Abstract
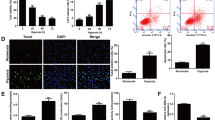
Human cytomegalovirus (HCMV) infection has been linked to the pathogenesis of vasculopathy by inducing dysfunction of vascular cells such as endothelial cells. Hcmv-miR-UL112 is the most well-characterized HCMV-encoded microRNA occurring in the plasma of patients with cardiovascular diseases such as hypertension, while the specific underlying pathophysiological mechanisms are yet to be defined. The current study investigated the effect of hcmv-miR-UL112 on the growth and proliferation of human umbilical vascular endothelial cells (HUVECs); it might also be associated with signaling pathways. An adenovirus vector was designed and synthesized to stably express hcmv-miR-UL112 in HUVECs. Cell Counting Kit-8 results showed that ectopically expressed hcmv-miR-UL112 can significantly increase the proliferation of HUVECs (p < 0.05). Flow cytometry revealed that the S-phase fraction in the cell cycle analysis was raised significantly after overexpression of hcmv-miR-UL112 (p < 0.05). Gene expression profile analysis, using the microarray technology, revealed 303 up-regulated and 62 down-regulated genes in HUVECs by comparing the AD-hcmv-miR-UL112-infected and control groups (p < 0.05 and > 2 fold change). Kyoto Encyclopedia of Genes and Genomes and Reactome Pathway, chosen as the functional annotation categories, were affected by hcmv-miR-UL112 adenovirus vector. The significantly altered pathways mainly include the mitogen-activated protein kinase signaling pathway, cell adhesion molecules, chemokine signaling pathway, cytokine–cytokine receptor interaction, circadian rhythm-mammal, mineral absorption, protein processing in the endoplasmic reticulum, proximal tubule bicarbonate reclamation, vasopressin-regulated water reabsorption, and arachidonic acid metabolism. In conclusion, hcmv-miR-UL112 could serve as a potential biomarker, and the miRNA-mediated regulation of signaling pathways might play significant roles in the physiological effects of hcmv-associated diseases.



Similar content being viewed by others
References
N. Stern-Ginossar, B. Weisburd, A. Michalski, V.T. Le, M.Y. Hein, S.X. Huang, M. Ma, B. Shen, S.B. Qian, H. Hengel, M. Mann, N.T. Ingolia, J.S. Weissman, Decoding human cytomegalovirus. Science 338, 1088–1093 (2012)
S. Li, J. Zhu, W. Zhang, Y. Chen, K. Zhang, L.M. Popescu, X. Ma, W.B. Lau, R. Rong, X. Yu, B. Wang, Y. Li, C. Xiao, M. Zhang, S. Wang, L. Yu, A.F. Chen, X. Yang, J. Cai, Signature microRNA expression profile of essential hypertension and its novel link to human cytomegalovirus infection. Circulation 124, 175–184 (2011)
L. Yi, D.X. Wang, Z.J. Feng, Detection of human cytomegalovirus in atherosclerotic carotid arteries in humans. J. Formos. Med. Assoc. 107, 774–781 (2008)
E. Xenaki, J. Hassoulas, S. Apostolakis, G. Sourvinos, D.A. Spandidos, Detection of cytomegalovirus in atherosclerotic plaques and nonatherosclerotic arteries. Angiology 60, 504–508 (2009)
M. Popović, K. Smiljanić, B. Dobutović, T. Syrovets, T. Simmet, E.R. Isenocvić, Human cytomegalovirus infection and atherothrombosis. J. Thromb. Thrombolysis 33, 160–172 (2012)
M.A. Jarvis, J.A. Nelson, Human cytomegalovirus persistence and latency in endothelial cells and macrophages. Curr. Opin. Microbiol. 5, 403–407 (2002)
C. Grahame-clarke, Human cytomegalovirus, endothelial function and atherosclerosis. Herpes. 12, 42–45 (2005)
M. Weis, T.N. Kledal, K.Y. Lin, S.N. Panchal, S.Z. Gao, H.A. Valantine, E.S. Mocarski, J.P. Cooke, Cytomegalovirus infection impairs the nitric oxide synthase pathway: role of asymmetric dimethylarginine in transplant arteriosclerosis. Circulation 109, 500–505 (2004)
S.M. Billstrom, R. Christensen, G.S. Worthen, Human cytomegalovirus protects endothelial cells from apoptosis induced by growth factor withdrawal. J. Clin. Virol. 25(Suppl 2), S149–S157 (2002)
P. Petrakopoulou, M. Kubrich, S. Pehlivanli, B. Meiser, B. Reichart, W. von Scheidt, M. Weis, Cytomegalovirus infection in heart transplant recipients is associated with impaired endothelial function. Circulation 110, II207–212 (2004)
L.P. Lim, M.E. Glasner, S. Yekta, C.B. Burge, D.P. Bartel, Vertebrate microRNA genes. Science 299, 1540 (2003)
D.P. Bartel, MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009)
F. Grey, L. Hook, J. Nelson, The functions of herpesvirus-encoded microRNAs. Med. Microbiol. Immunol. 197, 261–267 (2008)
F. Grey, A. Antoniewicz, E. Allen, J. Saugstad, A. McShea, J.C. Carrington, J. Nelson, Identification and characterization of human cytomegalovirus-encoded microRNAs. J. Virol. 79, 12095–12099 (2005)
F. Grey, H. Meyers, E.A. White, D.H. Spector, J. Nelson, A human cytomegalovirus-encoded microRNA regulates expression of multiple viral genes involved in replication. PLoS Pathog. 3, e163 (2007)
E. Murphy, J. Vanicek, H. Robins, T. Shenk, A.J. Levine, Suppression of immediate-early viral gene expression by herpesvirus-coded microRNAs: implications for latency. Proc. Natl. Acad. Sci. USA 105, 5453–5458 (2008)
N. Stern-Ginossar, N. Saleh, M.D. Goldberg, M. Prichard, D.G. Wolf, O. Mandelboim, Analysis of human cytomegalovirus-encoded microRNA activity during infection. J. Virol. 83, 10684–10693 (2009)
L.M. Hook, F. Grey, R. Grabski, R. Tirabassi, T. Doyle, M. Hancock, I. Landais, S. Jeng, S. McWeeney, W. Britt, J.A. Nelson, Cytomegalovirus miRNAs target secretory pathway genes to facilitate formation of the virion assembly compartment and reduce cytokine secretion. Cell Host Microbe 15, 363–373 (2014)
N. Stern-Ginossar, N. Elefant, A. Zimmermann, D.G. Wolf, N. Saleh, M. Biton, E. Horwitz, Z. Prokocimer, M. Prichard, G. Hahn, D. Goldman-Wohl, C. Greenfield, S. Yagel, H. Hengel, Y. Altuvia, H. Margalit, O. Mandelboim, Host immune system gene targeting by a viral miRNA. Science 317, 376–381 (2007)
D. Nachmani, D. Lankry, D.G. Wolf, O. Mandelboim, The human cytomegalovirus microRNA miR-UL112 acts synergistically with a cellular microRNA to escape immune elimination. Nat. Immunol. 11, 806–813 (2010)
Y. Huang, Y. Qi, Y. Ma, R. He, Y. Ji, Z. Sun, Q. Ruan, The expression of interleukin-32 is activated by human cytomegalovirus infection and down regulated by hcmv-miR-UL112-1. Virol J 10, 51 (2013)
H. Guo, N.T. Ingolia, J.S. Weissman, D.P. Bartel, Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466, 835–840 (2010)
J. Fan, W. Zhang, Q. Liu, Human cytomegalovirus-encoded miR-US25-1 aggravates the oxidised low density lipoprotein-induced apoptosis of endothelial cells. Biomed. Res. Int. 2014, 531979 (2014)
F. Sandra, Y.H. Oktaviono, M.A. Widodo, Y. Dirgantara, A. Chouw, D. Sargowo, Endothelial progenitor cells proliferated via MEK-dependent p42 MAPK signaling pathway. Mol. Cell. Biochem. 400, 201–206 (2015)
M. Katz, I. Amit, Y. Yarden, Regulation of MAPKs by growth factors and receptor tyrosine kinases. Biochim. Biophys. Acta 1773, 1161–1176 (2007)
L.M. Graves, H.I. Guy, P. Kozlowski, M. Huang, E. Lazarowski, R.M. Pope, M.A. Collins, E.N. Dahlstrand, H.S. 3rd Earp, D.R. Evans, Regulation of carbamoyl phosphate synthetase by MAP kinase. Nature 403, 328–332 (2000)
Y.H. Shen, L. Zhang, B. Utama, J. Wang, Y. Gan, X. Wang, J. Wang, L. Chen, G.M. Vercellotti, J.S. Coselli, J.L. Mehta, X.L. Wang, Human cytomegalovirus inhibits Akt-mediated eNOS activation through upregulating PTEN (phosphatase and tensin homolog deleted on chromosome 10). Cardiovasc. Res. 69, 502–511 (2006)
T. Murakami, C. Mataki, C. Nagao, M. Umetani, Y. Wada, M. Ishii, S. Tsutsumi, T. Kohro, A. Saiura, H. Aburatani, T. Hamakubo, T. Kodama, The gene expression profile of human umbilical vein endothelial cells stimulated by tumor necrosis factor alpha using dna microarray analysis. J Atheroscler Thromb 7, 39–44 (2000)
C.F. Krieglstein, D.N. Granger, Adhesion molecules and their role in vascular disease. Am. J. Hypertens. 14, 44S (2001)
I.F. Charo, M.B. Taubman, Chemokines in the pathogenesis of vascular disease. Circ. Res. 95, 858–866 (2004)
R.C. Friedman, K.K. Farh, C.B. Burge, D.P. Bartel, Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009)
L. Wang, A.L. Oberg, Y.W. Asmann, H. Sicotte, S.K. McDonnell, S.M. Riska, W. Liu, C.J. Steer, S. Subramanian, J.M. Cunningham, J.R. Cerhan, S.N. Thibodeau, Genome-wide transcriptional profiling reveals microRNA-correlated genes and biological processes in human lymphoblastoid cell lines. PLoS ONE 4, e5878 (2009)
T. Vaissiere, R.J. Hung, D. Zaridze, A. Moukeria, C. Cuenin, V. Fasolo, G. Ferro, A. Paliwal, P. Hainaut, P. Brennan, J. Tost, P. Boffetta, Z. Herceg, Quantitative analysis of DNA methylation profiles in lung cancer identifies aberrant DNA methylation of specific genes and its association with gender and cancer risk factors. Cancer Res. 69, 243–252 (2009)
A.M. Roccaro, A. Sacco, X. Jia, A.K. Azab, P. Maiso, H.T. Ngo, F. Azab, J. Runnels, P. Quang, I.M. Ghobrial, microRNA dependent modulation of histone acetylation in Waldenstrom macroglobulinemia. Blood 116, 1506–1514 (2010)
Acknowledgements
This work was supported by grants from the Zhejiang Provincial Natural Science Foundation of China [Nos. LY13H020001 and LQ13H100001] and the Science and Technology Bureau of Zhoushan [No. 2012C13024].
Author information
Authors and Affiliations
Contributions
KS and LX contributed equally in performing cytology experiments and data analysis, DC undertook flow cytometry, WT was responsible for microarray data analysis, and YH is the corresponding author of this article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Edited by Hartmut Hengel.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
The complete list of significant up-regulation genes in AD-hcmv-miR-UL112 transfected HUVECs. The gene expression profiling of AD-hcmv-miR-UL112 transfected HUVECs was compared with that of negative control groups. Supplementary material 1 (DOC 557 kb)
Table S2
The complete list of significant down-regulation genes in AD-hcmv-miR-UL112 transfected HUVECs. The gene expression profiling of AD-hcmv-miR-UL112 transfected HUVECs was compared with that of negative control groups. Supplementary material 2 (DOC 121 kb)
Rights and permissions
About this article
Cite this article
Shen, K., Xu, L., Chen, D. et al. Human cytomegalovirus-encoded miR-UL112 contributes to HCMV-mediated vascular diseases by inducing vascular endothelial cell dysfunction. Virus Genes 54, 172–181 (2018). https://doi.org/10.1007/s11262-018-1532-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-018-1532-9