Abstract
Introduction
Acetaminophen toxicity has been associated with elevation of microRNAs. The present study was to evaluate overall microRNA profiles and previously identified microRNAs to differentiate acetaminophen (APAP) toxicity from other causes of transaminase elevation.
Methods
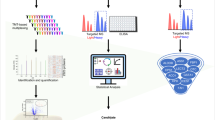
This was an observational study of adults with presumed acetaminophen toxicity at presentation. Serum samples were collected every 12 hours during hospitalization. Total miRNAs were extracted from plasma and levels of 327 microRNAs were quantified using real-time polymerase chain reaction. A standard measure of miRNA expression (delta-delta cycle threshold) was calculated for each microRNAs. A two-level cluster analysis was performed using a random k-means algorithm. Demographic and clinical characteristics of each cluster were compared using ANOVA, Wilcoxon rank sum, Kruskal-Wallis, and chi-square tests. Performance of specific miRNAs of interest was also evaluated.
Results
Twenty-seven subjects were enrolled (21 with a final diagnosis of acetaminophen toxicity), and a total of 61 samples were analyzed. Five clusters were identified, two of which demonstrated clear clinical patterns and included specific elevated miRNAs previously reported to be elevated in APAP toxicity patients. Features associated with clusters 1 and 5 included confirmed acetaminophen toxicity, high peak alanine aminotransferase, and late presentation. Clusters 2–4 contained lower peak microRNAs, lower peak alanine aminotransferase, and heterogeneous clinical characteristics.
Conclusions
Severe cases of acetaminophen toxicity showed two distinct patterns of microRNA elevation which were similar to previous work, while less severe cases were difficult to distinguish from non-acetaminophen-associated cases. Further work is needed to incorporate microRNA profiles into the diagnostic algorithm of acetaminophen toxicity.



Similar content being viewed by others
References
Budnitz DS, Lovegrove MC, Crosby AE. Emergency department visits for overdoses of acetaminophen-containing products. Am J Prev Med. 2011;40(6):585–92. https://doi.org/10.1016/j.amepre.2011.02.026.
Gummin DD, Mowry JB, Spyker DA, Brooks DE, Fraser MO, Banner W. 2016 Annual report of the American Association of Poison Control Centers’ National Poison Data System (NPDS): 34th annual report. Clin Toxicol (Philadelphia, Pa). 2017;55(10):1072–252. https://doi.org/10.1080/15563650.2017.1388087.
Amacher DE, Schomaker SJ, Aubrecht J. Development of blood biomarkers for drug-induced liver injury: an evaluation of their potential for risk assessment and diagnostics. Mol Diagn Ther. 2013;17(6):343–54. https://doi.org/10.1007/s40291-013-0049-0.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75(5):843–54.
Ward J, Szabo G, McManus D, Boyer E. Advanced molecular biologic techniques in toxicologic disease. J Med Toxicol. 2011;7(4):288–94. https://doi.org/10.1007/s13181-011-0189-8.
Osaki M, Kosaka N, Okada F, Ochiya T. Circulating microRNAs in drug safety assessment for hepatic and cardiovascular toxicity: the latest biomarker frontier? Mol Diagn Ther. 2014;18(2):121–6. https://doi.org/10.1007/s40291-013-0065-0.
Hammond SM. An overview of microRNAs. Adv Drug Deliv Rev. 2015;87:3–14. https://doi.org/10.1016/j.addr.2015.05.001.
Hydbring P, Badalian-Very G. Clinical applications of microRNAs. F1000Res. 2013;2:136. https://doi.org/10.12688/f1000research.2-136.v3.
Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16(3):203–22. https://doi.org/10.1038/nrd.2016.246.
Bala S, Petrasek J, Mundkur S, Catalano D, Levin I, Ward J, et al. Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology (Baltimore, Md). 2012;56(5):1946–57. https://doi.org/10.1002/hep.25873.
Ward J, Bala S, Petrasek J, Szabo G. Plasma microRNA profiles distinguish lethal injury in acetaminophen toxicity: a research study. World J Gastroenterol. 2012;18(22):2798–804. https://doi.org/10.3748/wjg.v18.i22.2798.
Su YW, Chen X, Jiang ZZ, Wang T, Wang C, Zhang Y, et al. A panel of serum microRNAs as specific biomarkers for diagnosis of compound- and herb-induced liver injury in rats. PLoS One. 2012;7(5):e37395. https://doi.org/10.1371/journal.pone.0037395.
Yang X, Greenhaw J, Shi Q, Su Z, Qian F, Davis K, et al. Identification of urinary microRNA profiles in rats that may diagnose hepatotoxicity. Toxicol Sci. 2012;125(2):335–44. https://doi.org/10.1093/toxsci/kfr321.
Fukushima T, Hamada Y, Yamada H, Horii I. Changes of micro-RNA expression in rat liver treated by acetaminophen or carbon tetrachloride--regulating role of micro-RNA for RNA expression. J Toxicol Sci. 2007;32(4):401–9.
Yu D, Wu L, Gill P, Tolleson WH, Chen S, Sun J, et al. Multiple microRNAs function as self-protective modules in acetaminophen-induced hepatotoxicity in humans. Arch Toxicol. 2018;92(2):845–58. https://doi.org/10.1007/s00204-017-2090-y.
Wang K, Zhang S, Marzolf B, Troisch P, Brightman A, Hu Z, et al. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc Natl Acad Sci U S A. 2009;106(11):4402–7. https://doi.org/10.1073/pnas.0813371106.
Krauskopf J, de Kok TM, Schomaker SJ, Gosink M, Burt DA, Chandler P, et al. Serum microRNA signatures as “liquid biopsies” for interrogating hepatotoxic mechanisms and liver pathogenesis in human. PLoS One. 2017;12(5):e0177928. https://doi.org/10.1371/journal.pone.0177928.
Wong A, Cheung B, Nejad C, Gantier M, Graudins A. Hepatotoxicity after paracetamol overdose in a patient with cystic fibrosis despite early acetylcysteine and utility of microRNA to predict hepatotoxicity. Clin Toxicol (Philadelphia, PA). 2018;56(10):904–6. https://doi.org/10.1080/15563650.2018.1454596.
Wong A, Nejad C, Gantier M, Choy KW, Doery J, Graudins A. MicroRNA from a 12-h versus 20-h acetylcysteine infusion for paracetamol overdose. Hum Exp Toxicol. 2019;38:646–54. https://doi.org/10.1177/0960327119833740.
Ward J, Kanchagar C, Veksler-Lublinsky I, Lee RC, McGill MR, Jaeschke H, et al. Circulating microRNA profiles in human patients with acetaminophen hepatotoxicity or ischemic hepatitis. Proc Natl Acad Sci U S A. 2014;111(33):12169–74. https://doi.org/10.1073/pnas.1412608111.
Starkey Lewis PJ, Dear J, Platt V, Simpson KJ, Craig DG, Antoine DJ, et al. Circulating microRNAs as potential markers of human drug-induced liver injury. Hepatology (Baltimore, Md). 2011;54(5):1767–76. https://doi.org/10.1002/hep.24538.
Antoine DJ, Dear JW, Lewis PS, Platt V, Coyle J, Masson M, et al. Mechanistic biomarkers provide early and sensitive detection of acetaminophen-induced acute liver injury at first presentation to hospital. Hepatology (Baltimore, Md). 2013;58(2):777–87. https://doi.org/10.1002/hep.26294.
Munakata C, Fuchigami Y, Hiroishi S, Haraguchi A, Hagimori M, Enomoto H, et al. Evaluation of miR-122 to predict high dose acetaminophen-induced liver injury in mice: the combination uses of 5-fluorouracil. Biol Pharm Bull. 2018;41(11):1732–5. https://doi.org/10.1248/bpb.b18-00504.
Harris PA, Taylor R, Thielke R, Payne J, Gonzalez N, Conde JG. Research electronic data capture (REDCap)--a metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inform. 2009;42(2):377–81. https://doi.org/10.1016/j.jbi.2008.08.010.
Liu P, Hwang JTG. Quick calculation for sample size while controlling false discovery rate with application to microarray analysis. Bioinformatics (Oxford, England). 2007;23(6):739–46. https://doi.org/10.1093/bioinformatics/btl664.
Sing T, Sander O, Beerenwinkel N, Lengauer T. ROCR: visualizing classifier performance in R. Bioinformatics (Oxford, England). 2005;21(20):3940–1. https://doi.org/10.1093/bioinformatics/bti623.
Vliegenthart ADB, Shaffer JM, Clarke JI, Peeters LEJ, Caporali A, Bateman DN, et al. Comprehensive microRNA profiling in acetaminophen toxicity identifies novel circulating biomarkers for human liver and kidney injury. Sci Rep. 2015;5:15501. https://doi.org/10.1038/srep15501 https://www.nature.com/articles/srep15501#supplementary-information.
Sources of Funding
This work was generously sponsored by an investigator-initiated grant from McNeil Consumer Healthcare. Specimen collection, storage, and processing was provided by the University of Massachusetts Medical School/UMass Memorial Healthcare biorepository (National Center for Advancing Translational Sciences/National Institutes of Health grant number UL1TR000161).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This protocol was reviewed and approved by the University of Massachusetts Medical School Institutional Review Board.
Conflicts of Interest
None.
Additional information
Supervising Editor: David M. Wood, MD
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(DOCX 139 kb)
Rights and permissions
About this article
Cite this article
Carreiro, S., Marvel-Coen, J., Lee, R. et al. Circulating microRNA Profiles in Acetaminophen Toxicity. J. Med. Toxicol. 16, 177–187 (2020). https://doi.org/10.1007/s13181-019-00739-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13181-019-00739-6