Abstract
The last resort for conservation of rare tree populations in refugial areas under high risk of climate driven extinction may be ex situ conservation and assisted translocation. Although such actions require detailed knowledge about the spatial scale and heterogeneity of the within-population distribution of genetic diversity, it is still unknown whether fine-scale spatial genetic structure (FSGS) is present in refugial populations of forest trees. In order to address this issue, we carried out the first whole-population genetic characterisation of a small and isolated refugial population of the IUCN red-listed Serbian spruce [Picea omorika (Panč.) Purk.] from the Balkans. All 418 adult individuals were georeferenced and genotyped at nuclear EST-SSRs and at a mitochondrial (mtDNA) locus. Spatial autocorrelation analyses provided only a simplified description of FSGS, which is concordant with findings in wind-pollinated species with limited seed dispersal. However, Bayesian analysis revealed three heterogeneous, highly differentiated (pairwise G’ ST > 0.3), and spatially localised sub-populations showing only partial overlap with the distribution of mtDNA haplotypes. Such complex structure in only 0.34 ha, resulting mainly from historical events, restrictions to gene flow and high local density, was undetected in previous work based on more traditional sampling schemes for population genetics surveys. We demonstrate the usefulness of sampling schemes leaning towards a whole-population genetic characterisation in mining the finest characteristics of FSGS, and argue that our understanding of genetic structuring in highly heterogeneous refugial regions at both macro- and micro-scales is still rather limited and often oversimplified. This has severe implications on conservation of plant biodiversity from these regions in terms of responses to global climate change.



Similar content being viewed by others
References
Aitken SN, Yeaman S, Holliday JA, Wang T, Curtis-McLane S (2008) Adaptation, migration or extirpation: climate change outcomes for tree populations. Evol Appl 1:95–111
Aleksić MJ (2008) Genetic structure of natural populations of Serbian spruce [Picea omorika (Panč.) Purk.]. Dissertation, University of Natural Resources and Applied Life Sciences, Austria
Aleksić MJ, Geburek T (2010) Mitochondrial DNA reveals complex genetic structuring in a stenoendemic conifer Picea omorika [(Panč.) Purk.] caused by its long persistence within the refugial Balkan region. Plant Syst Evol 285:1–11
Aleksić JM, Geburek T (2014) Quaternary population dynamics of an endemic conifer, Picea omorika, and their conservation implications. Cons Genet 15:87–107
Aleksić MJ, Schueler S, Mengl M, Geburek T (2009) EST-SSRs developed for other Picea species amplify in Picea omorika and reveal high genetic variation in two natural populations. Belg J Bot 142:89–95
Audigeos D, Brousseau L, Traissac S, Scotti - Saintagne C, Scotti I (2013) Molecular divergence in tropical tree populations occupying environmental mosaics. J Evol Biol 26:529–544
Austerlitz F, Mariette S, Machon N, Gouyon P-H, Godelle B (2000) Effects of colonization process on genetic diversity: differences between annual plant and tree species. Genetics 154:1309–1321
Brousseau L, Bonal D, Cigna J, Scotti I (2013) Highly local environmental variability promotes intrapopulation divergence of quantitative traits: an example from tropical rain forest trees. Ann Bot 112:1169–1179
Brousseau L, Postolache D, Lascoux M, Drouzas AD, Källman T, Leonarduzzi C et al (2016) Local adaptation in European firs assessed through extensive sampling across altitudinal gradients in southern Europe. PLoS One 11:e0158216
Christmas MJ, Breed MF, Lowe AJ (2016) Constraints to and conservation implications for climate change adaptation in plants. Cons Genet 17:305–320
Chybicki IJ, Dzialuk A (2014) Bayesian approach reveals confounding effects of population size and seasonality on outcrossing rates in a fragmented subalpine conifer. Tree Genet Genomes 10:1723–1737
Čolić DB (1987) Spontana obnova Pančićeve omorike (Picea omorika Panč.) posle požara. Zaštita prirode 40:37–56 (in Serbian with English summary)
Conifer Specialist Group (1998) Picea omorika. 2007 IUCN Red List of Threatened Species. http://www.iucnredlist.org/search/details.php/30313/summ
David A, Keathley D (1996) Inheritance of mitochondrial DNA in interspecific crosses of Picea glauca and Picea omorika. Can J For Res 26:428–432
de Lafontaine G, Ducousso A, Lefèvre S, Magnanou E, Petit RJ (2013) Stronger spatial genetic structure in recolonized areas than in refugia in the European beech. Mol Ecol 22:4397–4412
Dharmarajan G, Beatty WS, Rhodes OE (2013) Heterozygote deficiencies caused by a Wahlund effect: dispelling unfounded expectations. J Wildl Manage 77:226–234
Dizdarević M, Lakušić R, Grgić P, L Kutleša, Pavlović B, Jonlija R (1984) Ekološke osnove poimanja reliktnosti vrste Picea omorika Pančić. Bilten Društva ekologa Bosne i Herzegovine, Ser A 2:5–56 (in Bosnian with English abstract)
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214
Dyer RJ, Sork VL (2001) Pollen pool heterogeneity in shortleaf pine,Pinus echinataMill. Mol Ecol 10:859–866
Earl DA, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
El-Kassaby YA, Jaquish B (1996) Population density and mating pattern in western larch. J Hered 87:438–443
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fajardo A, Torres-Díaz C, Till-Bottraud I (2016) Disturbance and density-dependent processes (competition and facilitation) influence the fine-scale genetic structure of a tree species’ population. Ann Bot 117:67–77
FAO (2013) State of Mediterranean Forests 2013. FAO, Rome
Fukarek P (1951) Današnje rasprostranjenje Pančićeve omorike (Picea omorika Pančić) i neki podaci o njenim sastojinama. Godišnjak Biološkog Instituta u Sarajevu 3(1–2):141–198 (in Serbian with German summary)
Gajić M, Vilotić D, Karadžić D, Mihajlović L, Isajev V (1994) Serbian spruce–Picea omorika (Pančić) Purkyně on the territory of the National Park Tara. The National Park Tara, Bajina Bašta (in Serbian)
Gaut BS, Long AD (2003) The lowdown on linkage disequilibrium. Plant Cell 15:1502–1506
Grabherr RS, Gottfriend M, Pauli H (1994) Climatic effects on mountain plants. Nature 369:448–448
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Philos Trans R Soc Lond Ser B 351:1291–1298
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyze spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hardy OJ, Maggia L, Bandou E, Breyne P, Caron H, Chevallier M-H et al (2006) Fine-scale genetic structure and gene dispersal inferences in 10 neotropical tree species. Mol Ecol 15:559–571
Harms KE, Wright SJ, Calderón O, Hernández A, Herre EA (2000) Pervasive density-dependent recruitment enhances seedling diversity in a tropical forest. Nature 404:493–495
Hedrick PW (2005) A standardized genetic differentiation measure. Evol Int J org Evol 59:1633–1638
Hernández-Serrano A, Verdú M, González-Martínez SC, Pausas JG (2013) Fire structures pine serotiny at different scales. Am J Bot 100:2349–2356
Hewitt G (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978
Hijmans RJ, Guarino L, Marthur P (2012) DIVA-GIS version 7.5. http://www.diva-gis.org/
Hoban S, Schlarbaum S (2014) Optimal sampling of seeds from plant populations forex-situ conservation of genetic biodiversity, considering realistic population structure. Biol Cons 177:90–99
Hoban S, Strand A (2015) Ex situ seed collections will benefit from considering spatial sampling design and species’ reproductive biology. Biol Cons 187:182–191
Hossaert-McKey M, Valero M, Magda D, Jarry M, Cuguen J, Verne P (1996) The evolving genetic history of a population ofLathyrus sylvestris : evidence from temporal and spatial genetic structure. Evolution Int J org Evolution 50:1808–1821
Hubbell SP, Ahumada JA, Condit R, Foster RB (2001) Local neighborhood effects on long-term survival of individual trees in a Neotropical forest. Ecol Res 16:859–875
IPCC (2014) Summary for policymakers. In: Field CB, Barros VR, Dokken DJ et al (eds) Climate change 2014: impacts, adaptation, and vulnerability. Part A: global and sectoral aspects. Contribution of Working Group II to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change. Cambridge University Press, New York, pp 1–32
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Jansson S, Ingvarsson PK (2010) Cohort-structured tree populations. Heredity 105:331–332
Jevtić J (1960) Neka zapažanja o urodu semena Pančićeve omorike na Tari. Šumartsvo 13(1–2):79–84 (in Serbian with French summary)
Jump AS, Peñuelas J (2006) Genetic effects of chronic habitat fragmentation in a wind-pollinated tree. Proc Natl Acad Sci USA 103:8096–8100
Jump AS, Mátyás C, Peñuelas J (2009) The altitude-for-latitude disparity in the range retractions of woody species. Trends Ecol Evol 24:694–701
Jump AS, Huang TJ, Chou CH (2012) Rapid altitudinal migration of mountain plants in Taiwan and its implications for high altitude biodiversity. Ecography 35:204–210
Kalinowski ST (2005) hp-rare 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189
Kohlermann L (1950) Untersuchungen über die Windverbreitung der Früchte und Samen mitteleuropäischer Waldbäume. Forstwiss Cbl 69:606–624
Komlodi JM (1970) Studies on the vegetational history of Picea omorica Panc. on the Great Hungarian Plain. Annales Universitatis Scientiarum Budapestinensis de Rolando Eötvös Nominatae. Sectio biologica 12:143–156
Kremer A (1994) Diversité génétique et variabilité des caractéres phènotypiques chez les arbres forestiers. Genet Sel Evol 26(Suppl 1):105s–123s
Kremer A, Ronce O, Robledo-Arnuncio JJ, Guillaume F, Bohrer G et al (2012) Long-distance gene flow and adaptation of forest trees to rapid climate change. Ecol Lett 15:378–392
Kuittinen H, Savolainen O (1992) Picea omorika is a self-fertile but outcrossing conifer. Heredity 68:183–187
Lefèvre F, Koskela J, Hubert J, Kraigher H, Longauer R, Olrik DC, et al (2013) Dynamic conservation of forest genetic resources in 33 European countries. Cons Biol 27:373–384
Lenoir J, Gégout JC, Marquet PA, De Ruffray P, Brisse H (2008) A significant upward shift in plant species optimum elevation during the 20th century. Science 320:1768–1771
Leonardi S, Piovani P, Scalfi M, Piotti A, Giannini R, Menozzi P (2012) Effect of habitat fragmentation on the genetic diversity and structure of peripheral populations of beech in Central Italy. J Hered 103:408–417
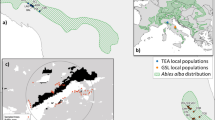
Liepelt S, Cheddadi R, de Beaulieu JL, Fady B, Gömöry D, Hussendörfer E, Konnert M, Litt T, Longauer R, Terhürne-Berson R, Ziegenhagen B (2009) Postglacial range expansion and its genetic imprints in Abies alba (Mill.)—A synthesis from palaeobotanic and genetic data. Rev Palaeobot Palyno 153:139–149
Litrico I, Ronfort J, Verlaque R, Thompson JD (2005) Spatial structure of genetic variation and primary succession in the pioneer tree species Antirhea borbonica on La Réunion. Mol Ecol 14:1575–1584
Loarie SR, Duffy PB, Hamilton H, Asner GP, Field CB, Ackerly DD (2009) The velocity of climate change. Nature 462:1052–1056
Lockwood JD, Aleksić MJ, Zou J, Wang J, Liu J, Renner SS (2013) A new phylogeny for the genus Picea from plastid, mitochondrial and nuclear sequences. Mol Phylogenet Evol 69:717–727
Loiselle BA, Sork VL, Nason J, Graham C (1995) Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae). Am J Bot 82:1420–1425
Malcolm JR, Markham A, Neilson RP, Garaci M (2002) Estimated migration rates under scenarios of global climate change. J Biogeogr 29:835–849
McCauley DE (1997) The relative contributions of seed and pollen movement to the local genetic structure ofSilene alba. J Hered 88:257–263
Morgante M, Vendramin GG, Rossi P (1991) Effects of stand density on outcrossing rate in two Norway spruce (Picea abies) populations. Can J Bot 69:2704–2708
Moritz C (1994) Defining ‘evolutionary significant units’ for conservation. Trends Ecol Evol 9:373–375
Oddou-Muratorio S, Klein EK, Demesure-Musch B, Austerlitz F (2006) Real-time patterns of pollen flow in the wild-service tree, Sorbus torminalis (Rosaceae). III. Mating patterns and the ecological maternal neighborhood. Am J Bot 93:1650–1659
Orr HA (2005) The genetic history of adaptation: a brief history. Nat Rev Genet 6:119–127
Parmesan C, Hanley ME (2015) Plants and climate change: complexities and surprises. Ann Bot 116:849–864
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinform 28 :2537–2539
Petit RJ, Hampe A (2006) Some evolutionary consequences of being a tree. Ann Rev Ecol Evol Syst 37:187–214
Petit RJ, El Mousadik A, Pons O (1998) Identifying populations for conservation on the basis of genetic markers. Cons Biol 12:844–855
Petit RJ, Hampe A, Cheddadi R (2005) Climate changes and tree phylogeography in the Mediterranean. Taxon 54:877–885
Piotti A, Leonardi S, Piovani P, Scalfi M, Menozzi P (2009) Spruce colonization at treeline: where do those seeds come from? Heredity 103:136–145
Piotti A, Leonardi S, Buiteveld J, Geburek T, Gerber S, Kramer K, et al (2012) Comparison of pollen gene flow among four European beech (Fagus sylvatica L.) populations characterized by different management regimes. Heredity 108:322–331
Piotti A, Leonardi S, Heuertz M, Buiteveld J, Geburek T, Gerber S et al (2013) Within-population genetic structure in beech (Fagus sylvatica L.) stands characterized by different disturbance histories: does forest management simplify population substructure? PLoS One 8:e73391
Piry S, Luikar G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90:502–503
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ran JH, Shen TT, Liu WJ, Wang PP, Wang XQ (2015) Mitochondrial introgression and complex biogeographic history of the genus Picea. Mol Phylogenet Evol 93:63–76
Ravazzi C (2002) Late Quaternary history of spruce in southern Europe. Rev Palaeobot Palynol 120:131–177
Richards CM, Antolin MF, Reilley A, Poole J, Walters C (2007) Capturing genetic diversity of wild populations for ex situ conservation: Texas wild rice (Zizania texana) as a model. Genet Resour Crop Evol 54:837–848
Rieseberg LH (1995) The role of hybridization in evolution: old wine in new skins. Am J Bot 82:944–953
Robledo-Arnuncio JJ, Collada C, Alia R, Gil L (2005) Genetic structure of montane isolates of Pinus sylvestris L. in a Mediterranean refugial area. J Biogeogr 32:595–605
Rousset F (2008) GENEPOP’007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Resour 8:103–106
Rungis D, Bérubé Y, Zhang J, Ralph S, Ritland CE, Ellis BE et al (2004) Robust simple sequence repeat markers for spruce (Picea spp.) from expressed sequence tags. Theor Appl Genet 109:1283–1294
Sagnard F, Oddou-Muratorio S, Pichot C, Vendramin GG, Fady B (2011) Effects of seed dispersal, adult tree and seedling density on the spatial genetic structure of regeneration at fine temporal and spatial scales. Tree Genet Genomes 7 :37–48
Sala OE, Chapin FS, Armesto JJ, Berlow E, Bloomfield J, Dirzo R et al (2000) Global biodiversity scenarios for the year 2100. Science 287:1770–1774
Scotti I, Gugerli F, Pastorelli R, Sebastiani F, Vendramin GG (2008) Maternally and paternally inherited molecular markers elucidate population patterns and inferred dispersal processes on a small scale within a subalpine stand of Norway spruce (Picea abies [L.] Karst.). For Ecol Manag 255:3806–3812
Scotti I, González-Martínez SC, Budde KB, Lalagüe H (2016) Fifty years of genetic studies: what to make of the large amounts of variation found within populations? Ann For Sci 73:69–75
Slavov GT, Leonardi S, Adams WT, Strauss SH, DiFazio SP (2010) Population substructure in continuous and fragmented stands of Populus trichocarpa. Heredity 105:348–357
Steinitz O, Troupin D, Vendramin GG, Nathan R (2011) Genetic evidence for a Janzen–Connell recruitment pattern in reproductive offspring of Pinus halepensistrees. Mol Ecol 20:4152–4164
Stewart JR, Lister AM, Barnes I, Dalén L (2010) Refugia revisited: individualistic responses of species in space and time. Proc R Soc B 277:661–671
Thomas CD, Cameron A, Green RE, Bakkenes M, Beaumont LJ, Collingham YC et al (2004) Extinction risk from climate change. Nature 427:145–148
Thuiller W (2007) Biodiversity: climate change and the ecologist. Nature 448:550–552
Trapnell DW, Hamrick JL, Ishibashi CD, Kartzinel TR (2013) Genetic inference of epiphytic orchid colonization; it may only take one. Mol Ecol 22:3680–3692
Troupin D, Nathan R, Vendramin GG (2006) Analysis of spatial genetic structure in an expanding Pinus halepensis population reveals development of fine-scale genetic clustering over time. Mol Ecol 15:3617–3630
Tucić B, Stojković B (2001) Shade avoidance syndrome in Picea omorika seedlings: a growth-room experiment. J Evol Biol 14:444–455
Unger GM, Konrad H, Geburek T (2011) Does spatial genetic structure increase with altitude? An answer from Picea abies in Tyrol, Austria. Plant Syst Evol 292:133–141
Vekemans X, Hardy OJ (2004) New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13:921–935
Vidaković M (1991) Conifers, morphology and variation, 2nd edn. Grafički zavod Hrvatske, Zagreb
Wahlund S (1928) Zusammensetzung von Populationen und Korrelationserscheinungen von Standpunkt der Vererbungslehre aus betrachtet. Hereditas 11:65–106 [English translation In: Weiss KM, Ballanoff PA (eds) Demographic genetics. Dowden, Hutchinson and Ross Inc, Stroudsburg, Pennsylvania, USA, pp 224–263]
Walther GR, Post E, Convey P, Menzel A, Parmesan C, Beebee TJ, et al (2002) Ecological responses to recent climate change. Nature 416:389–395
Wright SJ, Muller-Landau HC, Calderón O, Hernandéz A (2005) Annual and spatial variation in seedfall and seedling recruitment in a neotropical forest. Ecology 86:848–860
Acknowledgements
This work was financially supported by the Republic of Austria, Bioversity International, Research Grant 173005 of the Ministry of Education and Science of the Republic of Serbia, the COST Action FP1202 and the Italian MIUR project “Biodiversitalia” (RBAP10A2T4). The authors would like to thank M. Mengl, L. Weißenbacher, M. Josipović, D. Milekić, C. Leonarduzzi, C Bondani and Milovan for assistance in the field, and N. Kuzmanović for extracting climate data. All samples were collected with the permission of the Ministry of Energy, Development and Environmental Protection of the Republic of Serbia, and support of the Tara National Park, Republic of Serbia.
Author information
Authors and Affiliations
Corresponding author
Additional information
J. M. Aleksić and A. Piotti have contributed equally to the work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aleksić, J.M., Piotti, A., Geburek, T. et al. Exploring and conserving a “microcosm”: whole-population genetic characterization within a refugial area of the endemic, relict conifer Picea omorika . Conserv Genet 18, 777–788 (2017). https://doi.org/10.1007/s10592-017-0926-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-017-0926-x