Abstract
Human immunodeficiency virus type 1 (HIV-1) has been a global epidemic since 1983; yet, the virology and immunology related to HIV-1 remain elusive. Furthermore, as there is still no effective chemoprophylaxis or vaccine to treat patients with HIV-1, most research focuses on strategies to prevent HIV-1 infection, such as with antiviral drugs, novel therapeutics, or improved diagnostic kits. The HIV-1 Gag precursor protein (p55)—comprising the matrix (MA/p17), capsid (CA/p24), and nucleocapsid (NC/p7) protein domains—is the main structural HIV-1 protein, and is uniquely responsible for virion assembly within the virus life cycle. Recently, the immature and mature capsid structures were solved; however, the precursor protein structure is still unknown. Here, we expressed two subtypes of HIV-1 MA–CA stretch of the Gag protein, referred to as p41, in a bacterial expression system. We characterized the purified p41 protein, and showed its superior antigenicity over that of p24, highlighting the potential influence of the p17 domain on p24 structure. We further showed that p41 has good immunogenicity to induce an antibody response in mice. These results will aid future investigations into the HIV-1 capsid precursor structure, and potentially contribute to improving the design of diagnostic kits.
Similar content being viewed by others
1 Introduction
Human immunodeficiency virus type 1 (HIV-1) is a retrovirus that causes acquired immune deficiency syndrome (AIDS). Since 1983, HIV-1 has been a global epidemic, with more than 36.7 million people living with HIV-1 all over the world and about 2.1 million newly infected each year [1]. Currently, the primary treatment for HIV-1 infection is antiretroviral therapy (ART). Most of the antivirals against HIV-1 approved by the Food and Drug Administration are reverse transcriptase inhibitors (RTIs), protease inhibitors (PIs), and integrase inhibitors (INIs) [2]. Although suppressing HIV-1 replication can prolong a patient’s lifetime, these treatment options still cannot cure HIV-1 infection. Besides, with increasing rates of viral resistance to ART and the associated side effects of long-term therapy, it is now necessary to develop new antiviral drugs, novel therapeutics, and early detection techniques.
The HIV-1 Gag precursor protein is the main structural protein encoded by the gag gene. It is responsible for the assembly of virions and undergoes dramatic conformational changes during protein maturation [3,4,5]. Structurally, Gag is composed of several domains, including the matrix (MA/p17), capsid (CA/p24), spacer peptide 1 (SP1), nucleocapsid (NC/p7), SP2, and p6 domains [3,4,5,6,7].
In the immature virus particles, Gag molecules are packaged radially, such that the MA domain is bound to the inner leaflet of the viral membrane and the C-terminal region of Gag is oriented toward the center of the particle [8, 9]. The MA domain undergoes N-terminal myristoylation, which is critical targeting Gag to the assembly site [10, 11]. The MA domain also directs Gag to the plasma membrane for Gag oligomerization and for the incorporation of the Env trimeric envelope protein into virions [11, 12]. Cryo-electron microscopy (cryo-EM) and tomography (cryo-ET) analyses have shown that the Gag hexagonal lattice of the immature virus particles has a spacing of 8-nm, and that Gag hexamers are reinforced mainly by a six-helix bundle of SP1 peptides and CA C-terminal domain (CTD)–CTD interactions [8, 9].
When virions are released from the host cell plasma membrane, the immature particles undergo a dramatic morphological rearrangement over a cascade of proteolytic steps for maturation. A key proteolytic site is between the MA and CA domains, which triggers the structural rearrangement of CA to promote maturation. In these mature virus particles, the CA subunits are organized as a 2-dimensional lattice of hexamers, this time with a 10-nm spacing [13, 14]. The mature capsid is a cone-shaped shell that encompasses approximately 1,500 CA proteins, organized as approximately 250 hexamers and 12 pentamers [15]. The native capsid structure has been well defined by X-ray crystallography [16], with evidence for sixfold symmetry within the hexamers (intrahexamer), and threefold and twofold symmetry between neighboring hexamers (interhexamer). Yet, CA conformational variability exists [16], with subnanometer-resolution cryo-ET structures demonstrating the presence of hexameric and pentameric CA within intact HIV-1 mature particles; the hexamer structure is compatible with crystallography studies but the pentamer forms by means of different interfaces [17].
The fullerene cone capsid encloses the HIV-1 genome, the viral replicative enzymes and some accessory proteins [18,19,20], and therefore the stability of the capsid can influence aspects of HIV-1 infection, such as uncoating and reverse transcription [21, 22]. Indeed, some authors have suggested that the capsid may interact with numerous cellular factors within the host cell during HIV-1 infection, and this has made the CA domain a potential target for pharmacological inhibitors [23,24,25].
The structure of full-length Gag assembly is still unclear, primarily because of the intrinsic flexibility of Gag. Furthermore, although many high-resolution structures of CA in immature and mature virions have been solved [8, 9, 14,15,16,17, 26,27,28], the structural information of MA in the immature Gag lattice or the mature virions is still not yet available. This lack of knowledge makes it difficult for us to understand the underlying functions of the MA domain in HIV-1 life cycle and how its presence would be beneficial for the design of therapeutics or vaccines. In this study, we constructed and expressed two subtypes of the HIV-1 MA–CA stretch of the protein, hereafter referred to as p41 (subtype C MJ4-p41 and subtype B NL4-3-p41), using a bacterial expression system. The purified proteins were characterized by SDS–PAGE, western blotting, dot-blotting, ELISA, and mouse immunization. We found that p41 proteins showed good antigenicity and immunogenicity. These results will provide useful information for the further structural and functional exploration of capsid precursor proteins and help to improve the design of diagnostic kits based on the p41 protein.
2 Materials and Methods
2.1 Strains, Plasmids, Enzymes and Reagents
Escherichia coli DH5α and BL21 (DE3) were purchased from Invitrogen (Carlsbad, CA, USA). HIV-1 infection clone plasmids MJ4 and NL4-3 were kindly provided by Prof. Hua Zhu (Rutgers University, USA). pMD18-T plasmid, restriction endonucleases NdeI and XhoI, Taq-DNA polymerase, dNTP, PCR buffer and T4 DNA ligase were purchased from TAKARA Biotechnology Co., Ltd. (Dalian, China). Eight HIV-1-positive and three HIV-negative serum samples were provided by Beijing Wantai Biological Pharmacy Enterprise Co., Ltd. (Beijing, China), which is a high-tech enterprise engaged in the biological diagnosis reagents and vaccine development and production. Serum samples were from their in-house control panel for commercial HIV diagnostic kit, application of which was accordance with the standard detection guidelines. pTO-T7 plasmid, p24 protein and anti-p24 monoclonal antibodies were preserved in our lab.
2.2 Plasmid Construction
Using the full-length gene sequence of HIV-1 p55 (MJ4 and NL4-3), we designed oligonucleotide primers that included the restriction sites for NdeI and XhoI (underlined): MJ4-p41, F: 5′-CATATGGGTGCGAGAGCGTC-3′, R: 5′-CTCGAGCAATACTCTTGCTTTGTGGC-3′; for NL4-3-p41, F: 5′CATATGGGTGCGAGAGCGTC-3′, R: 5′-CTCGAGCAAAACTCTTGCTTTATGGC-3′.
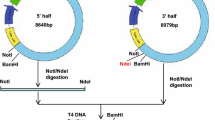
MJ4-p41 and NL4-3 genes were amplified using HIV-1 infection clone plasmids MJ4 and NL4-3 as template DNA. The purified PCR products were ligated into the pMD18-T plasmid. The pMD18-T-p41 plasmids and pTO-T7 plasmid were verified by double digestion with NdeI and XhoI restriction enzymes. The cleaved p41 and pTO-T7 were ligated by T4 DNA ligase to construct MJ4-p41 and NL4-3-p41 expression plasmids. Recombinant plasmids were verified by DNA sequencing.
2.3 Recombinant Protein Expression and Purification
The MJ4-p41 and NL4-3-p41 expression plasmids were transformed into E. coli BL21 (DE3) and cultured in LB medium containing kanamycin with shaking (180 rpm/min) at 37 °C until reaching an OD600 of 0.8. p41 protein expression was induced by 0.8 mM IPTG for 6 h with shaking at 37 °C. Strains were harvested by centrifugation (6000 rpm for 5 min), solubilized in 20 mM Tris–HCl pH 8.0, and sonicated. The insoluble material was removed by centrifugation and the supernatant was collected by 35% saturation ammonium sulfate precipitation. The precipitate was pelleted by centrifugation and resuspended for further purification on a HiTrap Ni2+ column using the AKTAPurifier System (GE Healthcare Life Sciences, PA, USA).
2.4 SDS–PAGE, Western Blotting and Dot-Blotting
The purified p41 proteins were analyzed by SDS–PAGE, western blotting and dot-blotting. Briefly, proteins were treated with 2 × reducing buffer (100 mM Tris–HCl, pH6.8, 200 mM β-mercaptoethanol, 4% SDS, 0.2% bromohenol blue and 20% glycerol) or non-reducing buffer (no β-mercaptoethanol), and loaded into the wells of 12% polyacrylamide gels. Gels were subjected to SDS–PAGE, and then stained with Coomassie blue or transferred onto nitrocellulose membranes (GE Healthcare, Little Chalfont, Buckinghamshire, UK) for immunoblotting. Membranes were blocked with 5% skim milk in Tris-buffered saline, and then incubated with anti-p24 monoclonal antibody (A10F9) [29] at room temperature for 2 h. The membranes were then washed and incubated with goat anti-mouse antibody conjugated with alkaline phosphatase (GAM-AP), at 1:2000 dilution for 1 h. Membranes were washed five times, and then incubated with chromogenic substrates.
For dot-blotting, 5-µL protein samples (0.5 mg/mL concentration) were absorbed onto nitrocellulose membranes. Membranes were air-dried, then blocked as described above, and incubated with HIV-1-positive serum (from the HIV-1 infected patients), -negative serum (from the HIV-1 uninfected individuals), or anti-p24 monoclonal antibodies. Secondary antibodies and chromogenic substrates were applied as above.
2.5 Indirect Enzyme-Linked Immunosorbent Assay (ELISA)
Wells of a 96-well ELISA plate were coated with 100 µL protein (1 µg/mL concentration) for 2 h at 37 °C. Wells were then blocked with blocking solution, and incubated with a serial dilution of immunized serum or anti-p24 mAbs for 1 h at 37 °C. The wells were then coated with horseradish peroxidase (HRP)-conjugated goat anti-mouse antibody (1:5000 dilution; made in-house) and incubated for 30 min at 37 °C. The OD was measured on an ELX800 Microplate Reader (Bio-Tek Instruments Inc., Winooski, VT).
Blocking ELISA was performed as described above, with modification. After coating and blocking the plate as above, the wells were then incubated with HIV-1-positive serum before adding the immunized serum. The wells were then incubated with goat anti-human antibody conjugated with HRP (1:5000 dilution; made in-house) for 30 min at 37 °C. The OD value was converted to percentage inhibition (%) using the following formula: Inhibition (%) = 100 − [(ODsample/ODcontrol × 100)], where the ODcontrol represents wells with PBS instead of immunized serum.
2.6 BALB/c Mice Immunization Assay
Four groups of female BALB/c mice (n = 3 per group) were respectively immunized with 100 µg MJ4-p24, MJ4-p41, NL4-3-p24 or NL4-3-p41 proteins subcutaneously on weeks 0, 1, 2 and 3. The proteins had been emulsified with complete Freund’s adjuvant in the prime immunization, and with incomplete Freund’s adjuvant in three booster immunizations. Immunized serum was collected for analysis. Antibody titers were measured using indirect ELISA (see above) using the respective proteins as coating antigens. The 4-week post-immunization serum was further evaluated by Blocking ELISA assay.
2.7 Statistical Analysis
ELISA assay data were processed with GraphPad Prism (Graphpad Software, CA, USA).
3 Results
3.1 Construction of Expression Plasmid
HIV-1 Gag full-length protein undergoes a cascade of proteolytic cleavage reactions to produce the mature and infectious virions (Fig. 1a). One of the most important proteolytic sites is between the MA and CA domains, which triggers the structural rearrangement of CA to promote the maturation process. Yet, despite the importance of this step, there is limited structural information pertaining to this MA–CA stretch of the protein. To investigate the MA–CA stretch, referred to p41, we first cloned and expressed the proteins for subsequent characterization. The p41 gene was derived from the HIV-1 infectious plasmids, MJ4 (subtype C) and NL4-3 (subtype B), and the amplified products were of approximately 1000 bp on agarose electrophoresis (Fig. 1b). The target genes were inserted into the pTO-T7 vector to construct the pTO-T7-p41 expression plasmid, and this was used to generate the p41 protein bearing a C-terminal His-tag. The resulting clones were designated MJ4-p41 and NL4-3-p41. The plasmids were verified by DNA sequencing.
HIV-1 p41 construction. a Schematic representation of HIV-1 Gag, p41, and p24. The five proteolytic sites are indicated by arrows in the full-length HIV-1 Gag protein. b Identification of PCR-amplified HIV-1 p41 genes by agarose electrophoresis. c SDS–PAGE and western blotting analysis of HIV-1 p41 protein expression. The anti-p24 monoclonal antibody (A10F9) was used as the detection antibody. The arrow indicates the p41 protein
3.2 Expression and Purification of Recombinant p41 Protein
The MJ4-p41 and NL4-3-p41 expression plasmids were transformed into E. coli BL21 (DE3), and p41 protein was expressed and characterized. The eluted fractions were pooled for further identification and analysis. As shown in Fig. 1c, an apparent molecular weight of ~ 41 kDa was confirmed by immunoblotting using anti-p24 antibody (A10F9).
The p41 protein was efficiently expressed in the soluble supernatant (data not shown). p41 was purified using the HiTrap Ni2+ affinity chromatography. As shown in Fig. 2a, the p41 protein was over 90% pure measured by Quantity One software, indicating a high-purity p41 protein. To our surprise, NL4-3-p41 migrated as a single band under reducing (with β-mercaptoethanol) and non-reducing (no β-mercaptoethanol) conditions; however, the MJ4-p41 protein migrated as either a dimer or a monomer under non-reducing conditions. We previously expressed and purified MJ4-p24 and NL4-3-p24 proteins, and similarly found that MJ4-p24 was a dimer in our crystal structure analysis [29]. Therefore, we speculate that the MJ4-p41 monomer may similarly spontaneously cross-link into homologous dimers by forming a disulfide bond between two Cys309 residues which are the same cysteines as p24.
Antigenicity analysis of HIV-1 p41. a SDS–PAGE and western blotting analysis of p41 and p24 proteins in reducing and non-reducing conditions using HIV-1 positive sera (from the HIV-1 infected patient no. 7). b Dot-blotting analysis of p41 and p24 proteins using eight samples of HIV-1 positive serum and five anti-p24 mAbs, respectively. Three HIV-1 negative serum samples (from the HIV-1 uninfected individuals) served as the negative control
3.3 Characterization of p41 Protein
We next sought to compare p41 and p24 for their reactivity and immunogenicity using HIV-1-positive serum and anti-p24 antibodies in western blotting and dot-blotting analyses. We found that p41 proteins showed better reactivity with the HIV-1-positive sera (from the HIV-1 infected patient no. 7) than did p24, irrespective of reducing or non-reducing conditions for western blotting (Fig. 2a). Using a dot-blotting assay, we found that MJ4-p41 and NL4-3-p41 reacted better than MJ4-p24 and NL4-3-p24 with all eight of the tested HIV-1-positive serum samples, consistent with the western blotting results (Fig. 2b). In addition, we tested five anti-p24 monoclonal antibodies (previously isolated from mice) [29], and found stronger immunoreactivity for p41 than p24, further supporting the above findings. None of the proteins reacted with the three control sera.
To further compare the p24 and p41 proteins, we evaluated the reactivities of p24 and p41 with a twofold serial dilution of four anti-p24 mAbs (starting at 1 µg/mL) using ELISA. The four mAbs named A10F9, 9E8, H5F4 and 16G12 were corresponding to Fig. 2b no. 13–16 mAbs which had reactivity with four proteins. The 50% maximal effective concentration (EC50) was calculated to directly reflect the binding activity. As shown in Fig. 3, the EC50 values for the A10F9, 9E8, H5F4 and 16G12 mAbs were 9.2, 11.3, 103, and 46.2 ng/mL, respectively, for NL4-3-p41, and 11.5, 33.3, > 200, and > 200 ng/mL, respectively, for NL4-3-p24. We found that the four anti-p24 mAbs showed lower EC50 values with NL4-3-p41 than with NL4-3-p24, suggesting that the p17 domain may influence the p24 epitope and improve its binding to NL4-3-p41. However, it is worth noting that the EC50 values were similar among the four mAbs with MJ4-p24 and MJ4-p41, with no significant differences in reactivity observed. This may be due to the aforementioned dimer formation for MJ4-p41 and MJ4-p24, which may affect antibody reactivity. To figure out whether the dimerization of MJ4 proteins is associated with the antigenicity difference between p41 and p24, we performed the EC50 test of MJ4 proteins in presence of reducing agent, β-mercaptoethanol. As shown in Fig. 4, the EC50 values for the A10F9, 9E8, H5F4 and 16G12 mAbs were 8.3, 7.2, 15.7 and 9.6 ng/mL, respectively, for MJ4-p41, and 19.8, 20.5, > 200, and 40.1 ng/mL, respectively, for MJ4-p24. The results demonstrated that four anti-p24 mAbs had lower EC50 values with monomeric MJ4-p41 than that of MJ4-p24 monomer, which was consistent to the EC50 discrepancy between NL4-3-p41 and NL4-3-p24 in monomeric form. Taken together, the p17 domain may pose effects on the antigenicity of p24 moiety in context of the full-length p41 .
Comparison of antigenicity between p41 and p24 proteins with four anti-p24 mAbs (corresponding to Fig. 2b no. 13–16 mAbs) using indirect ELISA: a A10F9, b 9E8, c H5F4 and d 16G12. The data were analyzed by GraphPad Prism software and the 50% maximal effective concentration (EC50) was calculated by a four-parameter logistic fit
Comparison of antigenicity between MJ4-p41 and MJ4-p24 in presence of reducing agent by indirect ELISA using four anti-p24 mAbs, a A10F9, b 9E8, c H5F4 and d 16G12. The data were analyzed by GraphPad Prism software and the 50% maximal effective concentration (EC50) was calculated by a four-parameter logistic fit
3.4 Immunization Analysis
We next evaluated the immunogenicity of the p41 and p24 proteins to immunize BALB/c mice. Three mice per group were inoculated four times with 100 µg of the recombinant MJ4-p24, MJ4-p41, NL4-3-p24 or NL4-3-p4 formulated with Freud’s adjuvant. The antibody response profiles showed that all four recombinant proteins elicited a high antibody titer for the 4-week post-immunization samples of approximately 7-log, 7.5-log, 6-log and 6.5-log for MJ4-p24, MJ4-p41, NL4-3-p24 and NL4-3-p41, respectively (Fig. 5a). Thus, both p41 and p24 proteins exhibited good immunogenicity to elicit an antibody response in BALB/c mice.
Immunogenicity assay. a Antibody titer for the immunized serum by indirect ELISA. The injection time points are indicated by black arrows. Both p24 and p41 proteins induced strong antibody titers as high as 106 at 4-weeks post-immunization. b Inhibition profiles using a Blocking ELISA assay. Serum taken from animals at 4-weeks post-immunization was blocked from binding to p41 or p24 by HIV-1 positive sera (from the HIV-1 infected patient no.7). The mean inhibition values are marked above the corresponding column at tenfold dilution of serum
Finally, we examined whether binding of the HIV-1-positive sera (from the HIV-1 infected patient no. 7) to p24 or p41 could be inhibited by the 4-week post-immunization serum. In the blocking ELISA assays, we found that antiserum from all four groups of mice could efficiently inhibit the binding of HIV-1-positive sera to p24 or p41 protein at tenfold dilution, with mean inhibition values of 64, 77.5, 67.5 and 76.5% for MJ4-p24, MJ4-p41, NL4-3-p24 and NL4-3-p41, respectively (Fig. 5b). Thus, antibodies raised via p24 and p41 immunization share similar epitopes with naturally acquired antibodies produced in response to HIV-1 infection.
4 Discussion
The HIV-1 p17 (MA) and p24 (CA) of the Gag protein are crucial in the production of infectious virions. Here, we constructed and tested the MA–CA stretch of the Gag protein—referred to as p41—for its antigenicity and capacity to elicit an immune response. We found that p41 offered better antigenicity over that of p24, suggestive of an effect of p17 on the structure of the p24 domain. Through immunization analysis, we also found that p41 can induce an antibody response in mice, raising epitopes similar to those naturally acquired in response to HIV-1 infection.
Recent advancements in cryo-EM and cryo-ET have significantly accelerated our understanding of the HIV-1 structure. The MA domain directs Gag proteins to their correct assembly sites, promotes Gag oligomerization, and recruits the envelope trimer protein into virions [4]. It is also part of the nuclear localization signal (NLS) and nuclear export signal (NES), which is important for HIV-1 infection and assembly [30, 31]. Mutations or deletions within this critical region interfere with virion assembly and result in the production of non-infectious virions [32,33,34,35]. Likewise, the CA domain has multifaceted roles in HIV-1 infection and assembly [36], and the detailed structural knowledge of its assembly has provided a potential target for pharmacological inhibitors [23,24,25]. Indeed, the structures of the immature or mature forms of the CA domain have been solved, and this has led to novel therapeutic antivirals against the capsid protein [37].
Our characterization of p41 highlighted the varied antigenicity of p24 against the mAbs or anti-serum when coupled to MA (as in p41) compared with its free form (p24). During the virion maturation process, it is believed that cleavage in the MA–CA junction triggers a conformation rearrangement in the CA domain, and this causes the new CA N-terminus to fold into a β-hairpin, stabilized by a buried salt bridge between the N-terminal proline and a conserved aspartate residue in helix 3 [38]. The formation of this β-hairpin promotes maturation of the capsid [39]. To our knowledge, the structure of the MA–CA stretch is different to that of the mature CA. Although various structures of MA and CA have been reported, the structure of the immature MA–CA p41 protein is not available. Solving the structure of p41 will uncover the conformational changes that occur during virion maturation.
As a first step toward this understanding, the present studies focused on the expression and characterization of two subtypes of recombinant HIV-1 p41 protein. We found that the p41 protein had better reactivity with HIV-1-positive serum than that of p24 protein, indicating that p41 has a remarkable advantage in detecting HIV-1-positive serum. This could be used to help improve diagnostic tests. Furthermore, in the ELISA assay of p41 against anti-p24 mAbs, we confirmed better antigenicity of p41 than p24, possibly due to influence of p17 on the p24 epitope. The immunization assay also showed high immunogenicity for p41 and the antiserum could block HIV-1-positive serum from binding with the p41 protein. Together, these results suggest that p41 may be a suitable, alternative candidate for HIV-1 diagnostics.
In conclusion, we show that the p41 (MA–CA) protein is highly reactive with HIV-1-positive serum and anti-p24 mAbs as compared with the p24 (CA alone) protein. This knowledge may offer an advantage in the design and development of diagnostic agents with enhanced reactivity. Future studies may also consider resolving the complex structure of the precursor MA–CA region of Gag with the anti-p41 antibody to gain further insight into this interaction.
References
WHO. Global Health Observatory (GHO) data of HIV/AIDS [EB/OL], http://www.who.int/gho/hiv/en/
Arts EJ, Hazuda DJ (2012) HIV-1 antiretroviral drug therapy. Cold Spring Harb Perspect Med 2:a007161
Ganser-Pornillos BK, Yeager M, Sundquist WI (2008) The structural biology of HIV assembly. Curr Opin Struct Biol 18:203–217
Bell NM, Lever AM (2013) HIV Gag polyprotein: processing and early viral particle assembly. Trends Microbiol 21:136–144
de Marco A, Muller B, Glass B, Riches JD, Krausslich HG, Briggs JA (2010) Structural analysis of HIV-1 maturation using cryo-electron tomography. PLoS Pathog 6:e1001215
Ganser-Pornillos BK, Yeager M, Pornillos O (2012) Assembly and architecture of HIV. Adv Exp Med Biol 726:441–465
Sundquist WI, Krausslich HG (2012) HIV-1 assembly, budding, and maturation. Cold Spring Harb Perspect Med 2:a006924
Briggs J, Riches J, Glass B, Bartonova V, Zanetti G, Kräusslich HG (2009) Structure and assembly of immature HIV. Proc Nat Acad Sci 106:11090–11095
Wright ER, Schooler JB, Ding HJ, Kieffer C, Fillmore C, Sundquist WI, Jensen GJ (2007) Electron cryotomography of immature HIV-1 virions reveals the structure of the CA and SP1 Gag shells. EMBO J 26:2218–2226
Saad JS, Miller J, Tai J, Kim A, Ghanam RH, Summers MF (2006) Structural basis for targeting HIV-1 Gag proteins to the plasma membrane for virus assembly. Proc Natl Acad Sci USA 103:11364–11369
Ghanam RH, Samal AB, Fernandez TF, Saad JS (2012) Role of the HIV-1 matrix protein in Gag intracellular trafficking and targeting to the plasma membrane for virus assembly. Front Microbiol 3:55
Tedbury PR, Novikova M, Ablan SD, Freed EO (2016) Biochemical evidence of a role for matrix trimerization in HIV-1 envelope glycoprotein incorporation. Proc Natl Acad Sci USA 113:E182-190
Briggs JA, Simon MN, Gross I, Krausslich HG, Fuller SD, Vogt VM, Johnson MC (2004) The stoichiometry of Gag protein in HIV-1. Nat Struct Mol Biol 11:672–675
Ganser-Pornillos BK, Cheng A, Yeager M (2007) Structure of full-length HIV-1 CA: a model for the mature capsid lattice. Cell 131:70–79
Zhao G, Perilla JR, Yufenyuy EL, Meng X, Chen B, Ning J, Ahn J, Gronenborn AM, Schulten K, Aiken C et al (2013) Mature HIV-1 capsid structure by cryo-electron microscopy and all-atom molecular dynamics. Nature 497:643–646
Gres AT, Kirby KA, KewalRamani VN, Tanner JJ, Pornillos O, Sarafianos SG (2015) X-ray crystal structures of native HIV-1 capsid protein reveal conformational variability. Science. https://doi.org/10.1126/science.aaa5936
Mattei S, Glass B, Hagen WJ, Krausslich HG, Briggs JA (2016) The structure and flexibility of conical HIV-1 capsids determined within intact virions. Science 354:1434–1437
Accola MA, Ohagen A, Gottlinger HG (2000) Isolation of human immunodeficiency virus type 1 cores: retention of Vpr in the absence of p6(gag). J Virol 74:6198–6202
Kotov A, Zhou J, Flicker P, Aiken C (1999) Association of Nef with the human immunodeficiency virus type 1 core. J Virol 73:8824–8830
Welker R, Hohenberg H, Tessmer U, Huckhagel C, Krausslich HG (2000) Biochemical and structural analysis of isolated mature cores of human immunodeficiency virus type 1. J Virol 74:1168–1177
Blair WS, Pickford C, Irving SL, Brown DG, Anderson M, Bazin R, Cao J, Ciaramella G, Isaacson J, Jackson L et al (2010) HIV capsid is a tractable target for small molecule therapeutic intervention. PLoS Pathog 6:e1001220
Shi J, Zhou J, Shah VB, Aiken C, Whitby K (2011) Small-molecule inhibition of human immunodeficiency virus type 1 infection by virus capsid destabilization. J Virol 85:542–549
Lee K, Mulky A, Yuen W, Martin TD, Meyerson NR, Choi L, Yu H, Sawyer SL, Kewalramani VN (2012) HIV-1 capsid-targeting domain of cleavage and polyadenylation specificity factor 6. J Virol 86:3851–3860
Valle-Casuso JC, Di Nunzio F, Yang Y, Reszka N, Lienlaf M, Arhel N, Perez P, Brass AL, Diaz-Griffero F (2012) TNPO3 is required for HIV-1 replication after nuclear import but prior to integration and binds the HIV-1 core. J Virol 86:5931–5936
Matreyek KA, Yucel SS, Li X, Engelman A (2013) Nucleoporin NUP153 phenylalanine-glycine motifs engage a common binding pocket within the HIV-1 capsid protein to mediate lentiviral infectivity. PLoS Pathog 9:e1003693
Schur FK, Hagen WJ, Rumlova M, Ruml T, Muller B, Krausslich HG, Briggs JA (2014) Structure of the immature HIV-1 capsid in intact virus particles at 8.8 Å resolution. Nature. https://doi.org/10.1038/nature13838
Wagner JM, Zadrozny KK, Chrustowicz J, Purdy MD, Yeager M, Ganser-Pornillos BK, Pornillos O (2016) Crystal structure of an HIV assembly and maturation switch. Elife 5. https://doi.org/10.7554/eLife.17063
Schur FK, Obr M, Hagen WJ, Wan W, Jakobi AJ, Kirkpatrick JM, Sachse C, Krausslich HG, Briggs JA (2016) An atomic model of HIV-1 capsid-SP1 reveals structures regulating assembly and maturation. Science. https://doi.org/10.1126/science.aaf9620
Gu Y, Cao F, Wang L, Hou W, Zhang J, Hew CL, Li S, Yuan YA, Xia N (2013) Structure of a novel shoulder-to-shoulder p24 dimer in complex with the broad-spectrum antibody A10F9 and its implication in capsid assembly. PLoS ONE 8:e61314
Fiorentini S, Giagulli C, Caccuri F, Magiera AK, Caruso A (2010) HIV-1 matrix protein p17: a candidate antigen for therapeutic vaccines against AIDS. Pharmacol Ther 128:433–444
Haffar OK, Popov S, Dubrovsky L, Agostini I, Tang H, Pushkarsky T, Nadler SG, Bukrinsky M (2000) Two nuclear localization signals in the HIV-1 matrix protein regulate nuclear import of the HIV-1 pre-integration complex. J Mol Biol 299:359–368
Saad JS, Loeliger E, Luncsford P, Liriano M, Tai J, Kim A, Miller J, Joshi A, Freed EO, Summers MF (2007) Point mutations in the HIV-1 matrix protein turn off the myristyl switch. J Mol Biol 366:574–585
Sanford B, Li Y, Maly CJ, Madson CJ, Chen H, Zhou Y, Belshan M (2014) Deletions in the fifth alpha helix of HIV-1 matrix block virus release. Virology 468–470:293–302
Facke M, Janetzko A, Shoeman RL, Krausslich HG (1993) A large deletion in the matrix domain of the human immunodeficiency virus gag gene redirects virus particle assembly from the plasma membrane to the endoplasmic reticulum. J Virol 67:4972–4980
Freed EO, Orenstein JM, Buckler-White AJ, Martin MA (1994) Single amino acid changes in the human immunodeficiency virus type 1 matrix protein block virus particle production. J Virol 68:5311–5320
Campbell EM, Hope TJ (2015) HIV-1 capsid: the multifaceted key player in HIV-1 infection. Nat Rev Microbiol 13:471–483
Waheed AA, Freed EO (2012) HIV type 1 Gag as a target for antiviral therapy. AIDS research human retroviruses 28:54–75
von Schwedler UK, Stemmler TL, Klishko VY, Li S, Albertine KH, Davis DR, Sundquist WI (1998) Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly. EMBO J 17:1555–1568
Gross I, Hohenberg H, Huckhagel C, Krausslich HG (1998) N-terminal extension of human immunodeficiency virus capsid protein converts the in vitro assembly phenotype from tubular to spherical particles. J Virol 72:4798–4810
Acknowledgements
The work was supported by the National Natural Science Foundation of China (Grant No. 81371818, 81671645).
Author information
Authors and Affiliations
Contributions
YG and NX conceived and designed the experiments. ZZ, LW, SB, JQ, HS, FH, SL and SG. performed the experiments. ZZ, LW, SL and NX analyzed the data. ZZ, SL and YG wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical Approval
All experimental procedures were approved by the Xiamen University Laboratory Animal Management Ethics Committee.
Rights and permissions
About this article
Cite this article
Zhang, Z., Wang, L., Bai, S. et al. Expression, Purification and Characterization of Hiv-1 Capsid Precursor Protein p41. Protein J 37, 194–202 (2018). https://doi.org/10.1007/s10930-018-9763-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-018-9763-0