Abstract
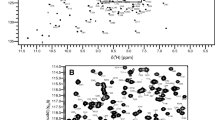
Human blood group A and B glycosyltransferases (GTA, GTB) are highly homologous glycosyltransferases. A number of high-resolution crystal structures is available showing that these enzymes convert from an open conformation into a catalytically active closed conformation upon substrate binding. However, the mechanism of glycosyltransfer is still under debate, and the precise nature as well as the time scales of conformational transitions are unknown. NMR offers a variety of experiments to shine more light on these unresolved questions. Therefore, in a first step we have assigned all methyl resonance signals in MILVA labeled samples of GTA and GTB, still a challenging task for 70 kDa homodimeric proteins. Assignments were obtained from methyl–methyl NOESY experiments, and from measurements of lanthanide-induced pseudocontact shifts (PCS) using high resolution crystal structures as templates. PCSs and chemical shift perturbations, induced by substrate analogue binding, suggest that the fully closed state is not adopted in the presence of lanthanide ions.








Similar content being viewed by others
References
Alfaro JA, Zheng RB, Persson M et al (2008) ABO(H) blood group A and B glycosyltransferases recognize substrate via specific conformational changes. J Biol Chem 283:10097–10108. https://doi.org/10.1074/jbc.M708669200
Amero C, Asunción Durá M, Noirclerc-Savoye M et al (2011) A systematic mutagenesis-driven strategy for site-resolved NMR studies of supramolecular assemblies. J Biomol NMR 50:229–236. https://doi.org/10.1007/s10858-011-9513-5
Atreya HS, Chary KVR (2001) Selective “unlabeling” of amino acids in fractionally 13C labeled proteins: an approach for stereospecific NMR assignments of CH3 groups in Val and Leu residues. J Biomol NMR 19:267–272. https://doi.org/10.1023/A:1011262916235
Ayala I, Sounier R, Usé N et al (2009) An efficient protocol for the complete incorporation of methyl-protonated alanine in perdeuterated protein. J Biomol NMR 43:111–119. https://doi.org/10.1007/s10858-008-9294-7
Chao F-A, Kim J, Xia Y et al (2014) FLAMEnGO 2.0: an enhanced fuzzy logic algorithm for structure-based assignment of methyl group resonances. J Magn Reson 245:17–23. https://doi.org/10.1016/j.jmr.2014.04.012
Cornilescu G, Marquardt JL, Ottiger M, Bax A (1998) Validation of protein structure from anisotropic carbonyl chemical shifts in a dilute liquid crystalline phase. J Am Chem Soc 120:6836–6837. https://doi.org/10.1021/ja9812610
Fiser A, Sali A (2003) ModLoop: automated modeling of loops in protein structures. Bioinformatics 19:2500–2501. https://doi.org/10.1093/bioinformatics/btg362
Gagnon SML, Meloncelli PJ, Zheng RB et al (2015) High resolution structures of the human ABO(H) blood group enzymes in complex with donor analogs reveal that the enzymes utilize multiple donor conformations to bind substrates in a stepwise manner. J Biol Chem 290:27040–27052. https://doi.org/10.1074/jbc.M115.682401
Gans P, Hamelin O, Sounier R et al (2010) Stereospecific isotopic labeling of methyl groups for NMR spectroscopic studies of high-molecular-weight proteins. Angew Chem Int Ed 49:1958–1962. https://doi.org/10.1002/anie.200905660
Gardner KH, Kay LE (1997) Production and incorporation of 15 N, 13 C, 2 H (1 H-δ1 methyl) isoleucine into proteins for multidimensional NMR studies. J Am Chem Soc 239:7599–7600. https://doi.org/10.1021/ja9706514
Gelis I, Bonvin AMJJ., Keramisanou D et al (2007) Structural basis for signal sequence recognition by the 204-kDa translocase motor Sec A determined by NMR. Cell 131:756–769. https://doi.org/10.1016/j.cell.2007.09.039
Goto NK, Gardner KH, Mueller GA et al (1999) A robust and cost-effective method for the production of Val, Leu, Ile (δ1) methyl-protonated 15N-, 13C-, 2H-labeled proteins. J Biomol NMR 13:369–374. https://doi.org/10.1023/A:1008393201236
Grimm LL, Weissbach S, Flügge F et al (2017) Protein NMR studies of substrate Binding to human blood group A and B glycosyltransferases. ChemBioChem 18:1260–1269. https://doi.org/10.1002/cbic.201700025
Han B, Liu Y, Ginzinger SW, Wishart DS (2011) SHIFTX2: significantly improved protein chemical shift prediction. J Biomol NMR 50:43–57. https://doi.org/10.1007/s10858-011-9478-4
Hearn VM, Smith ZG, Watkins WM (1968) An alpha-N-acetyl-D-galactosaminyltransferase associated with human blood-group A character. Biochem J 109:315–317
Hu W, Namanja AT, Wong S, Chen Y (2012) Selective editing of Val and Leu methyl groups in high molecular weight protein NMR. J Biomol NMR 53:113–124. https://doi.org/10.1007/s10858-012-9629-2
Isaacson RL, Simpson PJ, Liu M et al (2007) A new labeling method for methyl transverse relaxation-optimized spectroscopy NMR spectra of alanine residues. J Am Chem Soc 129:15428–15429. https://doi.org/10.1021/ja0761784
John M, Schmitz C, Ah YP et al (2007) Sequence-specific and stereospecific assignment of methyl groups using paramagnetic lanthanides. J Am Chem Soc 129:13749–13757. https://doi.org/10.1021/ja0744753
Lescanne M, Skinner SP, Blok A et al (2017) Methyl group assignment using pseudocontact shifts with PARAssign. J Biomol NMR. https://doi.org/10.1007/s10858-017-0136-3
Marcus SL, Polakowski R, Seto NOL et al (2003) A single point mutation reverses the donor specificity of human blood group B-synthesizing galactosyltransferase. J Biol Chem 278:12403–12405. https://doi.org/10.1074/jbc.M212002200
Monneau YR, Rossi P, Bhaumik A et al (2017) Automatic methyl assignment in large proteins by the MAGIC algorithm. J Biomol NMR 69:215–227. https://doi.org/10.1007/s10858-017-0149-y
Nicotra F, Panza L, Romaó A, Russo G (1992) Practical synthesis of disaccharide H. J Carbohydr Chem 11:397–399. https://doi.org/10.1080/07328309208018001
Palcic M, Heerze L, Pierce M, Hindsgaul O (1988) The use of hydrophobic synthetic glycosides as acceptors in glycosyltransferase assays. Glycoconj J 5:49–63. https://doi.org/10.1007/BF01048331
Pan YZ, Quade B, Brewer KD et al (2016) Sequence-specific assignment of methyl groups from the neuronal SNARE complex using lanthanide-induced pseudocontact shifts. J Biomol NMR 66:281–293. https://doi.org/10.1007/s10858-016-0078-1
Patenaude SI, Seto NOL, Borisova SN et al (2002) The structural basis for specificity in human ABO(H) blood group biosynthesis. Nat Struct Biol 9:685–690. https://doi.org/10.1038/nsb832
Pritišanac I, Degiacomi MT, Alderson TR et al (2017) Automatic assignment of methyl-NMR spectra of supramolecular machines using graph theory. J Am Chem Soc 139:9523–9533. https://doi.org/10.1021/jacs.6b11358
Proudfoot A, Frank AO, Ruggiu F et al (2016) Facilitating unambiguous NMR assignments and enabling higher probe density through selective labeling of all methyl containing amino acids. J Biomol NMR 65:15–27. https://doi.org/10.1007/s10858-016-0032-2
Rosenzweig R, Kay LE (2014) Bringing dynamic molecular machines into focus by methyl-TROSY NMR. Annu Rev Biochem 83:291–315. https://doi.org/10.1146/annurev-biochem-060713-035829
Sahakyan AB, Vranken WF, Cavalli A, Vendruscolo M (2011) Structure-based prediction of methyl chemical shifts in proteins. J Biomol NMR 50:331–346. https://doi.org/10.1007/s10858-011-9524-2
Schmitz C, Stanton-Cook MJ, Su XC et al (2008) Numbat: an interactive software tool for fitting ∆Χ-tensors to molecular coordinates using pseudocontact shifts. J Biomol NMR 41:179–189. https://doi.org/10.1007/s10858-008-9249-z
Schmitz C, Vernon R, Otting G et al (2012) Protein structure determination from pseudocontact shifts using ROSETTA. J Mol Biol 416:668–677. https://doi.org/10.1016/j.jmb.2011.12.056
Shannon RD (1976) Revised effective ionic radii and systematic studies of interatomic distances in halides and chalcogenides. Acta Crystallogr Sect A 32:751–767. https://doi.org/10.1107/S0567739476001551
Shoemaker GK, Kitova EN, Palcic MM, Klassen JS (2007) Equivalency of binding sites in protein-ligand complexes revealed by time-resolved tandem mass spectrometry. J Am Chem Soc 129:8674–8675
Sindhuwinata N, Munoz E, Munoz FJ et al (2010) Binding of an acceptor substrate analog enhances the enzymatic activity of human blood group B galactosyltransferase. Glycobiology 20:718–723 https://doi.org/10.1093/glycob/cwq019
Sindhuwinata N, Grimm LL, Weissbach S et al (2013) Thermodynamic signature of substrates and substrate analogs binding to human blood group B galactosyltransferase from isothermal titration calorimetry experiments. Biopolymers 99:784–795. https://doi.org/10.1002/bip.22297
Skinner SP, Moshev M, Hass MAS, Ubbink M (2013) PARAssign - Paramagnetic NMR assignments of protein nuclei on the basis of pseudocontact shifts. J Biomol NMR 55:379–389. https://doi.org/10.1007/s10858-013-9722-1
Soya N, Shoemaker GK, Palcic MM, Klassen JS (2009) Comparative study of substrate and product binding to the human ABO(H) blood group glycosyltransferases. Glycobiology 19:1224–1234. https://doi.org/10.1093/glycob/cwp114
Sprangers R, Kay LE (2007) Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature 445:618–622. https://doi.org/10.1038/nature05512
Sprangers R, Velyvis A, Kay LE (2007) Solution NMR of supramolecular complexes: providing new insights into function. Nat Methods 4:697–703. https://doi.org/10.1038/nmeth1080
Tugarinov V, Kay LE (2003) Ile, Leu, and Val methyl assignments of the 723-residue malate synthase g using a new labeling strategy and novel nmr methods. J Am Chem Soc 125:13868–13878. https://doi.org/10.1021/ja030345s
Tugarinov V, Kay LE (2004) An isotope labeling strategy for methyl TROSY spectroscopy. J Biomol NMR 28:165–172. https://doi.org/10.1023/B:JNMR.0000013824.93994.1f
Tugarinov V, Hwang PM, Ollerenshaw JE, Kay LE (2003) Cross-correlated relaxation enhanced 1H-13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J Am Chem Soc 125:10420–10428. https://doi.org/10.1021/ja030153x
Tugarinov V, Kay LE, Ibraghimov I, Orekhov VY (2005) High-resolution four-dimensional 1H-13C NOE spectroscopy using methyl-TROSY, sparse data acquisition, and multidimensional decomposition. J Am Chem Soc 127:2767–2775. https://doi.org/10.1021/ja044032o
Velyvis A, Schachman HK, Kay LE (2009) Assignment of Ile, Leu, and Val methyl correlations in supra-molecular systems: an application to aspartate transcarbamoylase. J Am Chem Soc 131:16534–16543. https://doi.org/10.1021/ja906978r
Velyvis A, Ruschak AM, Kay LE (2012) An economical method for production of 2H,13CH3-threonine for solution NMR studies of large protein complexes: application to the 670 kDa proteasome. PLoS ONE 7:1–8. https://doi.org/10.1371/journal.pone.0043725
Venditti V, Fawzi NL, Clore GM (2011) Automated sequence- and stereo-specific assignment of methyl-labeled proteins by paramagnetic relaxation and methyl-methyl nuclear overhauser enhancement spectroscopy. J Biomol NMR 51:319–328. https://doi.org/10.1007/s10858-011-9559-4
Vranken WF, Boucher W, Stevens TJ et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct Funct Genet 59:687–696. https://doi.org/10.1002/prot.20449
Weissbach S, Flügge F, Peters T (2018) Substrate binding drives active site closing of human blood group B galactosyltransferase as revealed by hot-spot labeling and NMR experiments. ChemBioChem. https://doi.org/10.1002/cbic.201800019
Wen J, Zhou P, Wu J (2012) Efficient acquisition of high-resolution 4-D diagonal-suppressed methyl-methyl NOESY for large proteins. J Magn Reson 218:128–132. https://doi.org/10.1016/j.jmr.2012.02.021
Xiao Y, Warner L, Latham MP et al (2015) Structure-based assignment of Ile, Leu and Val methyl groups in the active and inactive forms of the MAP Kinase ERK2. Biochemistry 54:4307–4319. https://doi.org/10.1021/acs.biochem.5b00506
Xu Y, Matthews S (2013) MAP-XSII: an improved program for the automatic assignment of methyl resonances in large proteins. J Biomol NMR 55:179–187. https://doi.org/10.1007/s10858-012-9700-z
Acknowledgements
This project has been funded by the German Research Council DFG (DFG Pe494/11−1). We acknowledge continuing support from the University of Lübeck. Professor Monica Palcic (University of Victoria, BC, Canada) is thanked for many stimulating discussions. We would like to thank Professor Marcellus Ubbink (University of Leiden, The Netherlands) and Dr. Alvaro Mallagaray (University of Lübeck) or stimulating discussions. We would also like to thank Dr. Thorsten Biet for excellent support with the NMR experiments, and Wilfried Hellebrandt for providing some of the synthetic oligosaccharides. We are grateful for support within the iNEXT Program (PID 1483 and PID 2254) giving us access to the NMR high-field facilities at the Bijvoet Center in Utrecht. In particular, we would like to thank Dr. Hans Wienk for performing the 4D methyl–methyl NOESY experiment in Utrecht.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Flügge, F., Peters, T. Complete assignment of Ala, Ile, Leu, Met and Val methyl groups of human blood group A and B glycosyltransferases using lanthanide-induced pseudocontact shifts and methyl–methyl NOESY. J Biomol NMR 70, 245–259 (2018). https://doi.org/10.1007/s10858-018-0183-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-018-0183-4