities of NF-αB. Mol Cell Biol 15: 2689–2696, 1995
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lin, R., Lin, R., Hiscott, J. et al. A role for casein kinase II phosphorylation in the regulation of IRF-1 transcriptional activity. Mol Cell Biochem 191, 169–180 (1999). https://doi.org/10.1023/A:1006850009017
Issue Date:
DOI: https://doi.org/10.1023/A:1006850009017