Abstract
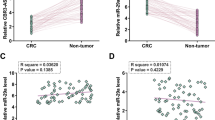
The biologic function as well as the mechanism of long noncoding RNAs (lncRNAs) in colorectal cancer (CRC) still remain largely unknown. Long noncoding RNA FGD5 antisense RNA 1 (FGD5-AS1) has been reported to have a promotive effect on other human cancers, but its function in CRC still remains unknown. The expression levels of long noncoding RNA FGD5-AS1, CDCA7 mRNA, and miR-302e were assessed by RT-qPCR. The protein levels of CDCA7 were assessed by Western blot. The function of FGD5-AS1 was detected using cell viability assay, 5-ethynyl-2′-deoxyuridine (EdU) assay, transwell, and caspase-3 activity assay. Additionally, the microRNAs (miRNAs) sponge potential of FGD5-AS1 was examined by RNA immunoprecipitation assay, RNA pull-down assay, and luciferase reporter assay. FGD5-AS1 was increased in colorectal cancer cell lines compared to normal cell lines. Inhibition of FGD5-AS1 suppressed cell proliferation, migration, invasion, and accelerated cell apoptosis in CRC. FGD5-AS1 competitively bound with miR-302e to modulate CDCA7. The inhibiting effects of FGD5-AS1 knockdown on CRC cell proliferation, migration, and invasion, and the promoting effects on CRC cell apoptosis could be revived by miR-302e suppression or CDCA7 upregulation. LncRNA FGD5-AS1 could promote CRC progression through sponging miR-302e and upregulating CDCA7. FGD5-AS1 might serve as a potential therapeutic target for CRC.




Similar content being viewed by others
References
Akhade VS, Pal D, Kanduri C (2017) Long noncoding RNA: genome organization and mechanism of action. Adv Exp Med Biol 1008:47–74
Chen T, Xie W, Xie L, Sun Y, Zhang Y, Shen Z, Sha N, Xu H, Wu Z, Hu H, Wu C (2015) Expression of long noncoding RNA lncRNA-n336928 is correlated with tumor stage and grade and overall survival in bladder cancer. Biochem Biophys Res Commun 468:666–670
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J (2016b) Cancer statistics in China, 2015. CA Cancer J Clin 66:115–132
Chen X, Xu Y, Liao X, Liao R, Zhang L, Niu K, Li T, Li D, Chen Z, Duan Y, Sun J (2016a) Plasma miRNAs in predicting radiosensitivity in non-small cell lung cancer. Tumour Biol 37:11927–11936
Chen ZP, Wei JC, Wang Q, Yang P, Li WL, He F, Chen HC, Hu H, Zhong JB, Cao J (2018) Long noncoding RNA 00152 functions as a competing endogenous RNA to regulate NRP1 expression by sponging with miRNA206 in colorectal cancer. Int J Oncol 53:1227–1236
Dai Q, Wei HL, Huang J, Zhou TJ, Chai L, Yang ZH (2016) KRAS polymorphisms are associated with survival of CRC in Chinese population. Tumour Biol: J Int Soc Oncodev Biol Med 37:4727–4734
Feng H, Zhang J, Shi Y, Wang L, Zhang C, Wu L (2018) Long noncoding RNA LINC-PINT is inhibited in gastric cancer and predicts poor survival. J Cell Biochem 120:9594–9600. https://doi.org/10.1002/jcb.28236
Guiu J, Bergen DJ, De Pater E, Islam AB, Ayllón V, Gama-Norton L, Ruiz-Herguido C, González J, López-Bigas N, Menendez P, Dzierzak E, Espinosa L, Bigas A (2014) Identification of Cdca7 as a novel notch transcriptional target involved in hematopoietic stem cell emergence. J Exp Med 211:2411–2423
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, Li R, West RB, van de Viiver MJ, Sukumar S, Chang HY (2010) Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464:1071–1076
Jimenez-P R, Martin-Cortazar C, Kourani O, Choido Y, Cordova R, Franjo MPD, Redondo JM, Iglesias T, Campanero MR (2018) CDCA7 is a critical mediator of lymphomagenesis that selectively regulates anchorage-independent growth. Haematologica 103:1669–1678
Jing H, Qu X, Liu L, Xia H (2018) A novel long noncoding RNA (lncRNA), LL22NC03-N64E9.1, promotes the proliferation of lung cancer cells and is a potential prognostic molecular biomarker for lung cancer. Med Sci Monit 24:4317–4323
Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, Bell I, Cheung E, Drenkow J, Dumais E, Patel S, Helt G, Ganesh M, Ghosh S, Piccolboni A, Semenchenko V, Tammana H, Gingeras TR(2007) RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science (New York, NY) 316:1484-1488
Kong Q, Qiu M (2018) Long noncoding RNA SNHG15 promotes human breast cancer proliferation, migration and invasion by sponging miR-211-3p. Biochem Biophys Res Commun 495:1594–1600
Li S, Liu X, Li H, Pan H, Acharya A, Deng Y, Yu Y, Haak R, Schmidt J, Schmalz G, Ziebolz D (2018) Integrated analysis of long noncoding RNA-associated competing endogenous RNA network in periodontitis. J Periodontal Res 53:495–505. https://doi.org/10.1111/jre.12539
Liu B, Pan S, Xiao Y, Liu Q, Xu J, Jia L (2018) LINC01296/miR-26a/GALNT3 axis contributes to colorectal cancer progression by regulating O-glycosylated MUC1 via PI3K/AKT pathway. J Exp Clin Cancer Res 37:316
Liu XH, Sun M, Nie FQ, Ge YB, Zhang EB, Yin DD, Kong R, Xia R, Lu KH, Li JH, De W, Wang KM, Wang ZX (2014) Lnc RNA HOTAIR functions as a competing endogenous RNA to regulate HER2 expression by sponging miR-331-3p in gastric cancer. Mol Cancer 13:92
Osthus RC, Karim B, Prescott JE, Smith BD, McDevitt M, Huso DL, Dang CV (2005) The Myc target gene JPO1/CDCA7 is frequently overexpressed in human tumors and has limited transforming activity in vivo. Cancer Res 65:5620–5627
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146:353–358
Siegle RL, Miller KD, Jemal A (2017a) Cancer statistics, 2017. CA Cancer J Clin 67:7–30
Siegle RL, Miller KD, Fedewasa SA, Ahnen DJ, Meester RGS, Barzi A, Jemal A (2017b) Colorectal cancer statistics, 2017. CA Cancer J Clin 67:177–193
Tay Y, Aqarwal V, Stefano J, Bartel DP, Stoffel M (2014b) Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol Cell 54:766–776
Tay Y, Rinn J, Pandolfi PP (2014a) The multilayered complexity of ceRNA crosstalk and competition. Nature 505:344–352
Wang KC, Chang HY (2011) Molecular mechanisms of long noncoding RNAs. Mol Cell 43:904–914
Wang W, Kandimalla R, Huang H, Zhu L, Li Y, Gao F, Goel A, Wang X (2018) Molecular subtyping of colorectal cancer: recent progress, new challenges and emerging opportunities. Semin Cancer Biol 55:37–52. https://doi.org/10.1016/j.semcancer.2018.05.002
Xiao L, Jiang L, Hu Q, Li Y (2018) MiR-302e attenuates allergic inflammation in vitro model by targeting RelA. Biosci Rep 38:BSR20180025. https://doi.org/10.1042/bsr20180025
Xu J, Zang R, Zhao J (2017) The novel long noncoding RNA TUSC7 inhibits proliferation by sponging MiR-211 in colorectal cancer. Cell Physiol Biochem : Int J Exp Cell Physiol, Biochem, Pharmacol 41:635–644
Yan ZY, Sun XC (2018) LincRNA-ROR functions as a ceRNA to regulate Oct4, Sox2, and Nanog expression by sponging miR-145 and its effect on biologic characteristics of colonic cancer stem cells. Chinese Journal of Pathology 47:284–290
Yang Q, Jia L, Li X, Guo R, Huang Y, Zheng Y, Li W (2018) Long noncoding RNAs: new players in the osteogenic differentiation of bone marrow- and adipose-derived mesenchymal stem cells. Stem Cell Rev 14:297–308
Ye L, Li F, Song Y, Yu D, Xiong Z, Li Y, Shi T, Yuan Z, Lin C, Wu X, Ren L, Li X, Song L (2018) Overexpression of CDCA7 predicts poor prognosis and induces EZH2-mediated progression of triple-negative breast cancer. Int J Cancer 143:2602–2613
Yu Y, Zhang X, Tian H, Zhang Z, Tian Y (2018) Knockdown of long non-coding RNA HOTAIR increases cisplatin sensitivity in ovarian cancer by inhibiting cisplatin-induced autophagy. J buon 23:1396–1401
Zhang G, He X, Ren C, Lin J, Wang Q (2018) Long noncoding RNA PCA3 regulates prostate cancer through sponging miR-218-5p and modulating high mobility group box 1. J Cell Physiol 234:13097–13109. https://doi.org/10.1002/jcp.27980
Zhu H, Lu J, Zhao H, Chen Z, Cui Q, Lin Z, Wang X, Wang J, Dong H, Wang S, Tan J (2018) Functional long noncoding RNAs (lncRNAs) in clear cell kidney carcinoma revealed by reconstruction and comprehensive analysis of the lncRNA-miRNA-mRNA regulatory network. Med Sci Monit 24:8250–8263
Acknowledgments
Authors deeply thanked all lab members.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Editor: Tetsuji Okamoto
Electronic Supplementary Material
ESM 1
(TXT 3 kb)
Supplementary Figure 1
(A) RT-qPCR detected FGD5-AS1 expression in CRC cells after transfection with pcDNA3.1/FGD5-AS1 or the empty vector. (B) CCK-8 assay detected CRC cell viability. (C) EdU assay evaluated CRC cell proliferation. (D) Casepase-3 activity assay measured CRC cell apoptosis. (E-F) Transwell assays assessed CRC cell migration and invasion. All the experiments were performed in triplicate. **P < 0.01, ***P < 0.001. (PNG 359 kb)
Supplementary Figure 2
(A) RT-qPCR detected the expression variation of the 16 miRNAs predicted to interact with FGD5-AS1. Data was presented as the sh-FGD5-AS1-2/sh-NC ratio. (B) RT-qPCR examined CDCA7 mRNA levels after transfection. (C) Western blot tested CDCA7 protein levels after transfection. All the experiments were performed in triplicate. *P < 0.05, **P < 0.01. (PNG 214 kb)
Supplementary Figure 3
(A-B) RT-qPCR and western blot confimed the knockdown efficiency of sh-CDCA7-1/2. (C) CCK-8 assay detected CRC cell viability. (D) EdU assay evaluated CRC cell proliferation. (E) Casepase-3 activity assay measured CRC cell apoptosis. (F-G) Transwell assays assessed CRC cell migration and invasion. All the experiments were performed in triplicate. *P < 0.05, **P < 0.01. (PNG 819 kb)
Rights and permissions
About this article
Cite this article
Li, D., Jiang, X., Zhang, X. et al. Long noncoding RNA FGD5-AS1 promotes colorectal cancer cell proliferation, migration, and invasion through upregulating CDCA7 via sponging miR-302e. In Vitro Cell.Dev.Biol.-Animal 55, 577–585 (2019). https://doi.org/10.1007/s11626-019-00376-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-019-00376-x