Abstract
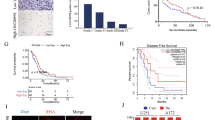
Glioma is the most frequent primary malignant brain tumor, which is characterized by high incidence and mortality, with a poor prognosis. Numerous studies have revealed the abnormal expression of long non-coding RNAs in gliomas. This study explored the effects and potential mechanism of LINC00663 in glioma. The LINC00663 levels and their prognostic values were analyzed from the GEO databases using bioinformatics. Also, LINC00663 expression in tissue samples and cell lines was measured using qRT-PCR. The roles of LINC00663 in glioma were confirmed using CCK8, EdU assay as well as Transwell tests. Moreover, the influences of LINC00663 on the AKT/mTOR signal cascades were detected using western blotting assay. LINC00663 expression was higher in both glioma tissues and cell lines than that in the normal brain tissues and human astrocytes. High expression of LINC00663 led to the low overall survival rate of patients with glioma. LINC00663 knockdown notably restrained cell proliferation, migration, and invasion abilities by decreasing the activation of AKT and mTOR. This study indicated that LINC00663 might have a cancer-promoting role in accelerating glioma development and progression through regulating AKT/mTOR pathway.




Similar content being viewed by others
Data Availability
All data generated or analyzed during the current study are included in this published article.
References
Ostrom QT, Cioffi G, Gittleman H, Patil N, Waite K, Kruchko C, Barnholtz-Sloan JS (2019) CBTRUS Statistical Report: primary brain and other central nervous system tumors diagnosed in the United States in 2012–2016. Neuro Oncol 21(Suppl 5):v1–v100. https://doi.org/10.1093/neuonc/noz150
Lapointe S, Perry A, Butowski NA (2018) Primary brain tumours in adults. Lancet 392(10145):432–446. https://doi.org/10.1016/S0140-6736(18)30990-5
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Tomiyama A, Ichimura K (2019) Signal transduction pathways and resistance to targeted therapies in glioma. Semin Cancer Biol 58:118–129. https://doi.org/10.1016/j.semcancer.2019.01.004
Cheng J, Meng JL, Zhu L, Peng Y (2020) Exosomal noncoding RNAs in glioma: biological functions and potential clinical applications. Mol Cancer 19(1):66. https://doi.org/10.1186/s12943-020-01189-3
Ng SY, Lin L, Soh BS, Stanton LW (2013) Long noncoding RNAs in development and disease of the central nervous system. Trends Genet 29(8):461–468. https://doi.org/10.1016/j.tig.2013.03.002
Uszczynska-Ratajczak B, Lagarde J, Frankish A, Guigó R, Johnson R (2018) Towards a complete map of the human long non-coding RNA transcriptome. Nat Rev Gene 19:535–548. https://doi.org/10.1038/s41576-018-0017-y
Shi XF, Sun M, Liu HB, Yao YW, Song Y (2013) Long non-coding RNAs: a new frontier in the study of human diseases. Cancer Lett 339(2):159–166. https://doi.org/10.1016/j.canlet.2013.06.013
Li JL, Li ZG, Zheng WY, Li XH, Wang ZD, Cui YF, Jiang XM (2017) LncRNA-ATB: An indispensable cancer-related long noncoding RNA. Cell Prolif 50(6):e12381. https://doi.org/10.1111/cpr.12381
Zhou K, Zhang C, Yao H, Zhang XW, Zhou YX, Che YJ, Huang YL (2018) Knockdown of long non-coding RNA NEAT1 inhibits glioma cell migration and invasion via modulation of SOX2 targeted by miR-132. Mol Cancer 17(1):105. https://doi.org/10.1186/s12943-018-0849-2
Lu W, Zhang HH, Niu YQ, Wu YF, Sun WJ, Li HY, Kong JL, Ding KF, Shen HM et al (2017) Long non-coding RNA linc00673 regulated non-small cell lung cancer proliferation, migration, invasion and epithelial-mesenchymal transition by sponging miR-150-5p. Mol Cancer 16(1):118. https://doi.org/10.1186/s12943-017-0685-9
Bester AC, Lee JD, Chavez A, Lee YR, Nachmani D, Vora S, Victor J, Sauvageau M, Monteleone E et al (2018) An integrated genome-wide CRISPRa approach to functionalize lncRNAs in drug resistance. Cell 173:649–664. https://doi.org/10.1016/j.cell.2018.03.052
Jia P, Cai H, Liu XB, Chen JJ, Ma J, Wang P, Liu YH, Zheng J, Xue YX (2016) Long non-coding RNA H19 regulates glioma angiogenesis and the biological behavior of glioma-associated endothelial cells by inhibiting microRNA-29a. Cancer Lett 381(2):359–369. https://doi.org/10.1016/j.canlet.2016.08.009
Shi J, Dong B, Cao JC, Mao YM, Guan W, Peng Y, Wang SN (2017) Long non-coding RNA in glioma: signaling pathways. Oncotarget 8(16):27582–27592. https://doi.org/10.18632/oncotarget.15175
Bozgeyik E, Igci YZ, Sami JM, Arman K, Gurses SA, Bozgeyik I, Pala E, Yumrutas O, Temiz E, Igci M (2016) A novel variable exonic region and differential expression of LINC00663 non-coding RNA in various cancer cell lines and normal human tissue samples. Tumour Biol 37(7):8791–8798. https://doi.org/10.1007/s13277-015-4782-3
Igci YZ, Arslan A, Akarsu E, Erkilic S, Igci M, Oztuzcu S, Cengiz B, Gogebakan B, Cakmak EA, Demiryurek AT (2011) Differential expression of a set of genes in follicular and classic variants of papillary thyroid carcinoma. Endoc Pathol 22(2):86–96. https://doi.org/10.1007/s12022-011-9157-8
Chen LL (2016) Linking long noncoding RNA localization and function. Trends Biochem Sci 41:761–772. https://doi.org/10.1016/j.tibs.2016.07.003
Choe G, Horvath S, Cloughesy TF, Crosby K, Seligson D, Palotie A, Inge L, Smith BL, Sawyers CL, Mischel PS (2003) Analysis of the phosphatidylinositol 3’-kinase signaling pathway in glioblastoma patients in vivo. Cancer Res 63(11):2742–2746
Zhang T, Ji DF, Wang P, Liang D, Jin L, Shi HL, Liu XJ, Meng QM, Yu RT, Gao SF (2018) The atypical protein kinase RIOK3 contributes to glioma cell proliferation/survival, migration/invasion and the AKT/mTOR signaling pathway. Cancer Lett 415:151–163. https://doi.org/10.1016/j.canlet.2017.12.010
Reon BJ, Anaya J, Zhang Y, Mandell J, Purow B, Abounader R, Dutta A (2016) Expression of lncRNAs in low-grade gliomas and glioblastoma multiforme: an in silico analysis. PLoS Med 13(12):e1002192. https://doi.org/10.1371/journal.pmed.1002192
Weidle UH, Birzele F, Kollmorgen G, Rüger R (2017) Long non-coding RNAS and their role in metastasis. Cancer Genomics Proteom 14(3):143–160
Tan SK, Pastori C, Penas C, Komotar RJ, Ivan ME, Wahlestedt C, Ayad NG (2018) Serum long noncoding RNA HOTAIR as a novel diagnostic and prognostic biomarker in glioblastoma multiforme. Mol Cancer 17(1):74. https://doi.org/10.1186/s12943-018-0822-0
Kim SS, Harford JB, Moghe M, Rait A, Pirollo KF, Chang EH (2018) Targeted nanocomplex carrying siRNA against MALAT1 sensitizes glioblastoma to temozolomide. Nucleic Acids Res 46(3):1424–1440. https://doi.org/10.1093/nar/gkx1221
Zheng J, Liu XB, Wang P, Xue YX, Ma J, Qu CB, Liu YH (2016) CRNDE promotes malignant progression of glioma by attenuating miR-384/PIWIL4/STAT3 Axis. Mol Ther 24(7):1199–1215. https://doi.org/10.1038/mt.2016.71
Qian X, Li XJ, Shi ZM, Xia Y, Cai QS, Xu DQ, Tan L, Du LY, Zheng YH et al (2019) PTEN suppresses glycolysis by dephosphorylating and inhibiting autophosphorylated PGK1. Mol Cell 76(3):516–527. https://doi.org/10.1016/j.molcel.2019.08.006
Zhao HF, Wang J, Shao W, Wu CP, Chen ZP, Tony ST, Li WP (2017) Recent advances in the use of PI3K inhibitors for glioblastoma multiforme: current preclinical and clinical development. Mol Cancer 16(1):100. https://doi.org/10.1186/s12943-017-0670-3
Manning BD, Toker A (2017) AKT/PKB signaling: navigating the network. Cell 169:381–405. https://doi.org/10.1016/j.cell.2017.04.001
Carnero A, Paramio JM (2014) The PTEN/PI3K/AKT pathway in vivo, cancer mouse models. Front Oncol 4:252. https://doi.org/10.3389/fonc.2014.00252
Lin AF, Hu QS, Li CN, Xing Z, Ma GL, Wang C, Li J, Ye Y, Yao J et al (2017) The LINK-A lncRNA interacts with PtdIns(3,4,5)P3 to hyperactivate AKT and confer resistance to AKT inhibitors. Nat Cell Biol 19(3):238–251. https://doi.org/10.1038/ncb3473
Yang N, Chen JQ, Zhang H, Wang XM, Yao H, Peng Y, Zhang WG (2017) LncRNA OIP5-AS1 loss-induced microRNA-410 accumulation regulates cell proliferation and apoptosis by targeting KLF10 via activating PTEN/PI3K/AKT pathway in multiple myeloma. Cell Death Dis 8(8):e2975. https://doi.org/10.1038/cddis.2017.358
Jin Y, Feng SJ, Qiu S, Shao N, Zheng JH (2017) LncRNA MALAT1 promotes proliferation and metastasis in epithelial ovarian cancer via the PI3K-AKT pathway. Eur Rev Med Pharmacol Sci 21(14):3176–3184
Liu CH, Zhang Y, She XL, Fan L, Li PY, Feng JB, Fu HJ, Liu Q, Liu Q et al (2018) A cytoplasmic long noncoding RNA LINC00470 as a new AKT activator to mediate glioblastoma cell autophagy. J Hematol Oncol 11(1):77. https://doi.org/10.1186/s13045-018-0619-z
Acknowledgements
The authors acknowledge and appreciate their colleagues for their valuable efforts and comments regarding this paper. Special thanks to professor Hui Zeng, Liuluan Zhu and Institute of Infectious Diseases, Beijing Ditan Hospital, Capital Medical University, for their kind support.
Funding
The study was supported by Grants from the National Natural Science Foundation of China (No. 81572474).
Author information
Authors and Affiliations
Contributions
MP performed the experiments, all statistical analyses, and drafted the manuscript. JS contributed to design, data acquisition, interpretation and critically revised the manuscript. SY, HM and CH conducted the experiments. YW designed, supervised the experimental work and critically revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
The study protocol was approved by the Ethics Committee of Beijing Tiantan Hospital, Capital Medical University.
Informed Consent
Written informed consent was obtained from all patients prior to sample collection.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pan, M., Shi, J., Yin, S. et al. The Effect and Mechanism of LINC00663 on the Biological Behavior of Glioma. Neurochem Res 46, 1737–1746 (2021). https://doi.org/10.1007/s11064-021-03311-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-021-03311-3