Abstract
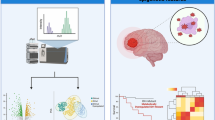
Glioblastoma multiforme (GBM) is one of the most lethal malignancies of the central nervous system characterized by high mortality rate. The complexity of GBM pathogenesis, progression, and prognosis is not fully understood yet. GBM-derived extracellular vesicles (EVs) carry several oncogenic elements that facilitate GBM progression. The purpose of this study was to identify systems level molecular signatures from GBM-derived EVs using integrative analysis of publicly available transcriptomic data generated from plasma and serum samples. The dataset contained 19 samples in total, of which 15 samples were from plasma (11 GBM patients and 4 healthy samples) and 4 samples were from serum (2 GBM and 2 healthy samples). We carried out statistical analysis to identify differentially expressed genes (DEGs), functional enrichment analysis of the DEGs, protein–protein interaction networks, module analysis, transcription factors and target gene regulatory networks analysis, and identification of hub genes. The differential expression of the identified hub genes were validated with the independent TCGA-GBM dataset. We have identified a few crucial genes and pathways associated with GBM prognosis and therapy resistance. The DEGs identified from plasma were associated with inflammatory processes and viral infection. On the other hand, the hub genes identified from the serum samples were significantly associated with protein ubiquitinylation processes and cytokine signaling regulation. The findings indicate that GBM-derived plasma and serum DEGs may be associated with distinct cellular processes and pathways which facilitate GBM progression. The findings will provide better understanding of the molecular mechanisms of GBM pathogenesis and progression. These results can further be utilized for developing and validating minimally invasive diagnostic and therapeutic molecular biomarkers for GBM.




Similar content being viewed by others
Abbreviations
- GBM:
-
Glioblastoma multiforme
- EVs:
-
Extracellular vesicles
- DEGs:
-
Differentially expressed genes
- PPIs:
-
Protein–protein interactions
- GO:
-
Gene ontology
- TFs:
-
Transcription factors
- OS:
-
Overall survival
References
Abdolmaleki M, Sohrabi A (2018) Characterization of JAK2 V617F (1849 G > T) mutation in cervical cancer related to human papillomavirus and sexually transmitted infections. J Cancer Prev 23:82–86. https://doi.org/10.15430/jcp.2018.23.2.82
Akhtar S, Vranic S, Cyprian FS, Al MAE (2018) Epstein-barr virus in gliomas: Cause, association, or artifact? Front Oncol 8:1–10. https://doi.org/10.3389/fonc.2018.00123
Alibek K, Kakpenova A, Baiken Y (2013) Role of infectious agents in the carcinogenesis of brain and head and neck cancers. Infect Agent Cancer 8:1. https://doi.org/10.1186/1750-9378-8-7
Baker BJ, Akhtar LN, Benveniste EN (2009) SOCS1 and SOCS3 in the control of CNS immunity. Trends Immunol 30:392–400. https://doi.org/10.1016/j.it.2009.07.001
Basu B, Ghosh MK (2019) Extracellular vesicles in glioma: From diagnosis to therapy. BioEssays 41:1–9. https://doi.org/10.1002/bies.201800245
Chen A, Zhong L, Lv J (2019) FOXL1 overexpression is associated with poor outcome in patients with glioma. OncolLett 18:751–757. https://doi.org/10.3892/ol.2019.10351
Chen J, Zhang C, Mi Y et al (2017) CREB1 regulates glucose transport of glioma cell line U87 by targeting GLUT1. Mol Cell Biochem 436:79–86. https://doi.org/10.1007/s11010-017-3080-3
Chin CH, Chen SH, Wu HH et al (2014) cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC SystBiol 8:S11. https://doi.org/10.1186/1752-0509-8-S4-S11
Cobbs CS, Harkins L, Samanta M et al (2002) Human cytomegalovirus infection and expression in human malignant glioma. Cancer Res 62:3347–3350
Dennis G, Sherman BT, Hosack DA, et al (2003) DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol 4:. https://doi.org/10.1186/gb-2003-4-9-r60
Duan H, Hu JL, Chen ZH et al (2020) Assessment of circulating tumor DNA in cerebrospinal fluid by whole exome sequencing to detect genomic alterations of glioblastoma. Chin Med J (Engl) 133:1415–1421. https://doi.org/10.1097/CM9.0000000000000843
Fornara O, Bartek J, Rahbar A et al (2016) Cytomegalovirus infection induces a stem cell phenotype in human primary glioblastoma cells: Prognostic significance and biological impact. Cell Death Differ 23:261–269. https://doi.org/10.1038/cdd.2015.91
Foster H, Cobbs CS (2019) Viruses and Glioblastoma: Affliction or Opportunity? 67–86. https://doi.org/https://doi.org/10.1007/978-3-030-04155-7_4
Goldszmid RS, Dzutsev A, Trinchieri G (2014) Host immune response to infection and cancer: Unexpected commonalities. Cell Host Microbe 15:295–305. https://doi.org/10.1016/j.chom.2014.02.003
Gonzalez H, Hagerling C, Werb Z (2018) Roles of the immune system in cancer: From tumor initiation to metastatic progression. Genes Dev 32:1267–1284. https://doi.org/10.1101/GAD.314617.118
Hoeller D, Hecker CM, Dikic I (2006) Ubiquitin and ubiquitin-like proteins in cancer pathogenesis. Nat Rev Cancer 6:776–788. https://doi.org/10.1038/nrc1994
Hu L, Li X, Liu Q et al (2017) UBE2S, a novel substrate of Akt1, associates with Ku70 and regulates DNA repair and glioblastomamultiforme resistance to chemotherapy. Oncogene 36:1145–1156. https://doi.org/10.1038/onc.2016.281
Kim D, Langmead B, Salzberg SL (2015) HISAT: A fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
Kinker GS, Thomas AM, Carvalho VJ et al (2016) Deletion and low expression of NFKBIA are associated with poor prognosis in lower-grade glioma patients. Sci Rep 6:1–9. https://doi.org/10.1038/srep24160
Kundu JK, Surh YJ (2008) Inflammation: Gearing the journey to cancer. Mutat Res - Rev Mutat Res 659:15–30. https://doi.org/10.1016/j.mrrev.2008.03.002
Lane R, Simon T, Vintu M, et al (2019) Cell-derived extracellular vesicles can be used as a biomarker reservoir for glioblastoma tumor subtyping. Commun Biol 2:. https://doi.org/10.1038/s42003-019-0560-x
Lee DH, Chung K, Song JA et al (2010) Proteomic identification of paclitaxel-resistance associated hnRNP A2 and GDI 2 proteins in human ovarian cancer cells. J Proteome Res 9:5668–5676. https://doi.org/10.1021/pr100478u
Leone R, Giussani P, De Palma S et al (2015) Proteomic analysis of human glioblastoma cell lines differently resistant to a nitric oxide releasing agent. MolBiosyst 11:1612–1621. https://doi.org/10.1039/c4mb00725e
Liggett (2014) 基因的改变NIH Public Access. Bone 23:1–7. https://doi.org/10.1038/jid.2014.371
Lombardi MY, Assem M (2017) Glioblastoma Genomics: A Very Complicated Story. Glioblastoma 3–25. https://doi.org/10.15586/codon.glioblastoma.2017.ch1
Love MI, Anders S, Huber W (2014) Differential analysis of count data - the DESeq2 package
Mani A, Gelmann EP (2005) The ubiquitin-proteasome pathway and its role in cancer. J ClinOncol 23:4776–4789. https://doi.org/10.1200/JCO.2005.05.081
Mantamadiotis T, Papalexis N, Dworkin S (2012) CREB signalling in neural stem/progenitor cells: Recent developments and the implications for brain tumour biology. BioEssays 34:293–300. https://doi.org/10.1002/bies.201100133
McFaline-Figueroa JR, Wen PY (2017) The viral connection to glioblastoma. Curr Infect Dis Rep 19:2–6. https://doi.org/10.1007/s11908-017-0563-z
Noreen M, Arshad M (2015) Association of TLR1, TLR2, TLR4, TLR6, and TIRAP polymorphisms with disease susceptibility. Immunol Res 62:234–252. https://doi.org/10.1007/s12026-015-8640-6
Ohgaki H, Kleihues P (2007) Genetic pathways to primary and secondary glioblastoma. Am J Pathol 170:1445–1453. https://doi.org/10.2353/ajpath.2007.070011
Onda M, Emi M, Yoshida A et al (2004) Comprehensive gene expression profiling of anaplastic thyroid cancers with cDNA microarray of 25 344 genes. EndocrRelat Cancer 11:843–854. https://doi.org/10.1677/erc.1.00818
Osti D, Del BM, Rappa G et al (2019) Clinical significance of extracellular vesicles in plasma from glioblastoma patients. Clin Cancer Res 25:266–276. https://doi.org/10.1158/1078-0432.CCR-18-1941
Paul Shannon 1, Andrew Markiel 1, Owen Ozier, 2 Nitin S. Baliga, 1 Jonathan T. Wang, 2 Daniel Ramage 2, et al (1971) Cytoscape: A Software Environment for Integrated Models. Genome Res 13:426. https://doi.org/10.1101/gr.1239303.metabolite
Pruitt KD, Hogue CW, Groll M et al (2001) Mcode. Nucleic Acids Res 29:137–140. https://doi.org/10.1093/nar/29.1.137
Puliyappadamba VT, Hatanpaa KJ, Chakraborty S, Habib AA (2014) The role of NF-κB in the pathogenesis of glioma. Mol Cell Oncol 1:. https://doi.org/10.4161/23723548.2014.963478
Raposo G, Stoorvogel W (2013) Extracellular vesicles exosomesmicrovesicles and friends. J Cell Biol 200:373–383. https://doi.org/10.1083/jcb.201211138
Ray SK, Patel SJ, Welsh CT et al (2002) Molecular evidence of apoptotic death in malignant brain tumors including glioblastomamultiforme: Upregulation of calpain and caspase-3. J Neurosci Res 69:197–206. https://doi.org/10.1002/jnr.10265
Reátegui E, Van Der Vos KE, Lai CP, et al (2018) Engineered nanointerfaces for microfluidic isolation and molecular profiling of tumor-specific extracellular vesicles. Nat Commun 9:. https://doi.org/10.1038/s41467-017-02261-1
Ryu JI, Han MH, Cheong JH et al (2017) Expression of the growth factor granulin in patients with brain tumors: Relevance to prognosis. Biomed Res 28:5583–5588
Santiago-Dieppa DR, Steinberg J, Gonda D et al (2014) Extracellular vesicles as a platform for “liquid biopsy” in glioblastoma patients. Expert Rev MolDiagn 14:819–825. https://doi.org/10.1586/14737159.2014.943193
Saugstad JA, Lusardi TA, Van Keuren-Jensen KR, et al (2017) Analysis of extracellular RNA in cerebrospinal fluid. J Extracell Vesicles 6:. https://doi.org/10.1080/20013078.2017.1317577
Schleifer RJ, Li S, Nechtman W et al (2018) KLHL5 knockdown increases cellular sensitivity to anticancer drugs. Oncotarget 9:37429–37438. https://doi.org/10.18632/oncotarget.26462
Sigi R, Wandkee C, Rauch V et al (2009) Loss of the mammalian APC/C activator FZR1 shortens G1 and lengthens S phase but has little effect on exit from mitosis. J Cell Sci 122:4208–4217. https://doi.org/10.1242/jcs.054197
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613. https://doi.org/10.1093/nar/gky1131
Tan X, Wang S, Zhu L et al (2012) cAMP response element-binding protein promotes gliomagenesis by modulating the expression of oncogenic microRNA-23a. ProcNatlAcadSci U S A 109:15805–15810. https://doi.org/10.1073/pnas.1207787109
Tang Z, Li C, Kang B et al (2017) GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45:W98–W102. https://doi.org/10.1093/nar/gkx247
Vachher M, Arora K, Burman A, Kumar B (2020) NAMPT, GRN, and SERPINE1 signature as predictor of disease progression and survival in gliomas. J Cell Biochem 121:3010–3023. https://doi.org/10.1002/jcb.29560
Ventero MP, Fuentes-Baile M, Quereda C et al (2019) Radiotherapy resistance acquisition in glioblastoma. Role of SOCS1 and SOCS3. PLoS ONE 14:1–21. https://doi.org/10.1371/journal.pone.0212581
Wan L, Chen M, Cao J et al (2017) The APC/C E3 ligase complex activator fzr1 restricts braf oncogenic function. Cancer Discov 7:424–441. https://doi.org/10.1158/2159-8290.CD-16-0647
Whitehead CA, Kaye AH, Drummond KJ et al (2020) Extracellular vesicles and their role in glioblastoma. Crit Rev Clin Lab Sci 57:227–252. https://doi.org/10.1080/10408363.2019.1700208
Xu J, Gu S, Wang S et al (2003) Characterization of a novel splicing variant of KLHL5*, a member of the kelch protein family. MolBiol Rep 30:239–242. https://doi.org/10.1023/A:1026372901766
Xu R, Rai A, Chen M et al (2018) Extracellular vesicles in cancer — implications for future improvements in cancer care. Nat Rev ClinOncol 15:617–638. https://doi.org/10.1038/s41571-018-0036-9
Yamabe K, Shimizu S, Ito T et al (1999) Cancer gene therapy using a pro-apoptotic gene, caspase-3. Gene Ther 6:1952–1959. https://doi.org/10.1038/sj.gt.3301041
Yamasaki M, Takemasa I, Komori T et al (2007) The gene expression profile represents the molecular nature of liver metastasis in colorectal cancer. Int J Oncol 30:129–138. https://doi.org/10.3892/ijo.30.1.129
Yang Z, Jiang S, Cheng Y et al (2017) FOXC1 in cancer development and therapy: Deciphering its emerging and divergent roles. TherAdv Med Oncol 9:797–816. https://doi.org/10.1177/1758834017742576
Zhou G, Soufan O, Ewald J et al (2019) NetworkAnalyst 3.0: A visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res 47:W234–W241. https://doi.org/10.1093/nar/gkz240
Zhou H, Miki R, Eeva M et al (2007) Reciprocal regulation of SOCS1 and SOCS3 enhances resistance to ionizing radiation in glioblastomamultiforme. Clin Cancer Res 13:2344–2353. https://doi.org/10.1158/1078-0432.CCR-06-2303
Acknowledgments
NR acknowledges Institutional Fellowship from Tezpur University for pursuing her PhD. PB acknowledges the infrastructural support provided from the Department of Molecular Biology and Biotechnology at the Tezpur University.
Funding
This work was supported by the Ramalingaswami Re-entry Fellowship Grant to Dr. Pankaj Barah by the Department of Biotechnology, Government of India.
Author information
Authors and Affiliations
Contributions
PB developed the concept. NR and MG analyzed the data. NR, MG, DKB, and PB interpreted the results and written the manuscript. NR and MG contributed equally to this work.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Publicly available gene expression data has been used for this analysis.
Consent for publication
All authors have seen and approved the manuscript for submission.
Competing interests
The authors of this manuscript declare no competing interest.
Availability of data and materials
The dataset used for this study is available at NCBI-GEO database (GEO Accession No. GSE106804).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Roy, N., Gaikwad, M., Bhattacharrya, D.K. et al. Identification of Systems Level Molecular Signatures from Glioblastoma Multiforme Derived Extracellular Vesicles. J Mol Neurosci 71, 1156–1167 (2021). https://doi.org/10.1007/s12031-020-01738-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-020-01738-x