Abstract
The worldwide COVID-19 pandemic is caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Nonstructural protein 3 (nsp3) has 1945 residues and is the largest protein encoded by SARS-CoV-2. It comprises more than a dozen independent domains with various functions. Many of these domains were studied in the closely-related virus SARS-CoV following an earlier outbreak. Nonetheless structural and functional information on the C-terminal region of nsp3 containing two transmembrane and three extra-membrane domains remains incomplete. This part of the protein appears to be involved in initiation of double membrane vesicle (DMV) formation, membranous organelles the virus builds to hide its replication-transcription complex from host immune defenses. Here we present the near-complete backbone and Ile, Leu, and Val methyl group chemical shift assignments of the most C-terminal domain of nsp3, CoV-Y. As the exact function and binding partners of CoV-Y remain unknown, our data provide a basis for future NMR studies of protein–protein interactions to elucidate the molecular mechanism of DMV formation.
Similar content being viewed by others
Biological context
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has been responsible for the current worldwide COVID-19 pandemic since 2019. Following the outbreaks of SARS-CoV in 2002–2003 and the Middle East respiratory syndrome (MERS-CoV) in 2012 this is the third instance of a global coronavirus outbreak.
Like other ( +) RNA viruses SARS-CoV-2 forms specialized, membranous organelles in infected cells. These double membrane vesicles (DMVs) create a favorable microenvironment for the viral replication-transcription complex by concentrating viral components and at the same time shielding them from the host immune defense (Shulla and Randall 2016). All SARS-CoV, SARS-CoV-2 and MERS-CoV specifically target the Endoplasmic Reticulum (ER) membrane and rearrange it into interconnected spherical 200–300 nm DMVs (Wolff et al. 2020). It was shown that SARS-CoV non-structural protein 3 (nsp3) together with nsp4 and nsp6 plays a key role in DMV formation (Angelini et al. 2013; Hagemeijer et al. 2014). In a hexameric form, nsp3 forms a molecular pore that connects the DMV interior with the cytosol (Wolff et al. 2020). Nsp3 of SARS-CoV and SARS-CoV-2 have 75.8% sequence identity and share the same domain organization. This 222 kDa protein consists of more than a dozen domains that can be roughly divided in two parts. The first, nsp3N, consists of ten N-terminal cytosolic domains with different functions including a papain-like protease (Lei et al. 2018) and a scaffold region that participates in the replication-transcription complex assembly (Imbert et al. 2008). The second, nsp3C, has two transmembrane domains (TM1 and TM2) with a luminal loop (Ecto3) between them and two cytosolic domains (Y1 and CoV-Y) following TM2. Nsp3C anchors nsp3 to the ER membrane and induces membrane rearrangement (Angelini et al. 2013; Hagemeijer et al. 2014). However, the molecular mechanism of nsp3C-induced DMV formation remains unknown because of a lack of structural information for nsp3C domains.
The focus of this work is CoV-Y, the most C-terminal domain of SARS-CoV-2 nsp3. This domain is present in all coronaviruses. The level of its conservation is close to that for the enzymatic domains of nsp3 and exceeds non-enzymatic ones (Neuman 2016). However, the function of the nsp3 CoV-Y domain is unknown. Here, we present the resonance assignments of the backbone 1HN, 15N, 13C, 13Cα, 13Cβ and methyl 1H and 13C (Ile, Leu, Val) nuclei of SARS-CoV-2 nsp3 CoV-Y domain. These assignments will facilitate investigation of nsp3 CoV-Y function and interactions with other SARS-CoV-2 proteins.
Methods and experiments
Construct design
We previously described expression and purification of nsp3 CoV-Y (residues 1638–1945) (Altincekic et al. 2021). This construct provided good quality 1H-15N TROSY-HSQC spectra but a long N-terminal unstructured region (approximately 30 residues) complicated the assignment. In this work we used a shorter construct of nsp3 CoV-Y that corresponds to residues 1660–1945 based on the NCBI reference sequence YP_009742610.1. The gene sequence was E. coli codon optimized and cloned into a pET28b( +) vector containing a removable tobacco etch virus (TEV) protease recognition site. After TEV cleavage, the final construct contained an artificial N-terminal glycine residue preceding the native CoV-Y sequence.
Sample preparation
For backbone assignment we produced partially deuterated U-[15N-13C]- nsp3 CoV-Y protein by growing BL21 (DE3) E. coli cells transformed with the pET28b( +) plasmid encoding CoV-Y domain in M9 media using D2O with 1 mg/L U-[15N]-labeled ammonium chloride and 4 mg/L U-[13C]-labeled D-glucose as sole sources of nitrogen and carbon, respectively.
For methyl labeling of the nsp3 CoV-Y domain we used the procedure described by Tugarinov et al. (Tugarinov and Kay 2004; Tugarinov et al. 2005). A U-[15N,13C,2H], Ile δ1-[13CH3], Leu, Val-[13CH3,12CD3]-labeled sample of nsp3 CoV-Y (U-[15N,13C,2H]-ILV) was produced using D2O M9 media with 1 mg/L U-[15N]-labeled ammonium chloride and 4 mg/L U-[13C,2H]-labeled D-glucose. For selective methyl protonation 70 mg/L of [13C4; 3,3-2H2]-alpha-ketobuterate and 140 mg/L of [1,2,3,4-13C4; 3,4′,4′,4′-2H4]-alpha-ketoisovaleriate were added 1 hour prior to induction. All isotopes were purchased from Cambridge Isotope Laboratories, Inc.
The protein was purified as described previously (Altincekic et al. 2021) with two modifications: (I) we added 1 mM of tris (2-carboxyethyl) phosphine (TCEP) after TEV-cleavage of the His6-tag to reduce cysteine side chains; (II) we changed the final buffer because the computed pI was reduced from 7.1 to 6.6 for the new shorter construct. All NMR samples were prepared in 20 mM MOPS buffer pH 6.4, 100 mM LiBr, 2 mM DTT and 0.01% NaN3.
NMR experiments
All NMR experiments were recorded at 25 °C on a Varian Inova 800 MHz spectrometer equipped with a cryogenic triple-resonance probe. For the backbone resonance assignments a sample of 0.35 mM partially deuterated U-[15N-13C]- nsp3 CoV-Y in NMR buffer with 10% D2O was used. A set of NMR experiments for backbone 15N, 13C and 1H resonance assignment consisted of 2D 1H-15N TROSY-HSQC and three-dimensional non-uniformly sampled TROSY-based HNCO (15%), HNCACO (15%), HNCA (16%), HNCACB (14%) (Yang and Kay 1999) and 15N-edited NOESY-HSQC (20%) with 140 ms mixing time (Zhang et al. 1994). For the assignment of Ile δ1, Leu δ1,2 and Val γ1,2 methyl groups a 0.25 mM U-[15N,13C,2H]-ILV sample was used. We recorded 2D 1H-13C HSQC and 3D HMCMCGCBCA (Tugarinov and Kay 2003) acquired with 20% NUS sampling coverage.
All spectra were processed using NMRpipe (Delaglio et al. 1995) and SMILE for NUS spectral reconstruction (Ying et al. 2017).
Assignment and data deposition
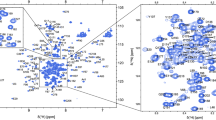
Analysis of the spectra and backbone resonance assignment were performed manually using CARA (Keller 2004). The 1H-15N TROSY-HSQC spectrum of nsp3 CoV-Y domain shows excellent signal dispersion for a 286-residue protein (Fig. 1). Backbone assignment is nearly complete (97% HN, 94% N, 98% C′, 98% Cα, 90% Cβ). There are nine missing residue assignments, K1660, N1680, N1854, T1858, Y1859, N1934-T1937 in the 1H-15N TROSY-HSQC spectrum. The HN peaks for missing residues are probably broadened beyond detection in the 1H-15N TROSY-HSQC. A few unassigned low intensity peaks in 1H-15N TROSY-HSQC spectrum don’t provide identifiable cross-peaks in 3D spectra for reliable correlation.
Assigned 1H-15N TROSY-HSQC spectrum of partially deuterated U-[15N-13C]- nsp3 CoV-Y domain at 0.35 mM concentration in 20 mM MOPS buffer pH 6.4, 100 mM LiBr, 2 mM DTT, 0.01% NaN3 and 10% D2O measured at 25 °C on a Varian Inova 800 MHz NMR spectrometer. The inset shows an expansion of the central region of the spectrum
Measured backbone chemical shifts were used as input for Talos + (Shen and Bax 2015) to calculate the random-coil-index-derived order parameters (RCI-S2) and secondary structure propensities (Fig. 2). Our results suggest that the CoV-Y domain possesses a well-ordered globular α/β fold with flexible termini (eight N-terminal residues and four C-terminal residues). These terminal residues as well as residues 1932–1936 have RCI-S2 < 0.65 (Fig. 2). This probably explains why peaks corresponding to N1934-T1937 are broadened beyond detection in the 1H-15N TROSY-HSQC spectrum.
Methyl groups assignment and figure preparation were done with CCPN Analysis (Vranken et al. 2005). All 14 Ile δ1 and 25 Leu δ1,2 groups were assigned as well as 24 out of 28 Val γ1,2 (Fig. 3). Due to lack of backbone assignment for V1935 and V1936, partial assignment of V1933, and overlap in spectra, the methyl groups of V1777, V1933, V1935 and V1936 were not assigned.
Assigned 1H-13C HSQC spectrum of U-[15N,13C,2H], Ile δ1-[13CH3], Leu,Val-[13CH3,12CD3]-labeled nsp3 CoV-Y domain at 0.25 mM concentration in 20 mM MOPS buffer pH 6.4, 100 mM LiBr, 2 mM DTT, 0.01% NaN3 and 10% D2O measured at 25 °C on a Varian Inova 800 MHz NMR spectrometer. For clarity methyl groups are shown with different colors: Ile δ1, dark green; Leu δ1,2, dark red; Val γ1,2, blue
All obtained backbone and methyl chemical shifts of SARS-CoV-2 Nsp3 CoV-Y have been deposited at the BMRB under accession number 51074.
Our results provide a foundation for future NMR studies of nsp3 CoV-Y domain to identify interaction partners and to understand the molecular mechanism of DMV formation.
Data availability
The values for 1H, 13C and 15N backbone chemical shifts and 1H, 13C for Ile δ1, Leu δ1,2 and Val γ1,2 methyl groups of SARS-CoV-2 Nsp3 CoV-Y have been deposited at the Biological Magnetic Resonance Data Bank (BMRB) (https://www.bmrb.wisc.edu) under Accession Number 51074.
References
Altincekic N, Korn SM, Qureshi NS et al (2021) Large-scale recombinant production of the SARS-CoV-2 proteome for high-throughput and structural biology applications. Front Mol Biosci 8:1–25. https://doi.org/10.3389/fmolb.2021.653148
Angelini MM, Akhlaghpour M, Neuman BW, Buchmeier MJ (2013) Severe acute respiratory syndrome coronavirus nonstructural proteins 3, 4, and 6 induce double-membrane vesicles. Mbio 4:1–10. https://doi.org/10.1128/mBio.00524-13
Delaglio F, Grzesiek S, Vuister GW et al (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Hagemeijer MC, Monastyrska I, Griffith J et al (2014) Membrane rearrangements mediated by coronavirus nonstructural proteins 3 and 4. Virology 458–459:125–135. https://doi.org/10.1016/j.virol.2014.04.027
Imbert I, Snijder EJ, Dimitrova M et al (2008) The SARS-coronavirus PLnc domain of nsp3 as a replication/transcription scaffolding protein. Virus Res 133:136–148. https://doi.org/10.1016/j.virusres.2007.11.017
Keller RLJ (2004) The computer aided resonance assignment tutorial. CANTINA Verlag, Goldau, Switzerland
Lei J, Kusov Y, Hilgenfeld R (2018) Nsp3 of coronaviruses: structures and functions of a large multi-domain protein. Antiviral Res 149:58–74. https://doi.org/10.1016/j.antiviral.2017.11.001
Neuman BW (2016) Bioinformatics and functional analyses of coronavirus nonstructural proteins involved in the formation of replicative organelles. Antiviral Res 135:97–107. https://doi.org/10.1016/j.antiviral.2016.10.005
Shen Y, Bax A (2015) Protein structural information derived from nmr chemical shift with the neural network program talos-n. Methods Mol Biol 1260:17–32. https://doi.org/10.1007/978-1-4939-2239-0_2
Shulla A, Randall G (2016) (+) RNA virus replication compartments: a safe home for (most) viral replication. Curr Opin Microbiol 32:82–88. https://doi.org/10.1016/j.mib.2016.05.003
Tugarinov V, Kay LE (2003) Ile, Leu, and Val Methyl assignments of the 723-residue malate synthase G using a new labeling strategy and novel NMR methods. J Am Chem Soc 125:13868–13878. https://doi.org/10.1021/ja030345s
Tugarinov V, Kay LE (2004) An isotope labeling strategy for methyl TROSY spectroscopy. J Biomol NMR 28:165–172. https://doi.org/10.1023/B:JNMR.0000013824.93994.1f
Tugarinov V, Kay LE, Ibraghimov I, Orekhov VY (2005) High-resolution four-dimensional 1H–13C NOE spectroscopy using methyl-TROSY, sparse data acquisition, and multidimensional decomposition. J Am Chem Soc 127:2767–2775. https://doi.org/10.1021/ja044032o
Vranken WF, Boucher W, Stevens TJ et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct Funct Genet 59:687–696. https://doi.org/10.1002/prot.20449
Wolff G, Limpens RWAL, Zevenhoven-Dobbe JC et al (2020) A molecular pore spans the double membrane of the coronavirus replication organelle. Science 369:1395–1398. https://doi.org/10.1101/2020.06.25.171686
Yang D, Kay LE (1999) Improved lineshape and sensitivity in the HNCO-family of triple resonance experiments. J Biomol NMR 14:273–276. https://doi.org/10.1023/A:1008381929212
Ying J, Delaglio F, Torchia DA, Bax A (2017) Sparse multidimensional iterative lineshape-enhanced (SMILE) reconstruction of both non-uniformly sampled and conventional NMR data. J Biomol NMR 68:101–118. https://doi.org/10.1007/s10858-016-0072-7
Zhang O, Kay LE, Olivier JP (1994) Backbone 1H and 15N resonance assignments of the N-terminal SH3 domain of drk in folded and unfolded states using enhanced-sensitivity pulsed field gradient NMR techniques. J Biomol NMR 4:845–858
Acknowledgements
This work is a contribution from the international covid19-nmr consortium (https://covid19-nmr.de)
Funding
This work was supported by the US National Science Foundation via Grant RAPID 2030601, and by the US National Institutes of Health Grants R01GM123249 and P41GM111135.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pustovalova, Y., Gorbatyuk, O., Li, Y. et al. Backbone and Ile, Leu, Val methyl group resonance assignment of CoV-Y domain of SARS-CoV-2 non-structural protein 3. Biomol NMR Assign 16, 57–62 (2022). https://doi.org/10.1007/s12104-021-10059-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-021-10059-y