Abstract
Fungi are the organisms present universally in the environment from ancient times. Being regarded as an essential part of human mycobiome either as commensals or as opportunistic pathogens they hold a spectacular place in our lives. Mycobiome is growing as a field of interest, but some questions still remain lurking like how fungi exactly modulate health and disease. Most of the studies focus on the interactions of the immune system with the bacterial and viral community rather than highlighting the communication with mycobiota. Crosstalk between the immune system and mycobiota have been found to be highly necessary for maintaining homeostasis, preventing susceptibility towards diseases, inducing immune tolerance, modulating microbial communities and generating signal responses by interacting with fungal molecular patterns. We are highlighting the latest advancement regarding the relationship of innate and adaptive branches of immunity with mycobiota, their role ranging from simple commensals to opportunistic pathogens and more importantly therapies designed to overcome fungal infections in this review.
Similar content being viewed by others
Introduction
Robert Urich quoted “the health outside starts from inside”, which highlights the importance of all processes that take place inside a human body and account for a huge impact on their lives. Human body is a reservoir of a large number of microorganisms, their genomes, enzymes, and metabolites clubbed together and referred to as "Microbiome", which is further housed in various human organs. Whenever one hears about the microbiome, only bacteria are often picturized because, since long, they have occupied the centre of research. However, the microbiome doesn’t only revolve around bacteria, it also consists of fungal, protozoan, parasitic, archaeal, and viral species that play significant roles even after staying behind curtains. Mycobiome (first voiced in 2009) "the fungal population" acquires only 0.1% of the total microbiome — counterbalance the number of bacterial species by being larger in size. (Qin et al. 2010) (Sam et al. 2017). They regulate health and disease significantly.
Table 1 highlights the distribution of commensal and pathogenic fungi across various body sites, including skin, oral cavity, intestine, etc. Initially, fungal diversity was evaluated by culture-dependent methods, but the inability to culture most fungal species created a major barrier. However, Internal transcribed spacer (ITS) sequencing-based method (a culture-independent method) replaced conventional ones for identifying fungi. These ITS regions are fungal rDNA sequences that are highly divergent among different fungal species in terms of length and sequence, helps to categorize fungi at the species level (Underhill and Iliev 2014). Studies have documented that maternal inheritance has also been observed in the case of mycobiota apart from bacteriome. An infant's birth mode influences the prevalence of site-specific mycobiome: vaginally born infants have higher levels of Candida albicans on their skin, whereas those delivered by caesarean-section have Candida orthopsilosis dominantly in their oral cavity (Schei et al. 2017) (Ward et al. 2018). According to the observations, mycobiome is transmitted vertically to infants from mother. Infant mycobiota, on the other hand, can also be altered by nutrition; research are in progress to understand more about human milk mycobiome composition, as C. albicans has been reported in the breast milk of mothers suffering from mammary candidiasis (Ward et al. 2017).
Notably, Colonization of fungi in early life develops homeostasis and susceptibility towards diseases. For example, Cladosporium colonization generates a state of protection against allergies, but in contrast colonization of C. albicans, Aspergillus, and Rhodotorula spp. are associated with the risk of childhood atopy (Speakman et al. 2020). Altogether, more studies should be conducted to deduce the relationship between mycobiome inheritance and its persistence.
Identification of fungal microbes
In general, Fungal Pathogen-Associated Molecular Patterns (PAMPs), consists of several layers of carbohydrates (β-glucans, chitins, and mannans) that vary in proportion among numerous fungal species. Immune response can be developed by human pattern-recognition receptors (PRRs) after encountering microbe specific molecules (Gow et al. 2017) (Erwig and Gow 2016). Numerous studies quoted 3 important PAMPs— chitin’s (polymers of N-acetylglucosamine) the size of which is significantly more important in S. cerevisiae to elicit an immunological response. Precisely, medium-sized chitin particles can induce IL-17 and TNFα production in the host. Alternatively, the small size chitin particles can induce anti-inflammatory responses (Da Silva et al. 2009). On the other hand, researches have shown that β-glucans (glucose polymers), specifically, β-1,3-glucans can interact with dectin-1 receptor which can further provoke an immune response against glucans. So far, no receptor for β-1,6-glucans (branches) has been recognized in the host. In case of Mannans (chains of mannose residues linked to fungal proteins via N-linkage or O-linkage) that are identified by host dectin-2, and mincle receptors to control fungi in numbers (Borriello et al. 2020). Overall, fungal PAMPs are the main elicitors of immunity, any failure in their recognition can be a potential cause of disease.
Fungi and morphology
Fungi often cycle amid yeast and hyphal morphologies (dimorphism). Various resources have proven that the conversion occurs either due to variation in temperature, pH or oxygen concentration of the host that drives fungi to their pathogenic state. Specifically, authors highlighted that macrophages can induce phagocytosis on encountering C. albicans yeast form, but subsequent CO2 production in macrophages promotes the transition to hyphal form, which eventually kills macrophages and ultimately disrupts the immune reactions (Hernández-Chávez et al. 2017). While in case of Malassezia, yeast form can survive as facultative intracellular parasite in keratinocytes by causing inhibition of inflammatory responses. However, sebaceous gland hyperactivity can induce morphological conversion Malassezia to hyphal form in vivo, which serves as an crucial basis for the development biofilms (Brand 2012). Meanwhile a study emphasizes that fungal morphology apart from inducing pathogenicity also governs T helper cell differentiation. For example, C. albicans yeast form, in vivo, induces TH17 responses while filamentous form induces the TH1 response. Contrastingly, in vitro, yeast form of C. albicans, via IL-12 induces TH1 response while hyphal form via IL-23 induces TH17 response (Speakman et al. 2020). Hence, these studies concluded that different morphologies are employed by fungal microbes in order to remain viable or pathogenic in host, but immune system keeps them in check.
Mycobiome and immunity
As discussed previously, Human body harbors various microorganisms, which maintains the human ecosystem through numerous interactions with the host. Fungal microbes are present all over the gut, lungs, skin etc., to maintain dynamics of host. In this part we will discuss about the receptors of host that communicate with fungi, crosstalk between immune cells and fungal species, other than that we will also examine the process of immune tolerance induced via mycobiota.
It is well documented that PRRs initiate an immune response after recognizing PAMPs of both pathogens and commensal microorganisms. C-type lectin receptors (CLRs), an antifungal PRR through Carbohydrate Recognition Domain (CRD) in the presence of calcium identify carbohydrate moiety (β-glucans, mannans) on fungal cells (Netea et al. 2015). Further, elicit innate immune response through integral immunoreceptor tyrosine-based activation motif (ITAM) or through FcRγ (Drickamer and Taylor 2015). Dectin-1 (gene ClEC7A), appears as an important CLR displayed on macrophages, mast cells, and dendritic cells which interacts with β-glucans present on fungal species (Saijo et al. 2010). Of utmost importance, Dectin-1 stimulates the release of cytokines associated with TH17, IL-22 which has anti-inflammatory as well as pro-inflammatory properties (A. et al. 2012). Evidences through studies disclosed that Dectin-1 gene polymorphism could increase the possibility of development of fungal diseases in patients who had undergone chemotherapy for acute myeloid leukemia. But more studies are required to refine about the relation of Dectin-1 gene with numerous fungal diseases (Fischer et al. 2016).
Dectin-2 (gene CLEC6A), another CLR expressed on dendritic cells, macrophages that recognize α-mannans in fungal cell wall and accelerates the release of cytokines through FcRγ and Syk-CARD9-NFκB signaling pathway. Clinical studies confirmed the presence of severe C. albicans infections in Dectin-2 knockout mice against the wild one. However, the response of dectin-2 deficient mice towards C. neoformans is normal indicating that mechanism of host defense is distinctive among different fungi. Another study by Saijo et al., validated that dectin-2 deficiency can enhance fungal load in kidney and subsequently lead to the development of hydronephrosis condition (Saijo et al. 2010). Apart from these, Cryptococcosis and aspergillosis are prominently reported in the individuals having dectin-2 polymorphism (Patin et al. 2019). Taken together, all these researches stress upon the significance of dectin-2 receptor in fungal infections.
Macrophage inducible C-type lectin (Mincle), distinctively recognizes mannans on fungal cell wall. Although they are not present on inactive macrophages rather their expression is stimulated by Macrophage C-type lectin (MCL) (Miyake et al. 2015). A study demonstrated that mincle can recognize Malassezia spp., C. albicans and potentiate the release of chemokines and cytokines against fungi. Their deficiency is often associated with C. albicans infections (Patin et al. 2019).
Toll-like Receptor (TLRs), another PRR serve as signaling receptor, are composed of two domains: N-terminal and C-terminal domain. Former one constitutes about 16–28 leucine-rich repeats, which recognize PAMPs while the latter one commences downstream signaling (Saijo et al. 2010). Multiple studies highlight distinctive functions of TLRs in recognition of fungus specific molecules, for example, TLR2 mainly recognizes fungal β-glucans and C. albicans phospholipomannans (PLMs) (Bourgeois and Kuchler 2012). Moreover, they are also responsible for monocyte-mediated apoptosis of C. albicans through caspase 3 and caspase 8 (Dreschers et al. 2016). On the other hand, TLR4 interacts with C. neoformans glucuronoxylomannan (GXM) and C. albicans mannans (O-linked). However, C. albicans can cause systemic infections in TLR4 knockout mice by impairing neutrophil recruitment and chemokine responses. All apart, TLR6 overcome fungal species by inducing TH17 antifungal response. Furthermore, it has been demonstrated that any polymorphism in TLR-4, TLR-6 can cause aspergillosis (Skevaki et al. 2015). Meanwhile, other TLRs are also concerned with fungal genome detection like TLR-7 gets activated by single-stranded fungal RNA, while TLR3 interacts with double-stranded fungal RNA and TLR-9 recognizes the unmethylated fungal DNA (CpG) (Bourgeois and Kuchler 2012). Overall, TLRs had displayed their potential function in revealing the identity of fungal microbes and initiates various mechanisms to break the chain of fungal evasion.
Involvement of Nod-like receptors (NLRs) is required to detect internalized microbial components of PAMPs and to further induce inflammatory caspases, along with cytokines IL-1β and IL-18 (Pathakumari et al. 2020). Recent research indicates that C. albicans’s morphological transition can activate both NLRP-3 and IL-1β. However, the authors also stated that NLRP-3 polymorphism can impair IL-1β production, which triggers the development of vulvovaginal candidiasis. Additionally, NLRP -/- mice had shown susceptibility towards C. albicans and C. neoformans infections (M. et al. 2017).
RIG-I-like Receptors (RLR), regarded as a cytosolic sensor, also shown to have a minor role in fungal recognition. It has 3 major receptors out of which MDA5 (Melanoma differentiation-associated gene 5) receptor altered expression is generally found in the case of chronic mucocutaneous candidiasis, owing to its crucial role in antifungal defense (M. et al. 2017).
Crosstalk between immune cells and fungal species
A latest study by de Zuani et al. confirmed that BMMCs (Bone marrow-derived mast cells) come into play after the detection of C. albicans and secrete some soluble factors to enhance the crawling movement of macrophages toward fungi. Additionally, ROS responses are also generated by BMMCs to overcome C. albicans population. Interestingly, When Dectin-1 knockout BMMCs are exposed to C. albicans (both hyphal and yeast forms), cytokine release is reduced but not totally suppressed, indicating that Dectin-1 is not the single receptor present on mast cells responsible for C. albicans identification. (De Zuani et al. 2018).
Macrophages (phagocytes derived from blood monocytes), a member of antigen presenting cells (APCs) that perform fungal phagocytosis and effectively generate reactive nitrogen intermediates and reactive oxygen species (antimicrobic components). Through research, it was concluded that on encounter with fungi, macrophages differentiate to M1 (classical activated macrophages) and M2 (alternative activated macrophages). Specifically, M1 macrophages generate anti-inflammatory responses, while M2 macrophages generate pro-inflammatory responses against fungi (Reales-Calderón et al. 2014). More studies can be performed to highlight the specific relationship between macrophages and mycobiota.
Dendritic cells (DCs) are effectual APCs, in general two types of DCs are prominent in host for combating fungal infections: Myeloid dendritic cells (mDCs) that offer fungal PAMPs to naïve T cells transforming them to TH1 cells. In contrast, plasmacytoid dendritic cells (pDCs), another subset of DCs, differentiate naïve T cells to TH2 cells and potentially generate IFN-α, TNF-α which inhibits hyphal growth of A. fumigatus. However, previous studies have reported that any imbalance in between pDCs and mDCs levels can increase the incidence of fungal infections among individuals (Pathakumari et al. 2020).
Natural killer cells (NK) constitute about 5–10% of the total lymphocytes but their role remains endless regarding the dialogue with mycobiome. A number of sources have proven that they utilize different receptors to crosstalk with fungi. For instance, through NKp30 activating receptor they elicit perforin degranulation of C. albicans, C. neoformans and further through NKp46 receptor they mediate destruction of C. glabrata (Li et al. 2013) (Schmidt et al. 2017). Nevertheless, they also induce cytokine response against pathogenic fungi as in case where neutrophils are activated by IFN-γ and GM-CSF (Granulocyte–macrophage colony stimulating factor) to overcome C. albicans infection in host. Many clinical trials have shown that mice without NK cells are more prone to aspergillosis and candidiasis. (Schmidt et al. 2017) (Bär et al. 2014).
Neutrophils are the initial line of defense against infections. According to the literature, neutrophils inhibit the morphological transition of C. albicans and kill them via production of neutrophil extracellular traps (NET) (Wheeler et al. 2017). Meanwhile, they also produce IL-17, besides TH17, to overcome pulmonary aspergillosis infection (Werner et al. 2011). As per preliminary research, neutropenia (a lack of neutrophils) and GI mucosal breaching enhance the risk of fungal infections (aspergillosis, candidiasis) and mortality.(Wheeler et al. 2017). However, neutropenia has not been identified as a concern in the case of cryptococcosis (Hirai et al. 2011).
T Cells
T cells have been widely investigated for their interaction with the fungal community. Specifically, TH1 cells are responsible for production of TNF, IFN-γ and GM-CSF which promotes phagocytosis, affects APC function and further, stimulates the discharge of antibodies from B-cells against fungal pathogens (Pathakumari et al. 2020) (Kroetz and Deepe 2012). The function of GM-CSF has been observed in GM-CSF knockout mice, where the levels of TH1 and TH2 cytokines drastically decrease upon encountering C. neoformans (Chen et al. 2007). However, in normal mice, GM-CSF promotes ROS response via macrophages by sequestering Zn through upregulation of Zn exporters. Zn deficiency decreases fungal proliferation, inhibits immune cell responses, and further restricts the growth of intracellular yeast in mice (Subramanian Vignesh et al. 2013).
It is well known that TH2 subclass obstructs the protective TH1 cell responses and reduces pro-inflammatory cytokine synthesis (Pathakumari et al. 2020). Prior research suggests that equilibrium amid TH2 and TH1 responses are worthy to safeguard against fungal pathogens. For instance, increased GATA-3 (TH2 cell transcription factor) expression can suppress IFN-γ production and develop the risk of systemic candidiasis. Furthermore, TH2 CD4 + T cells can compromise the function of lungs and subsequently cause allergic bronchopulmonary aspergillosis (ABPA). Overall, prevalent TH2 responses produced during fungal infections are deleterious and fatal (van de Veerdonk and Netea 2010). TH2 responses, on the other hand, can help retain humoral immunity and protect against helminthic infections by producing interleukins (IL-4, IL-5, and IL-13).(Nakayama et al. 2017).
TH17 cells get activated via MYD88, mannose receptor or by SYK-CARD9 signaling pathways induced by macrophages and dendritic cells (Romani 2004). Park et al. discovered that γδ T cells in the dermal region of skin produce IL-17 cytokine after confronting with C. albicans and after 7 days they switch to αβ TH17 effector T cells. Further, it is observed that after 30 days, these TH17 CD4 + resident memory T cells (TRM) predominate against C. albicans more rapidly than migratory memory CD4 T cells (Park et al. 2018). This research indicates that substitution of γδ T cells with αβ T cells holds importance for generation of protective immunity against fungi. Additionally, it is well documented that mice with a genetic mutation in IL-17A gene, could impair IL-17 pathway and thereby increase fungal burden (Sparber et al. 2019).
Regulatory T-cells (Treg) are regarded as the potent regulators of immune responses because they control inflammation via blocking the host tissue damage (Romani and Puccetti 2007). Two major Treg cells have been seen during fungal infections — natural (thymus derived-nTreg, CD4 + CD25 +), that release IL-10 and CTLA-4 cytokines at the initial stage of infection by A. fumigatus, to control inflammation triggered by neutrophils in the lungs. While another subset is induced under infection- (iTreg) that readily suppress the immune response mediated by TH2 at the later stage of aspergillosis (Montagnoli et al. 2006). Similarly, study by Whibley et al. highlighted that Forkhead box P3 (Foxp3 +) nTreg cells can induce TH17 and inhibit TH1, TH2 responses in systemic candidiasis infection (Whibley et al. 2014). Taken together, Treg cell subsets can control both arms of immunity to balance immune tolerance and inflammation in a coordinated manner during fungal infections (Romani and Puccetti 2007) (Fig. 1).
Immune tolerance and mycobiota
Tryptophan (Trp) is an essential amino acid procured from diet, which promotes synthesis of proteins and biogenic amines. Numerous studies have revealed that indoleamine 2,3‐dioxygenase 1 (IDO1), promotes oxidation of Trp through kynurenine pathway, which holds a vital position in generation of immune tolerance to fungi. Neutrophils, macrophages, tolerogenic DCs, epithelial cells express IDO1 and regulate immune homeostasis by mounting Kyn-mediated Treg cells through Aryl hydrocarbon Receptor (AhR) and repress inflammation by decreasing the Trp levels and finally controlling the growth of pathogen (Romani et al. 2014) (de Araújo et al. 2017). It is well documented that catabolism of Trp to L‐kynurenine is mediated by IDO1 after interaction with fungal microbes, promote the transcription of FOXP3 and dampen the transcript formation of RAR‐related orphan receptor gamma (RORγ‐t), in order to suppress the host immunity and promote the persistence of fungi, finally contributing to immune tolerance through Treg cells. However, TH17 cells inhibit Trp catabolism and favors pathology (de Araújo et al. 2017). Overall, during fungal infections, IDO1 holds the responsibility to maintain a level amid TH17 and Treg cells so as to generate a state of suppressed inflammation and immune tolerance to prevent tissue damage. Conversely, Fungi can obstruct the tolerogenic state of host, thus contributing in either commensalism or persistence of fungi.
Diseases caused by mycobiota
There has been growing concern in recent years regarding the role of fungi in the development of human diseases. Dysbiosis (an imbalance in which harmful microorganisms outweigh beneficial microbes) frequently causes fungal infections, disrupting the fungus-immune system balance, which can be lethal if ignored. Thus, in this section we will examine various mycobiota that have become causative agents of disease. Table 2 illustrates fungal diseases and their antifungal therapies.
Candidiasis
Candida spp. are natural inhabitants of human mycobiome, and it is evident from studies that the morphological transition can promote their conversion from commensals to pathogens, causing mucocutaneous, oral, and invasive infections. Specifically, invasive candidiasis results due to persistence of C. albicans in blood (candidemia) or peritonitis, or in osteomyelitis (deep-seated organ infections). From the findings, except C. albicans different species of Candida can also cause candidiasis in humans— C. glabrata, C. krusei, C. tropicalis, C. guilliermondii and C. parapsilosis, which differ in virulency. Candida spp. prevails in the body due to several reasons. First of which is extensive application of broad-spectrum antibiotics causing dysbiosis of bacterial species and flare up fungal species, second: cytotoxic chemotherapy-induced mucositis that ruptures GI and cutaneous layers, third being the presence of central venous catheters through which fungus can gain entry in the circulating blood and span to different regions (Pappas et al. 2018). A novel study conducted on murine model showed that kidney suffers from major candidiasis because of late initiation of innate immune response, minimal immune cells residing followed by their gradual enrollment (Hebecker et al. 2016). Meanwhile, recent study had documented a link between candidiasis and COVID-19, for example, it has been testified that due to prolonged ICU stays intrusive yeast infections have higher frequencies, especially the one’s which are caused by C. albicans, C. auris, C. glabrata, and C. parapsilosis were observed in patients with COVID-19 (Arastehfar et al. 2020).
Aspergillosis
Aspergillosis, the most prevalent cause of mortality across the globe, caused by ten different species. Some of which are Aspergillus fumigatus, Aspergillus flavus, Aspergillus niger, Aspergillus terreus, Aspergillus versicolor, etc. (Vivek-Ananth et al. 2018). Overall, these fungal microbes pave their way into the host via inhalation and can prove to be fatal if sustained in the host. For instance, Aspergillus conidial spores in case of immunocompromised patients operate as opportunistic pathogens while in heathy individual’s alveolar macrophages engulf and remove conidial spores effectively (Guerra et al. 2017). Subsequently, a study by Sainz et al. reported that patients with neutropenia, dectin-1 polymorphism, or who had received transplants (hematopoietic stem cell) had a higher chance of invasive aspergillosis (Sainz et al. 2012). Recently a study reported that ICU patients with influenza can develop influenza-associated pulmonary aspergillosis (IAPA). Furthermore, this study in patients with serious SARS-CoV-2 infection can give rise to the probability of COVID-19-associated pulmonary aspergillosis (CAPA) (Verweij et al. 2020). In this context, further research is needed to assess the impact of aspergillosis in patients with viral infections.
Cryptococcosis
Cryptococcosis, a widespread invasive fungal infection, responsible for 20–70% mortality across the globe. Cryptococcus is a pathogenic fungus that gains entry into human body through inhalation but in an immunocompromised state the immune system is inefficient to remove them and thus, C. neoformans and C. gatti become predominant and cause cryptococcosis (Iliev and Underhill 2013). Emerging studies indicate the presence of different types of cryptococcosis depending upon the site of infection which includes: cutaneous, peripheral, disseminated, and cerebral cryptococcosis. Among these, cerebral cryptococcosis has been reported as invasive opportunistic infection that can cause meningoencephalitis in HIV-infected patients (Pathakumari et al. 2020) (Iliev and Underhill 2013). Meanwhile, another study indicates that cell wall polysaccharides of Cryptococcus namely GXM, glucuronoxylomannogalactan and melanin, can alter the activity of macrophages in circulation (oxidative burst) (Pathakumari et al. 2020). Also, C. neoformans produce giant cells (titan) that subsequently decrease the phagocytic activity of macrophages thus, promoting survival of C. neoformans (Zaragoza and Nielsen 2013).
Fungal Biofilms
Generally, packed fungal communities organize themselves in biofilms—shielded by extracellular matrix (ECM) to promote their adherence to the surfaces and protect them against host immune responses. Such fungal biofilms can colonize mucosal surfaces like oral and vaginal areas as well as on implanted medical equipments like catheters, dentures, etc. (Kernien et al. 2018). Notably, Cryptococcus biofilms originate from surface attachment taken over by microcolonies, ECM production and eventually get mature (R. et al. 2015).
Studies demonstrated that shedding of polysaccharide capsules (GXM, galactoxylomannans, and mannoproteins) by C. neoformans can enhance ECM production. Additionally, presence of GXM in biofilm matrix provides resistance against NET, antifungals and further reduces the phagocytic response of macrophages. But still, it is unclear that whether the phagocytic response is sufficient enough to overcome Cryptococcus biofilms or not. Meanwhile, Candida biofilms can also inhibit the release of NET and ROS and altogether promote the growth of biofilms. Surprisingly, IL-17A has proven to induce the formation of C. albicans biofilms. While in contrast to Cryptococcus and Candida biofilms, Aspergillus biofilms have distinct ECM composition including: galactosaminogalactan (GAG) and galactomannan as core matrix polysaccharides. It is evident from studies that absence of GAG can reduce the probability of biofilm formation by A. fumigatus (Kernien et al. 2018).
Fusariosis
Another fungal species found in the environment, Fusarium that exists in mold form. Out of 70 species that are known, only 3 species (F. solani species, F. dimerum species, and F. oxysporum species) can cause disseminated (fusariosis) and superficial infections in transplant recipients, immunocompromised and diabetic patients (Guarro 2013) (Short et al. 2013) (Batista et al. 2020). Studies have documented that fusarium when penetrates via skin can generate multiple erythematous papules that further disseminates to whole body and proves to be fatal for immunocompromised patients. Additionally, Fusarium can also invade through contact lenses, which results in development of keratitis. The only therapy to eradicate fusariosis to date is Combination therapy (Batista et al. 2020).
Pityriasis versicolor (PV)
Malassezia is responsible for Pityriasis Versicolor (tinea versicolor), a common skin disease. The preliminary study concluded that the morphological transformation of Malassezia along with other factors such as hormonal imbalance, excessive use of antibiotics and high temperature can promote the susceptibility towards disease. Another study suggests that Pityriasis Versicolor (PV) triggers the development of hyperpigmentation or hypopigmented skin lesions (Dyląg et al. 2020). Specifically, in hypopigmented PV, Malassezia releases azelaic acid and malassezin that promotes apoptosis of melanocytes and subsequently halts the synthesis of melanin (Dolenc-Voljč 2017). However, the treatment of PV is possible through antifungal drugs, but results won’t appear spontaneously due to persistence of dead fungal cells on the skin (Dyląg et al. 2020).
Therapies against fungal infections
Fungal infections are more prominent worldwide and cause mortality with unacceptable high rates which shows a need for therapies to overcome fungal infections. In this section, we will discuss about some existing therapies and shortcomings associated with therapies.
Immunotherapy
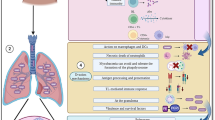
Currently, immunotherapy is one of the remarkable strategies to overcome fungal infections in which GM-CSF with IFN- γ protect against invasive aspergillosis. Precisely, GM-CSF can enhance antifungal ability and IFN- γ clears the pathogenic fungi in transplant recipients who had developed systemic fungal infections (Armstrong-James et al. 2017). A study by Jarvis et al. highlighted that adjunctive IFN- γ immunotherapy can eradicate C. neoformans from HIV-1 infected patient’s cerebrospinal fluid much faster than conventional therapies (Jarvis et al. 2012). Additionally, by restoring immunological function, adjunctive IFN- γ immunotherapy can be beneficial to individuals infected with invasive Candida or Aspergillus infections (Delsing et al. 2014). Although IFN- γ therapy have shown positive results in numerous studies but still more trials are needed to ensure its worth. Apart from these, PRR agonists and antagonists can also stimulate an immune response against fungi by subsequently limiting inflammation. For instance, conjugation of β-glucan laminarian with diphtheria toxoid (CRM 1197) can promote the development of antibodies against β-glucan in mice with systemic candidiasis (Armstrong-James et al. 2017) (Patin et al. 2019). Furthermore, researchers have employed CAR T cell technologies to produce cytotoxic T cells (Tc Cells) against pathogenic fungi. This research has given positive results in halting the growth of A. fumigatus hyphae after recognizing β-glucans moieties (Patin et al. 2019).
Antifungal drugs
Presently, antifungal drugs exist in three classes for the therapy of systemic fungal infections which includes azoles inhibiting ergosterol biosynthesis, echinocandins target fungal cell wall biosynthesis, and polyenes facilitate cell lysis (Revie et al. 2018). In a recent clinical study, antifungal drug posaconazole has proven to reduce the occurrence of invasive fungal infections in immunocompromised patients. Nevertheless, this drug is often linked with some side effects like nausea, vomiting, and malfunctioning of liver (Wong et al. 2020). Another study by Pierce et al. reported that compound 61894700 can halt the development of C. albicans filament and biofilms but is unable to inhibit the overall planktonic growth (Pierce et al. 2015). Additionally, two antifungal drugs had been developed namely sulfadiazine–pyrimethamine (SDZ/PYR) and sulfamethoxazole-trimethoprim (SMX/TMP) that repress growth and biofilm formation of C. gatti and C. neoformans (de Aguiar Cordeiro et al. 2013). Meanwhile, many researchers have repurposed some old drugs like tamoxifen, Cox2 inhibitors that had shown positive results against fungal infections. In addition to this, combinations of antifungal drugs are currently being employed to increase their efficacy, where the synergistic combinations make it effective even at low doses and reduce the resistance (Spitzer et al. 2017).
Vaccines
Vaccines act as potential weapons against fungal infections. For instance, live-attenuated fungal vaccines had given appreciating results during preclinical studies, but these are not approved for immunocompromised patients (Santos and Levitz 2014). A study by Spellberg et al. observed that the rAls3p-N (adhesion protein associated with C. albicans cell wall) based vaccine would protect against disseminated, mucosal, and vulvovaginal candidiasis, and are under phase-II trials (Spellberg et al. 2006). Apart from that, recombinant protein Asp f 16 (association with unmethylated CpG oligodeoxynucleotides) based vaccines also proved to be effective against invasive aspergillosis (Bozza et al. 2002).
Nanotechnology
Nanotechnology has been actively researched as a low-cost, one-of-a-kind approach for developing novel antifungals and vaccines (Souza and Amaral 2017). Antifungal chemotherapy employs nanoparticles (NPs) to deliver intrinsic antifungal effects with the objective of decreasing the medication concentration required for the treatment. NPs can be utilized to increase the stability and immunogenicity of antigens like peptides in the vaccine formulation (Zhao et al. 2014). Furthermore, silver nanoparticles demonstrated increased antifungal efficiency against C. albicans ATCC 10,231 and A. brasiliensis ATCC 16,404 after conjugation with nystatin (NYT) and fluconazole (Hussain et al. 2019). Another study discovered that combining posaconazole-derived compounds with PEG-imidazole modified G5 dendrimers produced a soluble and potent posaconazole conjugate, which seems to be a new and effective antifungal medication. However, in vivo studies are necessary before nanoparticles are to be used for medical purposes (Tang et al. 2021).
Shortcomings of therapies
Antifungal drugs that are commercially available are limited and their extensive use has resulted in the development of resistance, which is a major rising health concern. Genetic variations in the cellular target, upregulation of the target molecules, and antifungal efflux pump extrusion are all factors that trigger azole resistance (A. and B. 1999). For example, mutations in drug target genes, as observed in the case of ERG11, promote their overexpression and increase the azole resistance against C. albicans infections. Furthermore, mutation in the ERG3 gene encoding for Δ-5,6- desaturase enzyme Erg3 can accumulate ergosterol and further develop resistance against azoles and polyenes (Revie et al. 2018). However, resistance to polyenes is primarily caused by sterol modification, which occurs as a result of decrease in the cell's ergosterol content, substitution of the sterol with the one that has lower affinity for polyenes, or reorientation of ergosterol that sterically inhibits polyene binding (A. and B. 1999). Apart from that, echinocandin resistance develops after therapy as a result of amino acid alterations in the 1,3-D-glucan synthase, coded by the FKS1 and FKS2 genes, that diminish the enzyme's susceptibility to medications (Perlin 2015). Thus, all these studies emphasize on necessitating the creation of new drugs to tackle antifungal resistance.
Although vaccination serves as an important strategy to overcome fungal infections, but to date no vaccine is sanctioned to treat fungal infections. Besides this, there is still considerable uncertainty about whether fungal vaccines could generate herd immunity or not (Santos and Levitz 2014).
Fungal probiotics
When given in the proper dosage, probiotics (live microorganisms) are helpful to the host. As an example, Saccharomyces boulardii, the major fungal probiotic used for the treatment of several diseases like bacterial diarrhea, acute GI diseases, also exhibits positive results against Clostridium difficile infections (Kelesidis and Pothoulakis 2012) (Sen and Mansell 2020). Other than that, they also boost immunity and improve gut microflora specifically in IBD (Inflammatory Bowel Disease). Also, it helps in restoring the tight junctions in enterocytes and regulate the expression of p120 through E-cadherin (Sen and Mansell 2020) (Abbas et al. 2014) (Terciolo et al. 2017). Although S. boulardii is genetically similar to S. cerevisiae but certain characteristics adds advantage to their extensive use such as non-sporulating nature, survival at acidic pH, viability around 55 °C and ability to metabolize different carbon sources (Sen and Mansell 2020). However, safety concerns are associated with the use of fungal probiotics for immunocompromised patients.
Conclusion and future perspective
Recently, the World Health Organization (WHO) conducted a poll to establish the first fungal priority pathogen list for public health relevance in order to define R&D priorities. The selection of fungal pathogens was based on six criteria: mortality, transmission, community burden, treatability, health-care burden, and prevalence of medication resistance. For this reason, WHO has approached experts and individuals active in antifungal healthcare delivery and research for this purpose. Multi-criteria decision analysis (MCDA) was utilized in this approach, which combines scientific data and expert opinion. There are currently six fungal pathogens on the list: Candida auris, other azole-resistant Candida spp., azole-resistant A. fumigatus, C. neoformans, C. gatti, Pneumocystis jirovecii, and Mucormycetes. More fungal pathogens and criteria will be added, as recommended by the expert panel.
From this review, it is clear that both arms of immunity are crucial for maintaining a healthy human mycobiome. However, if the balance between the two is disturbed for any reason, it can manifest as a deadly disease. Nonetheless, research to discover the obscurity of the mycobiome should be continued. Further research is needed to investigate whether mycobiome alterations reported in a number of disorders either reflect disease pathogenesis or are a consequence of the disease process. New therapeutic approaches like the use of nanoparticles need to be adopted to treat fungal complications and, hence, potentiation of the immune system that helps in maintaining the fungi in commensal mode can serve as a lesson for devising better therapeutic strategies and preventing the development of drug resistance. Moreover, cross-reaction between auto-antigens of humans with commensal fungi can offer new insight into the role of commensal fungi, particularly in autoimmune disorders. In addition to this, how does interaction between numerous fungal communities across different body sites affect humans can be brought to light by future studies only.
References
Abbas, Z., Yakoob, J., Jafri, W., Ahmad, Z., Azam, Z., Usman, M.W., Shamim, S., Islam, M.: Cytokine and clinical response to Saccharomyces boulardii therapy in diarrhea-dominant irritable bowel syndrome: a randomized trial. Eur J Gastroenterol Hepatol 26(6), 630–639 (2014)
Arastehfar, A., Carvalho, A., Nguyen, M.-H., Hedayati, M., Netea, M., Perlin, D., Hoenigl, M.: COVID-19-associated Candidiasis (CAC): an underestimated complication in the absence of immunological predispositions? J Fungi J (2020). https://doi.org/10.3390/jof6040211
Armstrong-James, D., Brown, G.D., Netea, M.G., Zelante, T., Gresnigt, M.S., van de Veerdonk, F.L., Levitz, S.M.: Immunotherapeutic approaches to treatment of fungal diseases. Lancet Infect Dis 17, 393–402 (2017). https://doi.org/10.1016/S1473-3099(17)30442-5
Bär, E., Whitney, P.G., Moor, K., Reis e Sousa, C., LeibundGut-Landmann, S.: IL-17 regulates systemic fungal immunity by controlling the functional competence of NK cells. Immunity 40, 117–127 (2014). https://doi.org/10.1016/j.immuni.2013.12.002
Batista, B.G., de Chaves, M.A., Reginatto, P., Saraiva, O.J., Fuentefria, A.M.: Human fusariosis: an emerging infection that is difficult to treat. Rev Soc Bras Med Trop 53, e20200013 (2020). https://doi.org/10.1590/0037-8682-0013-2020
Borriello, F., Zanoni, I., Granucci, F.: Cellular and molecular mechanisms of antifungal innate immunity at epithelial barriers: the role of C-type lectin receptors. Eur J Immunol 50, 317–325 (2020). https://doi.org/10.1002/eji.201848054
Bourgeois, C., Kuchler, K.: Fungal pathogens-a sweet and sour treat for Toll-like receptors. Front Cell Infect Microbiol 2, 142 (2012). https://doi.org/10.3389/fcimb.2012.00142
Bozza, S., Gaziano, R., Lipford, G.B., Montagnoli, C., Bacci, A., Di Francesco, P., Kurup, V.P., Wagner, H., Romani, L.: Vaccination of mice against invasive aspergillosis with recombinant Aspergillus proteins and CpG oligodeoxynucleotides as adjuvants. Microbes Infect 4, 1281–1290 (2002). https://doi.org/10.1016/S1286-4579(02)00007-2
Brand, A.: Hyphal growth in human fungal pathogens and its role in virulence. Int J Microbiol 2012, 517529 (2012). https://doi.org/10.1155/2012/517529
Chen, G.-H., Olszewski, M.A., McDonald, R.A., Wells, J.C., Paine, R., Huffnagle, G.B., Toews, G.B.: Role of granulocyte macrophage colony-stimulating factor in host defense against pulmonary Cryptococcus neoformans infection during murine allergic bronchopulmonary mycosis. Am J Pathol 170, 1028–1040 (2007). https://doi.org/10.2353/ajpath.2007.060595
Da Silva, C.A., Chalouni, C., Williams, A., Hartl, D., Lee, C.G., Elias, J.A.: Chitin is a size-dependent regulator of macrophage TNF and IL-10 production. J Immunol 182, 3573–3582 (2009). https://doi.org/10.4049/jimmunol.0802113
Dambuza, I.M., Levitz, S.M., Netea, M.G., Brown, G.D.: Fungal recognition and host defense mechanisms. Microbiol Spectrum 5(4), 5–4 (2017)
de Aguiar Cordeiro, R., Mourão, C.I., Rocha, M.F.G., de Farias Marques, F.J., Teixeira, C.E.C., de Oliveira Miranda, D.F., Neto, L.V.P., Brilhante, R.S.N., de Jesus Pinheiro Gomes Bandeira, T., Sidrim, J.J.C.: Antifolates inhibit Cryptococcus biofilms and enhance susceptibility of planktonic cells to amphotericin B. Eur J Clin Microbiol Infect Dis 32, 557–564 (2013). https://doi.org/10.1007/s10096-012-1774-8
de Araújo, E.F., Loures, F.V., Feriotti, C., Costa, T., Vacca, C., Puccetti, P., Romani, L., Calich, V.L.G.: Disease tolerance mediated by phosphorylated indoleamine-2,3 dioxygenase confers resistance to a primary fungal pathogen. Front Immunol 8, 1522 (2017). https://doi.org/10.3389/fimmu.2017.01522
De Zuani, M., Paolicelli, G., Zelante, T., Renga, G., Romani, L., Arzese, A., Pucillo, C., Frossi, B.: Mast cells respond to Candida albicans infections and modulate macrophages phagocytosis of the fungus. Front Immunol (2018). https://doi.org/10.3389/fimmu.2018.02829
Delsing, C.E., Gresnigt, M.S., Leentjens, J., Preijers, F., Frager, F.A., Kox, M., Monneret, G., Venet, F., Bleeker-Rovers, C.P., van de Veerdonk, F.L., Pickkers, P., Pachot, A., Kullberg, B.J., Netea, M.G.: Interferon-gamma as adjunctive immunotherapy for invasive fungal infections: a case series. BMC Infect Dis 14, 166 (2014). https://doi.org/10.1186/1471-2334-14-166
Dolenc-Voljč M (2017) Diseases caused by Malassezia species in human beings. In: Kon K, Rai Soft Tissue, Bone and Joint Infections MBT-TM of S (eds) Clinical microbiology: diagnosis, treatments and prophylaxis of infections. Academic Press, New York, pp 77–91
Dreschers, S., Saupp, P., Hornef, M., Prehn, A., Platen, C., Morschhäuser, J., Orlikowsky, T.: Reduced PICD in monocytes mounts altered neonate immune response to Candida albicans. PLoS ONE 11, e0166648 (2016). https://doi.org/10.1371/journal.pone.0166648
Drickamer, K., Taylor, M.E.: Recent insights into structures and functions of C-type lectins in the immune system. Curr Opin Struct Biol 34, 26–34 (2015). https://doi.org/10.1016/j.sbi.2015.06.003
Dyląg, M., Leniak, E., Gnat, S., Szepietowski, J.C., Kozubowski, L.: A case of anti- pityriasis versicolor therapy that preserves healthy mycobiome. BMC Dermatol 20, 9 (2020). https://doi.org/10.1186/s12895-020-00106-x
Erwig, L., Gow, N.: Interactions of fungal pathogens with phagocytes. Nat Rev Microbiol (2016). https://doi.org/10.1038/nrmicro.2015.21
Fischer, M., Spies-Weisshart, B., Schrenk, K., Gruhn, B., Wittig, S., Glaser, A., Hochhaus, A., Scholl, S., Schnetzke, U.: Polymorphisms of Dectin-1 and TLR2 Predispose to Invasive Fungal Disease in Patients with Acute Myeloid Leukemia. PLoS ONE 11, e0150632–e0150632 (2016). https://doi.org/10.1371/journal.pone.0150632
Gessner, M.A., Werner, J.L., Lilly, L.M., Nelson, M.P., Metz, A.E., Dunaway, C.W., Chan, Y.R., Ouyang, W., Brown, G.D., Weaver, C.T., Steele, C.: Dectin-1-dependent interleukin-22 contributes to early innate lung defense against Aspergillus fumigatus. Infect Immunity 80(1), 410–417 (2012)
Ghannoum, M.A., Rice, L.B.: Antifungal agents: mode of action, mechanisms of resistance, and correlation of these mechanisms with bacterial resistance. Clin Microbiol Rev. 12(4), 501–17 (1999)
Gow, N., Munro, C., Latge, J.-P.: The fungal cell wall: structure, biosynthesis, and function. Microbiol Spectr (2017). https://doi.org/10.1128/microbiolspec.FUNK-0035-2016
Guarro, J.: Fusariosis, a complex infection caused by a high diversity of fungal species refractory to treatment. Eur J Clin Microbiol Infect Dis 32, 1491–1500 (2013). https://doi.org/10.1007/s10096-013-1924-7
Guerra, E., Lee, C., Specht, C., Yadav, B., Huang, H., Akalin, A., Huh, J., Mueller, C., Levitz, S.: Central role of IL-23 and IL-17 producing eosinophils as immunomodulatory effector cells in acute pulmonary aspergillosis and allergic asthma. PLOS Pathog 13, e1006175 (2017). https://doi.org/10.1371/journal.ppat.1006175
Hebecker, B., Vlaic, S., Conrad, T., Bauer, M., Brunke, S., Kapitan, M., Linde, J., Hube, B., Jacobsen, I.D.: Dual-species transcriptional profiling during systemic candidiasis reveals organ-specific host-pathogen interactions. Sci Rep 6, 36055 (2016). https://doi.org/10.1038/srep36055
Hernández-Chávez MJ, Pérez-García LA, Niño-Vega GA, Mora-Montes HM (2017) Fungal Strategies to Evade the Host Immune Recognition. J. fungi (Basel, Switzerland) 3
Hirai, Y., Ainoda, Y., Shoji, T., Fujita, T., Yoshinaga, K., Shiseki, M., Mori, N., Teramura, M., Totsuka, K., Motoji, T.: Disseminated cryptococcosis in a non-hodgkin’s lymphoma patient with late-onset neutropenia following rituximab-CHOP chemotherapy: a case report and literature review. Mycopathologia 172, 227–232 (2011). https://doi.org/10.1007/s11046-011-9423-9
Hussain, M.A., Ahmed, D., Anwar, A., Perveen, S., Ahmed, S., Anis, I., Shah, M.R., Khan, N.A.: Combination therapy of clinically approved antifungal drugs is enhanced by conjugation with silver nanoparticles. Int Microbiol 22, 239–246 (2019). https://doi.org/10.1007/s10123-018-00043-3
Iliev, I.D., Underhill, D.M.: Striking a balance: fungal commensalism versus pathogenesis. Curr Opin Microbiol 16, 366–373 (2013). https://doi.org/10.1016/j.mib.2013.05.004
Invitation to participate in survey to establish the first WHO fungal priority pathogens list (https://www.who.int/news-room/articles-detail/invitation-to-participate-in-survey-to-establish-the-first-who-fungal-priority-pathogens-list-(fppl)
Jarvis, J.N., Meintjes, G., Rebe, K., Williams, G.N., Bicanic, T., Williams, A., Schutz, C., Bekker, L.-G., Wood, R., Harrison, T.S.: Adjunctive interferon-γ immunotherapy for the treatment of HIV-associated cryptococcal meningitis: a randomized controlled trial. AIDS 26, 1105–1113 (2012). https://doi.org/10.1097/QAD.0b013e3283536a93
Kelesidis, T., Pothoulakis, C.: Efficacy and safety of the probiotic Saccharomyces boulardii for the prevention and therapy of gastrointestinal disorders. Ther Adv Gastroenterol 5, 111–125 (2012). https://doi.org/10.1177/1756283x11428502
Kernien, J.F., Snarr, B.D., Sheppard, D.C., Nett, J.E.: The Interface between fungal biofilms and innate immunity. Front Immunol 8, 1968 (2018). https://doi.org/10.3389/fimmu.2017.01968
Kroetz, D.N., Deepe, G.S.: The role of cytokines and chemokines in Histoplasma capsulatum infection. Cytokine 58, 112–117 (2012). https://doi.org/10.1016/j.cyto.2011.07.430
Li, S.S., Kyei, S.K., Timm-McCann, M., Ogbomo, H., Jones, G.J., Shi, M., Xiang, R.F., Oykhman, P., Huston, S.M., Islam, A., Gill, M.J., Robbins, S.M., Mody, C.H.: The NK receptor NKp30 mediates direct fungal recognition and killing and is diminished in NK cells from HIV-infected patients. Cell Host Microbe 14, 387–397 (2013). https://doi.org/10.1016/j.chom.2013.09.007
Miyake, Y., Oh-hora, M., Yamasaki, S.: C-type lectin receptor mcl facilitates mincle expression and signaling through complex formation. J Immunol 194, 5366–5374 (2015). https://doi.org/10.4049/jimmunol.1402429
Ml, R., Arturo, C., Mahmoud, G., Matthew, P., Marvin, W., Pranab, M.: Biofilm formation by Cryptococcus neoformans. Microbiol Spectr (2015). https://doi.org/10.1128/microbiolspec.MB-0006-2014
Montagnoli, C., Fallarino, F., Gaziano, R., Bozza, S., Bellocchio, S., Zelante, T., Kurup, W.P., Pitzurra, L., Puccetti, P., Romani, L.: Immunity and tolerance to Aspergillus involve functionally distinct regulatory T cells and tryptophan catabolism. J Immunol 176, 1712–1723 (2006). https://doi.org/10.4049/jimmunol.176.3.1712
Nakayama, T., Hirahara, K., Onodera, A., Endo, Y., Hosokawa, H., Shinoda, K., Tumes, D.J., Okamoto, Y.: Th2 cells in health and disease. Annu Rev Immunol 35, 53–84 (2017). https://doi.org/10.1146/annurev-immunol-051116-052350
Netea, M.G., Joosten, L.A.B., van der Meer, J.W.M., Kullberg, B.-J., van de Veerdonk, F.L.: Immune defence against Candida fungal infections. Nat Rev Immunol 15, 630–642 (2015). https://doi.org/10.1038/nri3897
Pappas, P.G., Lionakis, M.S., Arendrup, M.C., Ostrosky-Zeichner, L., Kullberg, B.J.: Invasive candidiasis. Nat Rev Dis Prim 4, 18026 (2018). https://doi.org/10.1038/nrdp.2018.26
Park, C.O., Fu, X., Jiang, X., Pan, Y., Teague, J.E., Collins, N., Tian, T., Oalley, J.T., Emerson, R.O., Kim, J.H., Jung, Y., Watanabe, R., Fuhlbrigge, R.C., Carbone, F.R., Gebhardt, T., Clark, R.A., Lin, C.P., Kupper, T.S.: Staged development of long-lived T-cell receptor αβ TH17 resident memory T-cell population to Candida albicans after skin infection. J Allergy Clin Immunol 142, 647–662 (2018). https://doi.org/10.1016/j.jaci.2017.09.042
Pathakumari, B., Liang, G., Liu, W.: Immune defence to invasive fungal infections: A comprehensive review. Biomed Pharmacother 130, 110550 (2020). https://doi.org/10.1016/j.biopha.2020.110550
Patin, E.C., Thompson, A., Orr, S.J.: Pattern recognition receptors in fungal immunity. Semin Cell Dev Biol 89, 24–33 (2019). https://doi.org/10.1016/j.semcdb.2018.03.003
Perlin, D.S.: Mechanisms of echinocandin antifungal drug resistance. Ann N Y Acad Sci 1354, 1–11 (2015). https://doi.org/10.1111/nyas.12831
Pierce, C.G., Chaturvedi, A.K., Lazzell, A.L., Powell, A.T., Saville, S.P., McHardy, S.F., Lopez-Ribot, J.L.: A novel small molecule inhibitor of Candida albicans biofilm formation, filamentation and virulence with low potential for the development of resistance. NPJ Biofilms Microbiomes 1, 15012 (2015). https://doi.org/10.1038/npjbiofilms.2015.12
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li J, Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Tap J, Lepage P, Bertalan M, Batto J-M, Hansen T, Le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu H, Yu C, Li S, Jian M, Zhou Y, Li Y, Zhang X, Li S, Qin N, Yang H, Wang J, Brunak S, Doré J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J, Consortium M, Bork P, Ehrlich SD, Wang J (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65. https://doi.org/10.1038/nature08821
Reales-Calderón JA, Aguilera-Montilla N, Corbí ÁL, Molero G, Gil C (2014) Proteomic characterization of human proinflammatory M1 and anti-inflammatory M2 macrophages and their response to Candida albicans. Proteomics 14:1503–1518. https://doi.org/10.1002/pmic.201300508
Revie NM, Iyer KR, Robbins N, Cowen LE (2018) Antifungal drug resistance: evolution, mechanisms and impact. Curr Opin Microbiol 45:70–76. https://doi.org/10.1016/j.mib.2018.02.005
Romani, L.: Immunity to fungal infections. Nat Rev Immunol 4, 11–24 (2004). https://doi.org/10.1038/nri1255
Romani, L., Puccetti, P.: Controlling pathogenic inflammation to fungi. Expert Rev Anti Infect Ther 5, 1007–1017 (2007). https://doi.org/10.1586/14787210.5.6.1007
Romani, L., Zelante, T., Luca, A., Ianniti, R., Moretti, S., Bartoli, A., Aversa, F., Puccetti, P.: Microbiota control of a tryptophan-AhR pathway in disease tolerance to fungi: highlights. Eur J Immunol (2014). https://doi.org/10.1002/eji.201344406
Saijo, S., Ikeda, S., Yamabe, K., Kakuta, S., Ishigame, H., Akitsu, A., Fujikado, N., Kusaka, T., Kubo, S., Chung, S., Komatsu, R., Miura, N., Adachi, Y., Ohno, N., Shibuya, K., Yamamoto, N., Kawakami, K., Yamasaki, S., Saito, T., Akira, S., Iwakura, Y.: Dectin-2 recognition of alpha-mannans and induction of Th17 cell differentiation is essential for host defense against Candida albicans. Immunity 32, 681–691 (2010). https://doi.org/10.1016/j.immuni.2010.05.001
Sainz, J., Muñoz, C.B., Segura-Catena, J., Vázquez, L., Ríos Tamayo, R., Oyonarte, S., Hemminki, K., Försti, A., Jurado, M.: Dectin-1 and DC-SIGN polymorphisms associated with invasive pulmonary aspergillosis infection. PLoS ONE 7, e32273 (2012). https://doi.org/10.1371/journal.pone.0032273
Sam, Q.H., Chang, M., Chai, L.: The fungal mycobiome and its interaction with gut bacteria in the host. Int J Mol Sci 18, 330 (2017). https://doi.org/10.3390/ijms18020330
Santos, E., Levitz, S.M.: Fungal vaccines and immunotherapeutics. Cold Spring Harb Perspect Med 4, a019711 (2014)
Schei, K., Avershina, E., Øien, T., Rudi, K., Follestad, T., Salamati, S., Ødegård, R.A.: Early gut mycobiota and mother-offspring transfer. Microbiome 5, 107 (2017). https://doi.org/10.1186/s40168-017-0319-x
Schmidt, S., Tramsen, L., Lehrnbecher, T.: Natural killer cells in antifungal immunity. Front Immunol 8, 1623 (2017). https://doi.org/10.3389/fimmu.2017.01623
Sen, S., Mansell, T.J.: Yeasts as probiotics: mechanisms, outcomes, and future potential. Fungal Genet Biol 137, 103333 (2020). https://doi.org/10.1016/j.fgb.2020.103333
Short, D.P.G., O’Donnell, K., Thrane, U., Nielsen, K.F., Zhang, N., Juba, J.H., Geiser, D.M.: Phylogenetic relationships among members of the Fusarium solani species complex in human infections and the descriptions of F. keratoplasticum sp. Nov. and F. petroliphilum stat. nov. Fungal Genet Biol 53, 59–70 (2013). https://doi.org/10.1016/j.fgb.2013.01.004
Skevaki, C., Pararas, M., Kostelidou, K., Tsakris, A., Routsias, J.G.: Single nucleotide polymorphisms of Toll-like receptors and susceptibility to infectious diseases. Clin Exp Immunol 180, 165–177 (2015). https://doi.org/10.1111/cei.12578
Souza, A.C.O., Amaral, A.C.: Antifungal therapy for systemic mycosis and the nanobiotechnology era: improving efficacy. Biodistribution and toxicity. Front Microbiol 8, 336 (2017). https://doi.org/10.3389/fmicb.2017.00336
Sparber, F., De Gregorio, C., Steckholzer, S., Ferreira, F.M., Dolowschiak, T., Ruchti, F., Kirchner, F.R., Mertens, S., Prinz, I., Joller, N., Buch, T., Glatz, M., Sallusto, F., LeibundGut-Landmann, S.: The skin commensal yeast malassezia triggers a type 17 response that coordinates anti-fungal immunity and exacerbates skin inflammation. Cell Host Microbe 25, 389–403 (2019). https://doi.org/10.1016/j.chom.2019.02.002
Speakman, E.A., Dambuza, I.M., Salazar, F., Brown, G.D.: T cell antifungal immunity and the role of C-type lectin receptors. Trends Immunol 41, 61–76 (2020). https://doi.org/10.1016/j.it.2019.11.007
Spellberg, B.J., Ibrahim, A.S., Avanesian, V., Fu, Y., Myers, C., Phan, Q.T., Filler, S.G., Yeaman, M.R., Edwards, J.E., Jr.: Efficacy of the anti-candida rAls3p-N or rAls1p-N vaccines against disseminated and mucosal candidiasis. J Infect Dis 194, 256–260 (2006). https://doi.org/10.1086/504691
Spitzer, M., Robbins, N., Wright, G.D.: Combinatorial strategies for combating invasive fungal infections. Virulence 8, 169–185 (2017). https://doi.org/10.1080/21505594.2016.1196300
Subramanian Vignesh, K., Landero Figueroa, J.A., Porollo, A., Caruso, J.A., Deepe, G.S.: Granulocyte macrophage-colony stimulating factor induced Zn sequestration enhances macrophage superoxide and limits intracellular pathogen survival. Immunity 39, 697–710 (2013). https://doi.org/10.1016/j.immuni.2013.09.006
Tang, S., Chen, J., Cannon, J., Cao, Z., Baker, J.R., Jr., Wang, S.H.: Dendrimer-based posaconazole nanoplatform for antifungal therapy. Drug Deliv 28, 2150–2159 (2021). https://doi.org/10.1080/10717544.2021.1986605
Terciolo, C., Dobric, A., Ouaissi, M., Siret, C., Breuzard, G., Silvy, F., Marchiori, B., Germain, S., Bonier, R., Hama, A., Owens, R., Lombardo, D., Rigot, V., André, F.: Saccharomyces boulardii CNCM I-745 Restores intestinal barrier integrity by regulation of E-cadherin recycling. J Crohn’s Colitis 11, 999–1010 (2017). https://doi.org/10.1093/ecco-jcc/jjx030
Underhill, D.M., Iliev, I.D.: The mycobiota: interactions between commensal fungi and the host immune system. Nat Rev Immunol 14, 405–416 (2014). https://doi.org/10.1038/nri3684
van de Veerdonk, F.L., Netea, M.G.: T-cell subsets and antifungal host defenses. Curr Fungal Infect Rep 4, 238–243 (2010). https://doi.org/10.1007/s12281-010-0034-6
Verweij, P.E., Rijnders, B.J.A., Brüggemann, R.J.M., Azoulay, E., Bassetti, M., Blot, S., Calandra, T., Clancy, C.J., Cornely, O.A., Chiller, T., Depuydt, P., Giacobbe, D.R., Janssen, N.A.F., Kullberg, B.-J., Lagrou, K., Lass-Flörl, C., Lewis, R.E., Liu, P.W.-L., Lortholary, O., Maertens, J., Martin-Loeches, I., Nguyen, M.H., Patterson, T.F., Rogers, T.R., Schouten, J.A., Spriet, I., Vanderbeke, L., Wauters, J., van de Veerdonk, F.L.: Review of influenza-associated pulmonary aspergillosis in ICU patients and proposal for a case definition: an expert opinion. Intensive Care Med 46, 1524–1535 (2020). https://doi.org/10.1007/s00134-020-06091-6
Vivek-Ananth, R.P., Mohanraj, K., Vandanashree, M., Jhingran, A., Craig, J.P., Samal, A.: Comparative systems analysis of the secretome of the opportunistic pathogen Aspergillus fumigatus and other Aspergillus species. Sci Rep 8, 6617 (2018). https://doi.org/10.1038/s41598-018-25016-4
Ward, T.L., Knights, D., Gale, C.A.: Infant fungal communities: current knowledge and research opportunities. BMC Med 15, 30 (2017). https://doi.org/10.1186/s12916-017-0802-z
Ward, T.L., Dominguez-Bello, M.G., Heisel, T., Al-Ghalith, G., Knights, D., Gale, C.A.: Development of the human mycobiome over the first month of life and across body sites. mSystems 3, 40–17 (2018). https://doi.org/10.1128/mSystems.00140-17
Werner, J.L., Gessner, M.A., Lilly, L.M., Nelson, M.P., Metz, A.E., Horn, D., Dunaway, C.W., Deshane, J., Chaplin, D.D., Weaver, C.T., Brown, G.D., Steele, C.: Neutrophils produce interleukin 17A (IL-17A) in a Dectin-1- and IL-23-dependent manner during invasive fungal infection. Infect Immun 79, 3966–3977 (2011). https://doi.org/10.1128/IAI.05493-11
Wheeler, M.L., Limon, J.J., Underhill, D.M.: Immunity to commensal fungi: detente and disease. Annu Rev Pathol Mech Dis 12, 359–385 (2017). https://doi.org/10.1146/annurev-pathol-052016-100342
Whibley, N., MacCallum, D.M., Vickers, M.A., Zafreen, S., Waldmann, H., Hori, S., Gaffen, S.L., Gow, N.A.R., Barker, R.N., Hall, A.M.: Expansion of Foxp3+ T-cell populations by Candida albicans enhances both Th17-cell responses and fungal dissemination after intravenous challenge. Eur J Immunol 44, 1069–1083 (2014). https://doi.org/10.1002/eji.201343604
Wong, T.Y., Loo, Y.S., Veettil, S.K., Wong, P.S., Divya, G., Ching, S.M., Menon, R.K.: Efficacy and safety of posaconazole for the prevention of invasive fungal infections in immunocompromised patients: a systematic review with meta-analysis and trial sequential analysis. Sci Rep 10, 14575 (2020). https://doi.org/10.1038/s41598-020-71571-0
Zaragoza, O., Nielsen, K.: Titan cells in Cryptococcus neoformans: cells with a giant impact. Curr Opin Microbiol 16, 409–413 (2013). https://doi.org/10.1016/j.mib.2013.03.006
Zhao, L., Seth, A., Wibowo, N., Zhao, C.-X., Mitter, N., Yu, C., Middelberg, A.P.J.: Nanoparticle vaccines. Vaccine 32, 327–337 (2014). https://doi.org/10.1016/j.vaccine.2013.11.069
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The work has been performed with funding from Delhi Technological University to Dr. Asmita Das. No animal or human materials have been used, and all ethical norms of the university have been adhered to.
Rights and permissions
About this article
Cite this article
Sachdeva, G., Das, A. Communication between immune system and mycobiota impacts health and disease. Proc.Indian Natl. Sci. Acad. 88, 250–262 (2022). https://doi.org/10.1007/s43538-022-00082-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43538-022-00082-5