Abstract
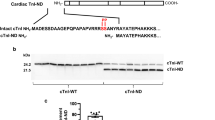
In this study we aimed to provide an in-depth proteomic analysis of differentially expressed proteins in the hearts of transgenic mouse models of pathological and physiological cardiac hypertrophy using tandem mass tag labeling and liquid chromatography tandem mass spectrometry. The Δ43 mouse model, expressing the 43-amino-acid N-terminally truncated myosin essential light chain (ELC) served as a tool to study the mechanisms of physiological cardiac remodeling, while the pathological hypertrophy was investigated in A57G (Alanine 57 → Glycine) ELC mice. The results showed that 30 proteins were differentially expressed in Δ43 versus A57G hearts as determined by multiple pair comparisons of the mutant versus wild-type (WT) samples with P < 0.05. The A57G hearts showed differential expression of nine mitochondrial proteins involved in metabolic processes compared to four proteins for ∆43 hearts when both mutants were compared to WT hearts. Comparisons between ∆43 and A57G hearts showed an upregulation of three metabolically important mitochondrial proteins but downregulation of nine proteins in ∆43 hearts. The physiological model of cardiac hypertrophy (∆43) showed no changes in the levels of Ca2+-binding proteins relative to WT, while the pathologic model (A57G) showed the upregulation of three Ca2+-binding proteins, including sarcalumenin. Unique differences in chaperone and fatty acid metabolism proteins were also observed in Δ43 versus A57G hearts. The proteomics data support the results from functional studies performed previously on both animal models of cardiac hypertrophy and suggest that the A57G- and not ∆43- mediated alterations in fatty acid metabolism and Ca2+ homeostasis may contribute to pathological cardiac remodeling in A57G hearts.



Similar content being viewed by others
References
Allard MF, Schonekess BO, Henning SL, English DR, Lopaschuk GD (1994) Contribution of oxidative metabolism and glycolysis to ATP production in hypertrophied hearts. Am J Physiol 267:H742–H750
Amsellem V et al. (2014) ICAM-2 regulates vascular permeability and N-cadherin localization through ezrin-radixin-moesin (ERM) proteins and Rac-1 signalling. Cell Commun Signal 12:12. doi:10.1186/1478-811X-12-12
Barefield D, Kumar M, de Tombe PP, Sadayappan S (2014) Contractile dysfunction in a mouse model expressing a heterozygous MYBPC3 mutation associated with hypertrophic cardiomyopathy. Am J Physiol Heart Circ Physiol 306:H807–815. doi:10.1152/ajpheart.00913.2013
Barger PM, Kelly DP (1999) Fatty acid utilization in the hypertrophied and failing heart: molecular regulatory mechanisms. Am J Med Sci 318:36–42
Bernardo BC, Weeks KL, Pretorius L, McMullen JR (2010) Molecular distinction between physiological and pathological cardiac hypertrophy: experimental findings and therapeutic strategies. Pharmacol Ther 128:191–227. doi:10.1016/j.pharmthera.2010.04.005
Bers DM, Guo T (2005) Calcium signaling in cardiac ventricular myocytes. Ann N Y Acad Sci 1047:86–98. doi:10.1196/annals.1341.008
Chang AN et al (2015) Constitutive phosphorylation of cardiac myosin regulatory light chain in vivo. J Biol Chem. doi:10.1074/jbc.M115.642165
Chen CP et al (2015a) In vivo roles for myosin phosphatase targeting subunit-1 phosphorylation sites T694 and T852 in bladder smooth muscle contraction. J Physiol 593:681–700. doi:10.1113/jphysiol.2014.283853
Chen TH et al (2015b) Neonatal death and heart failure in mouse with transgenic HSP60 expression. Biomed Res Int 2015:539805. doi:10.1155/2015/539805
Cho H et al (2014) Expression of stress-induced phosphoprotein1 (STIP1) is associated with tumor progression and poor prognosis in epithelial ovarian cancer. Genes Chromosom Cancer 53:277–288. doi:10.1002/gcc.22136
Chung E, Leinwand LA (2014) Pregnancy as a cardiac stress model. Cardiovasc Res 101:561–570. doi:10.1093/cvr/cvu013
Cui Z, Dewey S, Gomes AV (2011) Cardioproteomics: advancing the discovery of signaling mechanisms involved in cardiovascular diseases. Am J Cardiovasc Dis 1:274–292
Cui Z, Scruggs SB, Gilda JE, Ping P, Gomes AV (2014) Regulation of cardiac proteasomes by ubiquitination, SUMOylation, and beyond. J Mol Cell Cardiol 71:32–42. doi:10.1016/j.yjmcc.2013.10.008
Del Arco A, Agudo M, Satrústegui J (2000) Characterization of a second member of the subfamily of calcium-binding mitochondrial carriers expressed in human non-excitable tissues. Biochem J 345:725–732
den Boer ME, Dionisi-Vici C, Chakrapani A, van Thuijl AO, Wanders RJ, Wijburg FA (2003) Mitochondrial trifunctional protein deficiency: a severe fatty acid oxidation disorder with cardiac and neurologic involvement. J Pediatr 142:684–689. doi:10.1067/mpd.2003.231
Depre C et al (2006) Activation of the cardiac proteasome during pressure overload promotes ventricular hypertrophy. Circulation 114:1821–1828. doi:10.1161/CIRCULATIONAHA.106.637827
Dewey S, Lai X, Witzmann FA, Sohal M, Gomes AV (2013) Proteomic analysis of hearts from Akita mice suggests that increases in soluble epoxide hydrolase and antioxidative programming are key changes in early stages of diabetic cardiomyopathy. J Proteome Res 12:3920–3933. doi:10.1021/pr4004739
Doenst T et al (2010) Decreased rates of substrate oxidation ex vivo predict the onset of heart failure and contractile dysfunction in rats with pressure overload. Cardiovasc Res 86:461–470. doi:10.1093/cvr/cvp414
Facundo HT, Brainard RE, Watson LJ, Ngoh GA, Hamid T, Prabhu SD, Jones SP (2012) O-GlcNAc signaling is essential for NFAT-mediated transcriptional reprogramming during cardiomyocyte hypertrophy. Am J Physiol Heart Circ Physiol 302:H2122–H2130. doi:10.1152/ajpheart.00775.2011
Fan GC, Gregory KN, Zhao W, Park WJ, Kranias EG (2004) Regulation of myocardial function by histidine-rich, calcium-binding protein. Am J Physiol Heart Circ Physiol 287:H1705–H1711. doi:10.1152/ajpheart.01211.2003
Garrington TP, Johnson GL (1999) Organization and regulation of mitogen-activated protein kinase signaling pathways. Curr Opin Cell Biol 11:211–218
Geeves MA, Holmes KC (2005) The molecular mechanism of muscle contraction. Adv Protein Chem 71:161–193. doi:10.1016/S0065-3233(04)71005-0
Gomes AV et al. (2009) Contrasting proteome biology and functional heterogeneity of the 20 S proteasome complexes in mammalian tissues. Mol Cell Proteomics 8:302–315. doi:10.1074/mcp.M800058-MCP200
Gomes AV et al (2006) Mapping the murine cardiac 26S proteasome complexes. Circ Res 99:362–371. doi:10.1161/01.RES.0000237386.98506.f7
Harvey PA, Leinwand LA (2011) Cellular mechanisms of cardiomyopathy. J Cell Biol 194:355–365. doi:10.1083/jcb.201101100
Heineke J, Molkentin JD (2006) Regulation of cardiac hypertrophy by intracellular signalling pathways. Nat Rev Mol Cell Biol 7:589–600
Hernandez OM, Jones M, Guzman G, Szczesna-Cordary D (2007) Myosin essential light chain in health and disease. Am J Physiol Heart Circ Physiol 292:H1643–H1654. doi:10.1152/ajpheart.00931.2006
Hunter JJ, Chien KR (1999) Signaling pathways for cardiac hypertrophy and failure. N Engl J Med 341:1276–1283. doi:10.1056/NEJM199910213411706
Ibdah JA et al (2001) Lack of mitochondrial trifunctional protein in mice causes neonatal hypoglycemia and sudden death. J Clin Invest 107:1403–1409. doi:10.1172/JCI12590
Kaski JP et al (2009) Prevalence of sarcomere protein gene mutations in preadolescent children with hypertrophic cardiomyopathy. Circ Cardiovasc Genet 2:436–441. doi:10.1161/CIRCGENETICS.108.821314
Kazmierczak K et al. (2013) Discrete effects of A57G-myosin essential light chain mutation associated with familial hypertrophic cardiomyopathy American journal of physiology. Heart Circ Physiol 305:H575–H589
Kazmierczak K, Xu Y, Jones M, Guzman G, Hernandez OM, Kerrick WGL, Szczesna-Cordary D (2009) The role of the N-terminus of the myosin essential light chain in cardiac muscle contraction. J Mol Biol 387:706–725. doi:10.1016/j.jmb.2009.02.006
Kazmierczak K, Yuan C-C, Liang J, Huang W, Rojas AI, Szczesna-Cordary D (2014) Remodeling of the heart in hypertrophy in animal models with myosin essential light chain mutations. Front Physiology 5:353. doi:10.3389/fphys.2014.00353
Keller A, Nesvizhskii AI, Kolker E, Aebersold R (2002) Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 74:5383–5392
Lee W et al (2001) Different expressivity of a ventricular essential myosin light chain gene Ala57Gly mutation in familial hypertrophic cardiomyopathy. Am Heart J 141:184–189
Li Q, Sarna SK (2009) Nuclear myosin II regulates the assembly of preinitiation complex for ICAM-1 gene transcription. Gastroenterology 137(1051–1060):e1051–e1053. doi:10.1053/j.gastro.2009.03.040
Li Z, Adams RM, Chourey K, Hurst GB, Hettich RL, Pan C (2012) Systematic comparison of label-free, metabolic labeling, and isobaric chemical labeling for quantitative proteomics on LTQ Orbitrap Velos. J Proteome Res 11:1582–1590. doi:10.1021/pr200748h
Lorell BH, Carabello BA (2000) Left ventricular hypertrophy: pathogenesis, detection, and prognosis. Circulation 102:470–479. doi:10.1161/01.CIR.102.4.470
Maejima H, Nagashio R, Yanagita K, Hamada Y, Amoh Y, Sato Y, Katsuoka K (2014) Moesin and stress-induced phosphoprotein-1 are possible sero-diagnostic markers of psoriasis. PLoS One 9:e101773
Maron BJ (2002) The young competitive athlete with cardiovascular abnormalities: causes of sudden death, detection by preparticipation screening, and standards for disqualification. Card Electrophysiol Rev 6:100–103. doi:10.1023/A:1017903709361
McMullen JR et al (2007) Protective effects of exercise and phosphoinositide 3-kinase(p110alpha) signaling in dilated and hypertrophic cardiomyopathy. Proc Natl Acad Sci USA 104:612–617. doi:10.1073/pnas.0606663104
Megger DA et al (2014) Comparison of label-free and label-based strategies for proteome analysis of hepatoma cell lines. Biochim Biophys Acta 1844:967–976. doi:10.1016/j.bbapap.2013.07.017
Miller MS et al (2005) The essential light chain N-terminal extension alters force and fiber kinetics in mouse cardiac muscle. J Biol Chem 280:34427–34434. doi:10.1074/jbc.M508430200
Moolman JC, Corfield VA, Posen B, Ngumbela K, Seidman C, Brink PA, Watkins H (1997) Sudden death due to troponin T mutations. J Am Coll Cardiol 29:549–555
Morita H et al (2008) Shared genetic causes of cardiac hypertrophy in children and adults. N Engl J Med 358:1899–1908. doi:10.1056/NEJMoa075463
Muthu P et al (2011) Structural and functional aspects of the myosin essential light chain in cardiac muscle contraction. FASEB J 25:4394–4405. doi:10.1096/fj.11-191973
Nesvizhskii AI, Keller A, Kolker E, Aebersold R (2003) A statistical model for identifying proteins by tandem mass spectrometry. Anal Chem 75:4646–4658
Olson TM, Karst ML, Whitby FG, Driscoll DJ (2002) Myosin light chain mutation causes autosomal recessive cardiomyopathy with mid-cavitary hypertrophy and restrictive physiology. Circulation 105:2337–2340
Poetter K et al (1996) Mutations in either the essential or regulatory light chains of myosin are associated with a rare myopathy in human heart and skeletal muscle. Nat Genet 13:63–69
Powell SR, Herrmann J, Lerman A, Patterson C, Wang X (2012) The ubiquitin-proteasome system and cardiovascular disease. Prog Mol Biol Trans Sci 109:295–346. doi:10.1016/B978-0-12-397863-9.00009-2
Rauniyar N, Yates JR 3rd (2014) Isobaric labeling-based relative quantification in shotgun proteomics. J Proteome Res 13:5293–5309. doi:10.1021/pr500880b
Rayment I et al. (1993) Three-dimensional structure of myosin subfragment-1: a molecular motor. Science 261:50–58. doi:10.1126/science.8316857
Razeghi P, Young ME, Alcorn JL, Moravec CS, Frazier OH, Taegtmeyer H (2001) Metabolic gene expression in fetal and failing human heart. Circulation 104:2923–2931
Richard P et al. (2003) Hypertrophic cardiomyopathy: Distribution of disease genes, spectrum of mutations, and implications for a molecular diagnosis strategy. Circulation 107:2227–2232, and erratum (2004) 2109(2225):3258
Sanzen Y et al (2010) Functional proteomic analysis of experimental autoimmune myocarditis-induced chronic heart failure in the rat. Biol Pharm Bull 33:477–486
Sarikas A et al (2005) Impairment of the ubiquitin-proteasome system by truncated cardiac myosin binding protein C mutants. Cardiovasc Res 66:33–44. doi:10.1016/j.cardiores.2005.01.004
Shadforth IP, Dunkley TP, Lilley KS, Bessant C (2005) i-Tracker: for quantitative proteomics using iTRAQ. BMC Genom 6:145. doi:10.1186/1471-2164-6-145
Smith SH, Kramer MF, Reis I, Bishop SP, Ingwall JS (1990) Regional changes in creatine kinase and myocyte size in hypertensive and nonhypertensive cardiac hypertrophy. Circ Res 67:1334–1344
Spirito P, Bellone P, Harris KM, Bernabo P, Bruzzi P, Maron BJ (2000) Magnitude of left ventricular hypertrophy and risk of sudden death in hypertrophic cardiomyopathy. N Engl J Med 342:1778–1785
Stanley WC, Lopaschuk GD, Hall JL, McCormack JG (1997) Regulation of myocardial carbohydrate metabolism under normal and ischaemic conditions. Cardiovasc Res 33:243–257
Timson DJ (2003) Fine tuning the myosin motor: the role of the essential light chain in striated muscle myosin. Biochimie 85:639–645
Uehara T et al (1998) Myocardial glucose metabolism in patients with hypertrophic cardiomyopathy: assessment by F-18-FDG PET study. Ann Nucl Med 12:95–103
van der Velden J, Ho CY, Tardiff JC, Olivotto I, Knollmann BC, Carrier L (2015) Research priorities in sarcomeric cardiomyopathies. Cardiovasc Res 105:449–456. doi:10.1093/cvr/cvv019
VanBuren P, Waller GS, Harris DE, Trybus KM, Warshaw DM, Lowey S (1994) The essential light chain is required for full force production by skeletal muscle myosin. Proc Natl Acad Sci USA 91:12403–12407
Watson LJ et al (2010) O-linked beta-N-acetylglucosamine transferase is indispensable in the failing heart. Proc Natl Acad Sci USA 107:17797–17802. doi:10.1073/pnas.1001907107
Young ME et al (2007) Proposed regulation of gene expression by glucose in rodent heart. Gene Regul Syst Biol 1:251–262
Yuan CC et al (2015) Constitutive phosphorylation of cardiac myosin regulatory light chain prevents development of hypertrophic cardiomyopathy in mice. Proc Natl Acad Sci USA. doi:10.1073/pnas.1505819112
Yujiri T, Fanger GR, Garrington TP, Schlesinger TK, Gibson S, Johnson GL (1999) MEK kinase 1 (MEKK1) transduces c-Jun NH2-terminal kinase activation in response to changes in the microtubule cytoskeleton. J Biol Chem 274:12605–12610
Zhang YS et al (2015a) Nuclear cardiac myosin light chain 2 modulates NADPH oxidase 2 expression in myocardium: a novel function beyond muscle contraction. Basic Res Cardiol 110:38. doi:10.1007/s00395-015-0494-5
Zhang YS et al (2015b) A novel function of nuclear nonmuscle myosin regulatory light chain in promotion of xanthine oxidase transcription after myocardial ischemia/reperfusion. Free Radic Biol Med 83:115–128. doi:10.1016/j.freeradbiomed.2015.02.013
Acknowledgments
This work was supported by U.S. National Institutes of Health (NIH) Grants HL108343 and HL123255 (DSC); HL096819 (AVG), and the American Heart Association Grants 15PRE23020006 (CCY) and 15POST25080302 (ZZ).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gomes, A.V., Kazmierczak, K., Cheah, J.X. et al. Proteomic analysis of physiological versus pathological cardiac remodeling in animal models expressing mutations in myosin essential light chains. J Muscle Res Cell Motil 36, 447–461 (2015). https://doi.org/10.1007/s10974-015-9434-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10974-015-9434-0