Abstract
Introduction
German shepherd dogs (GSDs) are a popular breed affected by numerous disorders. Few studies have explored genetic variations that influence canine blood metabolite levels.
Objectives
To investigate genetic variants affecting the natural metabolite variation in GSDs.
Methods
A total of 82 healthy GSDs were genotyped on the Illumina CanineHD Beadchip, assaying 173,650 markers. For each dog, 74 metabolites were measured through liquid and gas chromatography mass spectrometry (LC–MS and GC–MS) and were used as phenotypes for genome-wide association analyses (GWAS). Sliding window and homozygosity analyses were conducted to fine-map regions of interest, and to identify haplotypes and gene dosage effects.
Results
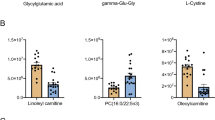
Summary statistics for 74 metabolites in this population of GSDs are reported. Forty-one metabolites had significant associations at a false discovery rate of 0.05. Two associations were located around genes which encode for enzymes for the relevant metabolites: 4-hydroxyproline was significantly associated to D-amino acid oxidase (DAO), and threonine to l-threonine 3-dehydrogenase (LOC477365). Three of the top ten haplotypes associated to 4-hydroxyproline included at least one SNP on DAO. These haplotypes occurred only in dogs with the highest 15 measurements of 4-hydroxyproline, ranging in frequency from 16.67 to 20%. None of the dogs were homozygous for these haplotypes. The top two haplotypes associated to threonine included SNPs on LOC477365 and were also overrepresented in dogs with the highest 15 measurements of threonine. These haplotypes occurred at a frequency of 90%, with 80% of these dogs homozygous for the haplotypes. In dogs with the lowest 15 measurements of threonine, the haplotypes occurred at a frequency of 26.67% and 0% homozygosity.
Conclusion
DAO and LOC477365 were identified as candidate genes affecting the natural plasma concentration of 4-hydroxyproline and threonine, respectively. Further investigations are needed to validate the effects of the variants on these genes.

Similar content being viewed by others
References
Adamski, J. (2012). Genome-wide association studies with metabolomics. Genome Medicine, 4(34), 1–7.
Allaway, D., Kamlage, B., Gilham, M. S., Hewson-Hughes, A. K., Wiemer, J. C., Colyer, A., et al. (2013). Effects of dietary glucose supplementation on the fasted plasma metabolome in cats and dogs. Metabolomics, 9(5), 1096–1108. https://doi.org/10.1007/s11306-013-0527-8.
Asher, L., Diesel, G., Summers, J. F., McGreevy, P. D., & Collins, L. M. (2009). Inherited defects in pedigree dogs. Part 1: Disorders related to breed standards. Veterinary Journal, 182, 402–411. https://doi.org/10.1016/j.tvjl.2009.11.002.
Bagheri, M., Farzadfar, F., Qi, L., Yekaninejad, M. S., Chamari, M., Zeleznik, O. A., et al. (2018). Obesity-related metabolomic profiles and discrimination of metabolically unhealthy obesity. Journal of Proteome Research, 17(4), 1452–1462. https://doi.org/10.1021/acs.jproteome.7b00802.
Ballevre, O., Cadenhead, A., Calder, A. G., Rees, W. D., Lobley, G. E., Fuller, M. F., et al. (1990). Quantitative partition of threonine oxidation in pigs: Effect of dietary threonine. American Journal of Physiology-Endocrinology and Metabolism, 259(4), E483–E491. https://doi.org/10.1152/ajpendo.1990.259.4.e483.
Bauer, D., Hamacher, K., Bröer, S., Pauleit, D., Palm, C., Zilles, K., et al. (2005). Preferred stereoselective brain uptake of d-serine—A modulator of glutamatergic neurotransmission. Nuclear Medicine and Biology, 32(8), 793–797. https://doi.org/10.1016/j.nucmedbio.2005.07.004.
Beckmann, M., Enot, D. P., Overy, D. P., Scott, I. M., Jones, P. G., Allaway, D., et al. (2010). Metabolite fingerprinting of urine suggests breed-specific dietary metabolism differences in domestic dogs. British Journal of Nutrition, 103(8), 1127–1138. https://doi.org/10.1017/S000711450999300X.
Benevides, G. P., Pimentel, E. R., Toyama, M. H., Novello, J. C., Marangoni, S., & Gomes, L. (2004). Biochemical and biomechanical analysis of tendons of caged and penned chickens. Connective Tissue Research, 45(4–5), 206–215. https://doi.org/10.1080/03008200490522997.
Biancalana, A., Veloso, L., & Gomes, L. (2010). Obesity affects collagen fibril diameter and mechanical properties of tendons in Zucker rats. Connective Tissue Research, 51(3), 171–178. https://doi.org/10.3109/03008200903191312.
Bird, M. I., & Nunn, P. B. (1983). Metabolic homoeostasis of l-threonine in the normally-fed rat. Importance of liver threonine dehydrogenase activity. Biochemical Journal, 214(3), 687–694. https://doi.org/10.1042/bj2140687.
Boughton, B. A., Callahan, D. L., Silva, C., Bowne, J., Nahid, A., Rupasinghe, T., et al. (2011). Comprehensive profiling and quantitation of amine group containing metabolites. Analytical Chemistry, 83(19), 7523–7530. https://doi.org/10.1021/ac201610x.
Breen, M., & Modiano, J. F. (2008). Evolutionarily conserved cytogenetic changes in hematological malignancies of dogs and humans—Man and his best friend share more than companionship. Chromosome Research, 16(1), 145–154. https://doi.org/10.1007/s10577-007-1212-4.
Burns, R. A., & Milner, J. A. (1982). Threonine, tryptophan and histidine requirements of immature Beagle dogs. Journal of Nutrition, 112(3), 447–452. https://doi.org/10.1093/jn/112.3.447.
Burns, R. A., Milner, J. A., & Corbin, J. E. (1981). Arginine: An indispensable amino acid for mature dogs. The Journal of Nutrition, 111(6), 1020–1024. https://doi.org/10.1093/jn/111.6.1020.
Bushell, K. R., Kim, Y., Chan, F. C., Ben-Neriah, S., Jenks, A., Alcaide, M., et al. (2015). Genetic inactivation of TRAF3 in canine and human B-cell lymphoma. Blood, 125(6), 999–1005. https://doi.org/10.1182/blood-2014-10-602714.
Campbell, C. L., Bhérer, C., Morrow, B. E., Boyko, A. R., & Auton, A. (2016). A pedigree-based map of recombination in the domestic dog genome. G3: Genes, Genomes, Genetics, 6(11), 3517–3524. https://doi.org/10.1534/g3.116.034678.
Chen, Y. P., Cheng, Y. F., Li, X. H., Yang, W. L., Wen, C., Zhuang, S., et al. (2017). Effects of threonine supplementation on the growth performance, immunity, oxidative status, intestinal integrity, and barrier function of broilers at the early age. Poultry Science, 96(2), 405–413. https://doi.org/10.3382/ps/pew240.
Cirulli, E. T., Guo, L., Leon Swisher, C., Shah, N., Huang, L., Napier, L. A., et al. (2019). Profound perturbation of the metabolome in obesity is associated with health risk. Cell Metabolism, 29(2), 488–500. https://doi.org/10.1016/j.cmet.2018.09.022.
Colyer, A., Gilham, M. S., Kamlage, B., Rein, D., & Allaway, D. (2011). Identification of intra- and inter-individual metabolite variation in plasma metabolite profiles of cats and dogs. British Journal of Nutrition, 106(S1), S146–S149. https://doi.org/10.1017/S000711451100081X.
Corfield, A. P., Myerscough, N., Longman, R., Sylvester, P., Arul, S., & Pignatelli, M. (2000). Mucins and mucosal protection in the gastrointestinal tract: New prospects for mucins in the pathology of gastrointestinal disease. Gut, 47(4), 589–594. https://doi.org/10.1136/gut.47.4.589.
D’Aniello, A. (2007). d-Aspartic acid: An endogenous amino acid with an important neuroendocrine role. Brain Research Reviews, 53(2), 215–234. https://doi.org/10.1016/j.brainresrev.2006.08.005.
D’Aniello, A., Vetere, A., & Petrucelli, L. (1993). Further study on the specificity of d-amino acid oxidase and of d-aspartate oxidase and time course for complete oxidation of d-amino acids. Comparative Biochemistry and Physiology, 105(3–4), 731–734. https://doi.org/10.1016/0305-0491(93)90113-J.
Defa, L., Changting, X., Shiyan, Q., Jinhui, Z., Johnson, E. W., & Thacker, P. A. (1999). Effects of dietary threonine on performance, plasma parameters and immune function of growing pigs. Animal Feed Science and Technology, 78(3–4), 179–188. https://doi.org/10.1016/S0377-8401(99)00005-X.
Dias, D., Hill, C., Jayasinghe, N., Atieno, J., Sutton, T., & Roessner, U. (2015). Quantitative profiling of polar primary metabolites of two chickpea cultivars with contrasting responses to salinity. Journal of Chromatography, B: Analytical Technologies in the Biomedical and Life Sciences, 1000, 1–13.
Drost, H.-G. G., & Paszkowski, J. (2017). Biomartr: genomic data retrieval with R. Bioinformatics, 33(18), 1216–1217. https://doi.org/10.1093/bioinformatics/btw821.
Durinck, S., Moreau, Y., Kasprzyk, A., Davis, S., De Moor, B., Brazma, A., et al. (2005). BioMart and Bioconductor: A powerful link between biological databases and microarray data analysis. Bioinformatics. https://doi.org/10.1093/bioinformatics/bti525.
Durinck, S., Spellman, P. T., Birney, E., & Huber, W. (2009). Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nature Protocols. https://doi.org/10.1038/nprot.2009.97.
Edgar, A. J. (2002). The human l-threonine 3-dehydrogenase gene is an expressed pseudogene. BMC Genetics, 3(1), 18. https://doi.org/10.1186/1471-2156-3-18.
Endo, M., Ohashi, K., & Mizuno, K. (2007). LIM kinase and slingshot are critical for neurite extension. Journal of Biological Chemistry, 282(18), 13692–13702. https://doi.org/10.1074/jbc.M610873200.
Fiehn, O. (2002). Metabolomics—The link between genotypes and phenotypes. Plant Molecular Biology, 48(1–2), 155–171. https://doi.org/10.1023/A:1013713905833.
Fleischer, S., Sharkey, M., Mealey, K., Ostrander, E. A., & Martinez, M. (2008). Pharmacogenetic and metabolic differences between dog breeds: Their impact on canine medicine and the use of the dog as a preclinical animal model. The AAPS Journal, 10(1), 110–119. https://doi.org/10.1208/s12248-008-9011-1.
Genchi, G. (2017). An overview on d-amino acids. Amino Acids, 49(9), 1521–1533. https://doi.org/10.1007/s00726-017-2459-5.
Gieger, C., Geistlinger, L., Altmaier, E., Hrabé de Angelis, M., Kronenberg, F., Meitinger, T., et al. (2008). Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum. PLoS Genetics, 4(11), e1000282. https://doi.org/10.1371/journal.pgen.1000282.
Hammer, V. A., Rogers, Q. R., & Freedland, R. A. (1996). Threonine is catabolized by l-threonine 3-dehydrogenase and threonine dehydratase in hepatocytes from domestic cats (Felis domestica). The Journal of Nutrition, 126(9), 2218–2226. https://doi.org/10.1093/jn/126.9.2218.
Hanna, V. S., & Hafez, E. A. A. (2018). Synopsis of arachidonic acid metabolism: A review. Journal of Advanced Research, 11, 23–32. https://doi.org/10.1016/j.jare.2018.03.005.
Heilmann, R. M., McNiel, E. A., Grützner, N., Lanerie, D. J., Suchodolski, J. S., & Steiner, J. M. (2017). Diagnostic performance of the urinary canine calgranulins in dogs with lower urinary or urogenital tract carcinoma. BMC Veterinary Research, 13(1), 112. https://doi.org/10.1186/s12917-017-1032-5.
Kathrani, A., Werling, D., & Allenspach, K. (2011). Canine breeds at high risk of developing inflammatory bowel disease in the South-Eastern UK. Veterinary Record, 169(24), 635. https://doi.org/10.1136/vr.d5380.
Kawabe, M., Baba, Y., Tamai, R., Yamamoto, R., Komori, M., Mori, T., et al. (2015). Profiling of plasma metabolites in canine oral melanoma using gas chromatography-mass spectrometry. Journal of Veterinary Medical Science, 77(8), 1025–1028. https://doi.org/10.1292/jvms.14-0641.
Kettunen, J., Demirkan, A., Würtz, P., Draisma, H. H. M., Haller, T., Rawal, R., et al. (2016). Genome-wide study for circulating metabolites identifies 62 loci and reveals novel systemic effects of LPA. Nature Communications, 7, 1–9. https://doi.org/10.1038/ncomms11122.
Krejsgaard, T., Vetter-Kauczok, C. S., Woetmann, A., Kneitz, H., Eriksen, K. W., Lovato, P., et al. (2009). Ectopic expression of B-lymphoid kinase in cutaneous T-cell lymphoma. Blood, 113(23), 5896–5904. https://doi.org/10.1182/blood-2008-09-181024.
Lindblad-Toh, K., Wade, C. M., Mikkelsen, T. S., Karlsson, E. K., Jaffe, D. B., Kamal, M., et al. (2005). Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature, 438(7069), 803–819. https://doi.org/10.1038/nature04338.
Linder, D. E., Freeman, L. M., Holden, S. L., Biourge, V., & German, A. J. (2013). Status of selected nutrients in obese dogs undergoing caloric restriction. BMC Veterinary Research, 9(1), 219. https://doi.org/10.1186/1746-6148-9-219.
Liu, Yun, Ke, X., Kang, H. Y., Wang, X. Q., Shen, Y., & Hong, S. L. (2016). Genetic risk of TNFSF4 and FAM167A-BLK polymorphisms in children with asthma and allergic rhinitis in a Han Chinese population. Journal of Asthma, 53(6), 567–575. https://doi.org/10.3109/02770903.2015.1108437.
Liu, Y., Wang, X., & Hu, C. A. A. (2017). Therapeutic potential of amino acids in inflammatory bowel disease. Nutrients. https://doi.org/10.3390/nu9090920.
Lloyd, A. J., Beckmann, M., Tailliart, K., Brown, W. Y., Draper, J., & Allaway, D. (2016). Characterisation of the main drivers of intra- and inter- breed variability in the plasma metabolome of dogs. Metabolomics, 12(4), 1–12. https://doi.org/10.1007/s11306-016-0997-6.
Lloyd, A. J., Beckmann, M., Wilson, T., Tailliart, K., Allaway, D., & Draper, J. (2017). Ultra high performance liquid chromatography–high resolution mass spectrometry plasma lipidomics can distinguish between canine breeds despite uncontrolled environmental variability and non-standardized diets. Metabolomics, 13(2), 1–11. https://doi.org/10.1007/s11306-016-1152-0.
Machiela, M. J., & Chanock, S. J. (2014). GWAS is going to the dogs. Genome Biology, 15(3), 105. https://doi.org/10.1186/gb4166.
Madrigal-Ruíz, P.-M., Navarro-Hernández, R.-E., Petri, M.-H., Chavarría-Ávila, E., Ríos-Ibarra, C., Castro-Albarrán, J., et al. (2016). Inverse relationship of the CMKLR1 relative expression and chemerin serum levels in obesity with dysmetabolic phenotype and insulin resistance. Mediators of Inflammation, 2016, 1–9. https://doi.org/10.1155/2016/3085390.
Malinovsky, A. V. (2017). Reason for indispensability of threonine in humans and other mammals in comparative aspect. Biochemistry (Moscow), 82(9), 1055–1060. https://doi.org/10.1134/S0006297917090097.
Mao, X., Zeng, X., Qiao, S., Wu, G., & Li, D. (2011). Specific roles of threonine in intestinal mucosal integrity and barrier function. Frontiers in Bioscience (Elite Edition), 3, 1192–1200.
Milner, J. A. (2018). Assessment of the essentiality of methionine, threonine, tryptophan, histidine and isoleucine in immature dogs. The Journal of Nutrition, 109(8), 1351–1357. https://doi.org/10.1093/jn/109.8.1351.
Mizuno, K., Niwa, R., Shuin, M., Ohashi, K., Kaji, N., & Uemura, T. (2003). Cell cycle-associated changes in slingshot phosphatase activity and roles in cytokinesis in animal cells. Journal of Biological Chemistry, 278(35), 33450–33455. https://doi.org/10.1074/jbc.m305802200.
Molina, H. (2002). The murine complement regulator Crry: New insights into the immunobiology of complement regulation. CMLS Cellular and Molecular Life Sciences, 59, 220–229.
Moncada, S., & Higgs, A. (1993). The l-arginine-nitric oxide pathway. New England Journal of Medicine, 329(27), 2002–2012. https://doi.org/10.1056/NEJM199312303292706.
Mortlock, S. A., Booth, R., Mazrier, H., Khatkar, M. S., & Williamson, P. (2016). Visualization of genome diversity in German Shepherd dogs. Bioinformatics and Biology Insights, 9, 37–42. https://doi.org/10.4137/BBI.S30524.
Moundras, C., Bercovici, D., Rémésy, C., & Demigné, C. (1992). Influence of glucogenic amino acids on the hepatic metabolism of threonine. Biochimica et Biophysica Acta (BBA)—General Subjects, 1115(3), 212–219. https://doi.org/10.1016/0304-4165(92)90056-Z.
Nakamura, N., & Hirose, S. (2008). Regulation of mitochondrial morphology by USP30, a deubiquitinating enzyme present in the mitochondrial outer membrane. Molecular Biology of the Cell, 19(5), 1903–1911. https://doi.org/10.1091/mbc.e07-11-1103.
O’Kell, A. L., Garrett, T. J., Wasserfall, C., & Atkinson, M. A. (2017). Untargeted metabolomic analysis in naturally occurring canine diabetes mellitus identifies similarities to human Type 1 diabetes. Scientific Reports, 7(1), 1–8. https://doi.org/10.1038/s41598-017-09908-5.
Ollier, W. E. R., Kennedy, L. J., Thomson, W., Barnes, A. N., Bell, S. C., Bennett, D., et al. (2001). Dog MHC alleles containing the human RA shared epitope confer susceptibility to canine rheumatoid arthritis. Immunogenetics, 53(8), 669–673. https://doi.org/10.1007/s002510100372.
Olmstead, I. L. D., Hill, D. R. A., Dias, D. A., Jayasinghe, N. S., Callahan, D. L., Kentish, S. E., et al. (2013). A quantitative analysis of microalgal lipids for optimization of biodiesel and omega-3 production. Biotechnology and Bioengineering, 110(8), 2096–2104. https://doi.org/10.1002/bit.24844.
Parker, H. G. (2012). Genomic analyses of modern dog breeds. Mammalian Genome, 23(1–2), 19–27. https://doi.org/10.1007/s00335-011-9387-6.
Peiravan, A., Bertolini, F., Rothschild, M. F., Simpson, K. W., Jergens, A. E., Allenspach, K., et al. (2018). Genome-wide association studies of inflammatory bowel disease in German Shepherd dogs. PLoS ONE, 13(7), e0200685. https://doi.org/10.1371/journal.pone.0200685.
Petersen, D. L., Berthelsen, J., Willerslev-Olsen, A., Fredholm, S., Dabelsteen, S., Bonefeld, C. M., et al. (2017). A novel BLK-induced tumor model. Tumor Biology, 39(7), 101042831771419. https://doi.org/10.1177/1010428317714196.
Polley, M. J., Nachman, R. L., & Weksler, B. B. (1981). Human complement in the arachidonic acid transformation pathway in platelets. Journal of Experimental Medicine, 153(2), 257–268.
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M. A. R., Bender, D., et al. (2007). PLINK: A tool set for whole-genome association and population-based linkage analyses. American Journal of Human Genetics, 81(3), 559–575. https://doi.org/10.1086/519795.
Puurunen, J., Sulkama, S., Tiira, K., Araujo, C., Lehtonen, M., Hanhineva, K., et al. (2016a). A non-targeted metabolite profiling pilot study suggests that tryptophan and lipid metabolisms are linked with ADHD-like behaviours in dogs. Behavioral and Brain Functions, 12(1), 27. https://doi.org/10.1186/s12993-016-0112-1.
Puurunen, J., Tiira, K., Lehtonen, M., Hanhineva, K., & Lohi, H. (2016b). Non-targeted metabolite profiling reveals changes in oxidative stress, tryptophan and lipid metabolisms in fearful dogs. Behavioral and Brain Functions. https://doi.org/10.1186/s12993-016-0091-2.
R Special Interest Group on Databases (R-SIG-DB), Wickham, H., & Müller, K. (2018). DBI: R Database Interface. R package. Retrieved from https://cran.r-project.org/package=DBI.
Rémond, D., Buffière, C., Godin, J.-P., Mirand, P. P., Obled, C., Papet, I., et al. (2009). Intestinal inflammation increases gastrointestinal threonine uptake and mucin synthesis in enterally fed minipigs. The Journal of Nutrition, 139(4), 720–726. https://doi.org/10.3945/jn.108.101675.
Rhee, E. P., Ho, J. E., Chen, M. H., Shen, D., Cheng, S., Larson, M. G., et al. (2013). A genome-wide association study of the human metabolome in a community-based cohort. Cell Metabolism, 18(1), 130–143. https://doi.org/10.1016/j.cmet.2013.06.013.
Richards, K. L., & Suter, S. E. (2015). Man’s best friend: What can pet dogs teach us about non-Hodgkin’s lymphoma? Immunological Reviews, 263(1), 173–191. https://doi.org/10.1111/imr.12238.
Richards, S. E., Wang, Y., Claus, S. P., Lawler, D., Kochhar, S., Holmes, E., et al. (2013). Metabolic phenotype modulation by caloric restriction in a lifelong dog study. Journal of Proteome Research, 12(7), 3117–3127. https://doi.org/10.1021/pr301097k.
Rimbault, M., & Ostrander, E. A. (2012). So many doggone traits: Mapping genetics of multiple phenotypes in the domestic dog. Human Molecular Genetics, 21(1), 52–57. https://doi.org/10.1093/hmg/dds323.
Rowell, J. L., McCarthy, D. O., & Alvarez, C. E. (2011). Dog models of naturally occurring cancer. Trends in Molecular Medicine, 17(7), 380–388. https://doi.org/10.1016/j.molmed.2011.02.004.
Ruth, M. R., & Field, C. J. (2013). The immune modifying effects of amino acids on gut-associated lymphoid tissue. Journal of Animal Science and Biotechnology, 4(1), 27. https://doi.org/10.1186/2049-1891-4-27.
Samuelsson, B. (1991). Arachidonic acid metabolism: Role in inflammation. Zeitschrift für Rheumatologie, 50(Suppl 1), 3–6.
Shen, K. Z., Cox, B. A., & Johnson, S. W. (1997). L-arginine potentiates GABA-mediated synaptic transmission by a nitric oxide-independent mechanism in rat dopamine neurons. Neuroscience, 79(3), 649–658. https://doi.org/10.1016/S0306-4522(97)00024-9.
Shin, H. Y., Lee, D. C., Chu, S. H., Jeon, J. Y., Lee, M. K., Im, J. A., et al. (2012). Chemerin levels are positively correlated with abdominal visceral fat accumulation. Clinical Endocrinology, 77(1), 47–50. https://doi.org/10.1111/j.1365-2265.2011.04217.x.
Simpfendorfer, K. R., Armstead, B. E., Shih, A., Li, W., Curran, M., Manjarrez-Orduño, N., et al. (2015). Autoimmune disease-associated haplotypes of BLK exhibit lowered thresholds for B cell activation and expansion of Ig class-switched B cells. Arthritis and Rheumatology, 67(11), 2866–2876. https://doi.org/10.1002/art.39301.
Soder, J., Hagman, R., Dicksved, J., Lindase, S., Malmlof, K., Agback, P., et al. (2017). The urine metabolome differs between lean and overweight Labrador Retriever dogs during a feed-challenge. PLoS ONE, 12(6), 1–17. https://doi.org/10.1371/journal.pone.0180086.
Song, G. G., & Lee, Y. H. (2017). Association between BLK polymorphisms and susceptibility to SLE. Zeitschrift für Rheumatologie, 76(2), 176–182. https://doi.org/10.1007/s00393-016-0072-8.
Storey, J. (2015). qvalue: Q-value estimation for false discovery rate control. R package. https://doi.org/10.1002/ejoc.201200111.
Summers, J. F., Diesel, G., Asher, L., McGreevy, P. D., & Collins, L. M. (2010). Inherited defects in pedigree dogs. Part 2: Disorders that are not related to breed standards. Veterinary Journal, 183(1), 39–45. https://doi.org/10.1016/j.tvjl.2009.11.002.
Takano, T., & Cybulsky, A. V. (2000). Complement C5b-9-mediated arachidonic acid metabolism in glomerular epithelial cells: Role of cyclooxygenase-1 and -2. American Journal of Pathology, 156(6), 2091–2101. https://doi.org/10.1016/S0002-9440(10)65080-8.
Tamai, R., Furuya, M., Hatoya, S., Akiyoshi, H., Yamamoto, R., Komori, Y., et al. (2014). Profiling of serum metabolites in canine lymphoma using gas chromatography mass spectrometry. Journal of Veterinary Medical Science, 76(11), 1513–1518. https://doi.org/10.1292/JVMS.14-0043.
Tang, L., Tong, Y., Cao, H., Xie, S., Yang, Q., Zhang, F., et al. (2014). The MTMR9 rs2293855 polymorphism is associated with glucose tolerance, insulin secretion, insulin sensitivity and increased risk of prediabetes. Gene, 546(2), 150–155. https://doi.org/10.1016/j.gene.2014.06.028.
Trang, L. E., Fürst, P., Odeback, A. C., & Lövgren, O. (1985). Plasma amino acids in rheumatoid arthritis. Scandinavian Journal of Rheumatology, 14(4), 393–402. https://doi.org/10.3109/03009748509102044.
Tsai, K. L., Noorai, R. E., Starr-Moss, A. N., Quignon, P., Rinz, C. J., Ostrander, E. A., et al. (2012). Genome-wide association studies for multiple diseases of the German Shepherd Dog. Mammalian Genome, 23(1–2), 203–211. https://doi.org/10.1007/s00335-011-9376-9.
Turner, S. D. (2018). qqman: An R package for visualizing GWAS results using Q-Q and manhattan plots. Journal of Open Source Software. https://doi.org/10.21105/joss.00731.
Vaysse, A., Ratnakumar, A., Derrien, T., Axelsson, E., Pielberg, G. R., Sigurdsson, S., et al. (2011). Identification of genomic regions associated with phenotypic variation between dog breeds using selection mapping. PLoS Genetics. https://doi.org/10.1371/journal.pgen.1002316.
Viant, M. R., Ludwig, C., Rhodes, S., Günther, U. L., & Allaway, D. (2007). Validation of a urine metabolome fingerprint in dog for phenotypic classification. Metabolomics, 3(4), 453–463. https://doi.org/10.1007/s11306-007-0092-0.
Vilson, Å., Bonnett, B., Hansson-Hamlin, H., & Hedhammar, Å. (2013). Disease patterns in 32,486 insured German Shepherd dogs in Sweden: 1995-2006. Veterinary Record, 173(5), 116. https://doi.org/10.1136/vr.101577.
Wahl, J. M., Herbst, S. M., Clark, L. A., Tsai, K. L., & Murphy, K. E. (2008). A review of hereditary diseases of the German Shepherd dog. Journal of Veterinary Behavior: Clinical Applications and Research, 3(6), 255–265. https://doi.org/10.1016/j.jveb.2008.05.004.
Wang, X., Qiao, S. Y., Liu, M., & Ma, Y. X. (2006). Effects of graded levels of true ileal digestible threonine on performance, serum parameters and immune function of 10-25 kg pigs. Animal Feed Science and Technology, 129(3–4), 264–278. https://doi.org/10.1016/j.anifeedsci.2006.01.003.
Warnes, G. R., Bolker, B., Bonebakker, L., Gentleman, R., Liaw, W. H. A., Lumley, T., et al. (2016). Package “gplots”: Various R programming tools for plotting data. R package. https://doi.org/10.1038/nnano.2010.132.
White, M. E., Hayward, J. J., Stokol, T., & Boyko, A. R. (2015). Genetic mapping of novel loci affecting canine blood phenotypes. PLoS ONE, 10(12), e0145199. https://doi.org/10.1371/journal.pone.0145199.
Wickham, H. (2016). Package ‘ggplot2’: Elegant Graphics for Data Analysis. New York: Springer-Verlag. https://doi.org/10.1093/bioinformatics/btr406.
Wu, G., Bazer, F. W., Burghardt, R. C., Johnson, G. A., Woo, S., Darrell, K., et al. (2011). Proline and hydroxyproline metabolism: Implications for animal and human nutrition. Amino Acids, 40(4), 1053–1063. https://doi.org/10.1007/s00726-010-0715-z.
Wu, G., Bazer, F. W., Davis, T. A., Kim, S. W., Li, P., Marc Rhoads, J., et al. (2009). Arginine metabolism and nutrition in growth, health and disease. Amino Acids, 37(1), 153–168. https://doi.org/10.1007/s00726-008-0210-y.
Yanagiya, T., Tanabe, A., Iida, A., Saito, S., Sekine, A., Takahashi, A., et al. (2007). Association of single-nucleotide polymorphisms in MTMR9 gene with obesity. Human Molecular Genetics, 16(24), 3017–3026. https://doi.org/10.1093/hmg/ddm260.
Yang, J., Lee, S. H., Goddard, M. E., & Visscher, P. M. (2011). GCTA: A tool for genome-wide complex trait analysis. The American Journal of Human Genetics, 88(1), 76–82. https://doi.org/10.1016/j.ajhg.2010.11.011.
Yao, J., de la Iglesia, H. O., & Bajjalieh, S. M. (2013). Loss of the SV2-like protein SVOP produces no apparent deficits in laboratory mice. PLoS ONE, 8(7), e68215. https://doi.org/10.1371/journal.pone.0068215.
Zhang, H., Wang, L., Huang, Y., Zhuang, C., Zhao, G., Liu, R., et al. (2012a). Influence of BLK polymorphisms on the risk of rheumatoid arthritis. Molecular Biology Reports, 39(11), 9965–9970. https://doi.org/10.1007/s11033-012-1865-8.
Zhang, J., Wei, S., Liu, L., Nagana Gowda, G. A., Bonney, P., Stewart, J., et al. (2012b). NMR-based metabolomics study of canine bladder cancer. Biochimica et Biophysica Acta (BBA)—Molecular Basis of Disease, 1822(11), 1807–1814. https://doi.org/10.1016/j.bbadis.2012.08.001.
Zhou, Y., Li, X., Wang, G., & Li, X. (2016). Association of FAM167A-BLK rs2736340 polymorphism with susceptibility to autoimmune diseases: A meta-analysis. Immunological Investigations, 45(4), 336–348. https://doi.org/10.3109/08820139.2016.1157812.
Acknowledgements
This work was supported by the Canine Research Foundation. This research is supported by an Australian Government Research Training Program (RTP) Scholarship. Metabolites were extracted and analysed from plasma at Metabolomics Australia (School of BioSciences, University of Melbourne, Australia), a National Collaborative Research Infrastructure Strategy (NCRIS) initiative under Bioplatforms Australia, Pty Ltd. The authors would like to thank Himasha Mendis, Nirupama Jayasinghe and Alice Ng from Metabolomics Australia who extracted and analysed metabolites. The authors would also like to thank the owners and dogs that donated samples for this study.
Author information
Authors and Affiliations
Contributions
PXYS performed research, analysed the data, and wrote the manuscript. JMMC, CJJ, and SM contributed to performing research, analysis of data, and writing the manuscript. SM, RB, BC and SC collected the samples and data, and conceived the study. MSK contributed in the analysis, interpretations and writing. UR and SN advised on metabolomics analysis and contributed to writing. PW conceived the study, performed research, contributed to the analysis of the data and writing of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflicts of interest.
Ethical Approval
All protocols in this study was conducted in accordance with the guidelines of the Animal Research Act, NSW, Australia, approved by the University of Sydney’s Animal Ethics Committee under protocols 444 and 4949.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11306_2019_1586_MOESM1_ESM.docx
Supplementary material 1—ESM_1: Details on plasma preparation, machines used, and analysis methods for LC–MS and GC–MS for amines, sugars, organic acids, and fatty acids (DOCX 18 kb)
11306_2019_1586_MOESM2_ESM.pdf
Supplementary material 2—ESM_2: Summary statistics of amino acids. Outliers were defined as measurements beyond 4 standard deviations from the mean (PDF 205 kb)
11306_2019_1586_MOESM3_ESM.pdf
Supplementary material 3—ESM_3: Summary statistics of fatty acids. Outliers were defined as measurements beyond 4 standard deviations from the mean (PDF 158 kb)
11306_2019_1586_MOESM4_ESM.pdf
Supplementary material 4—ESM_4: Summary statistics of sugars. Outliers were defined as measurements beyond 4 standard deviations from the mean (PDF 166 kb)
11306_2019_1586_MOESM5_ESM.pdf
Supplementary material 5—ESM_5: Summary statistics of organic acids. Outliers were defined as measurements beyond 4 standard deviations from the mean (PDF 78 kb)
11306_2019_1586_MOESM6_ESM.pdf
Supplementary material 6—ESM_6: Correlation plots for all metabolites, and for each group of metabolites – amino acids, fatty acids, sugars and organic acids respectively. Complete observations and Spearman’s rank correlation were used from the dataset adjusted for normality and pruned for outliers (PDF 266 kb)
11306_2019_1586_MOESM7_ESM.pdf
Supplementary material 7—ESM_7: Genome-wide Manhattan plots of amino acids using output from the mixed linear model analysis. Red line indicates a genome-wide q-value cut-off of 0.05 (PDF 8702 kb)
11306_2019_1586_MOESM8_ESM.pdf
Supplementary material 8—ESM_8: Manhattan plots of significant chromosome associations for amino acids at a chromosome-wide q-value cut-off of 0.05. Where present, red line indicates a q-value cut-off of 0.05, blue line indicates a q-value cut-off of 0.01 (PDF 494 kb)
11306_2019_1586_MOESM9_ESM.pdf
Supplementary material 9—ESM_9: Genome-wide Manhattan plots of fatty acids using output from the mixed linear model analysis. Red line indicates a genome-wide q-value cut-off of 0.05 (PDF 4513 kb)
11306_2019_1586_MOESM10_ESM.pdf
Supplementary material 10—ESM_10: Manhattan plots of significant chromosome associations for fatty acids at a chromosome-wide q-value cut-off of 0.05. Where present, red line indicates a q-value cut-off of 0.05, blue line indicates a q-value cut-off of 0.01 (PDF 254 kb)
11306_2019_1586_MOESM11_ESM.pdf
Supplementary material 11—ESM_11: Genome-wide Manhattan plots of sugars using output from the mixed linear model analysis. Red line indicates a genome-wide q-value cut-off of 0.05 (PDF 6025 kb)
11306_2019_1586_MOESM12_ESM.pdf
Supplementary material 12—ESM_12: Manhattan plots of significant chromosome associations for sugars at a chromosome-wide q-value cut-off of 0.05. Where present, red line indicates a q-value cut-off of 0.05, blue line indicates a q-value cut-off of 0.01 (PDF 136 kb)
11306_2019_1586_MOESM13_ESM.pdf
Supplementary material 13—ESM_13: Genome-wide Manhattan plots of organic acids using output from the mixed linear model analysis. Red lines indicate a genome-wide q-value cut-off of 0.05 (PDF 3010 kb)
11306_2019_1586_MOESM14_ESM.pdf
Supplementary material 14—ESM_14: Manhattan plots of significant chromosome associations for organic acids at a chromosome-wide q-value cut-off of 0.05. Where present, red line indicates a q-value cut-off of 0.05, blue line indicates a q-value cut-off of 0.01 (PDF 246 kb)
11306_2019_1586_MOESM16_ESM.xlsx
Supplementary material 16—ESM_16: Sliding window output of 2, 4 or 6 SNP haplotypes (p < 0.0001) for each significant region and the homozygosity of each haplotype in the top 15 and lowest 15 measurements for each metabolite (XLSX 925 kb)
11306_2019_1586_MOESM22_ESM.xlsx
Supplementary material 22—ESM_22: Metadata (age and sex) for GSDs in this study. Originally from Mortlock et al. 2016 (XLSX 10 kb)
11306_2019_1586_MOESM23_ESM.xlsx
Supplementary material 23—ESM_23: Summary statistics for all metabolites after adjusting for normality, pruning for outliers, and back-transforming the data. SD = Standard deviation; Q1 = first quartile; Q3 = third quartile; IQR = Interquartile range; CV = coefficient of variation(XLS X 21 kb)
Rights and permissions
About this article
Cite this article
Soh, P.X.Y., Marin Cely, J.M., Mortlock, SA. et al. Genome-wide association studies of 74 plasma metabolites of German shepherd dogs reveal two metabolites associated with genes encoding their enzymes. Metabolomics 15, 123 (2019). https://doi.org/10.1007/s11306-019-1586-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-019-1586-2