Abstract
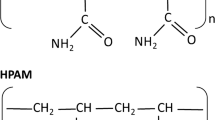
The increasing usage of partially hydrolyzed polyacrylamide (HPAM) in oilfields as a flooding agent to enhance oil recovery at so large quantities is an ecological hazard to the subsurface ecosystem due to persistence and inertness. Biodegradation of HPAM is a potentially promising strategy for dealing with this problem among many other methods available. To understand the responsible microorganisms and mechanism of HPAM biodegradation under anaerobic conditions, an enrichment culture from production waters of oil reservoirs were established with HPAM as the sole source of carbon and nitrogen incubated for over 328 days, and analyzed using both molecular microbiology and chemical characterization methods. Gel permeation chromatography, High-pressure liquid chromatography and Fourier-transformed infrared spectroscopy results indicated that, after 328 days of anaerobic incubation, some of the amide groups on HPAM were removed and released as ammonia/ammonium and carboxylic groups, while the carbon backbone of HPAM was converted to smaller polymeric fragments, including oligomers and various fatty acids. Based on these results, the biochemical process of anaerobic biodegradation of HPAM was proposed. The phylogenetic analysis of 16S rRNA gene sequences retrieved from the enrichments showed that Proteobacteria and Planctomycetes were the dominant bacteria in the culture with HPAM as the source of carbon and nitrogen, respectively. For archaea, Methanofollis was more abundant in the anaerobic enrichment. These results are helpful for understanding the process of HPAM biodegradation and provide significant insights to the fate of HPAM in subsurface environment and for possible bioremediation.






Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bao M, Chen Q, Li Y, Jiang G (2010) Biodegradation of partially hydrolyzed polyacrylamide by bacteria isolated from production water after polymer flooding in an oil field. J Hazard Mater 184:105–110
Bauer M et al (2006) Whole genome analysis of the marine Bacteroidetes ‘Gramella forsetii’ reveals adaptations to degradation of polymeric organic matter. Environ Microbiol 8:2201–2213
Chang LL, Raudenbush DL, Dentel SK (2001) Aerobic and anaerobic biodegradability of a flocculant polymer. Water Sci Technol 44:461–468
Cheng L, Shi S, Li Q, Chen J, Zhang H, Lu Y (2013) Progressive degradation of crude oil n -alkanes coupled to methane production under mesophilic and thermophilic conditions. PLoS ONE 9:e113253–e113253
Dai X, Luo F, Yi J, He Q, Dong B (2014) Biodegradation of polyacrylamide by anaerobic digestion under mesophilic condition and its performance in actual dewatered sludge system. Bioresour Technol 153:55–61
Dai X, Luo F, Zhang D, Dai L, Chen Y, Dong B (2015) Waste-activated sludge fermentation for polyacrylamide biodegradation improved by anaerobic hydrolysis and key microorganisms involved in biological polyacrylamide removal. Sci Rep 5:11675. https://doi.org/10.1038/srep11675
Deshmukh SR, Chaturvedi PN, Singh RP (1985) The turbulent drag reduction by graft copolymers of guargum and polyacrylamide. J Appl Polym Sci 30:4013–4018
Dianou D, Miyaki T, Asakawa SH, Nagaoka K, Oyaizu H (2001) Methanoculleus chikugoensis sp nov., a novel methanogenic archaeon isolated from paddy field soil in Japan, and DNA-DNA hybridization among Methanoculleus species. Int J Syst Evolut Microbiol 51:1663–1669
Elmamouni R, Frigon JC, Hawari J, Marroni D, Guiot SR (2002) Combining photolysis and bioprocesses for mineralization of high molecular weight polyacrylamides. Biodegradation 13:221–227
Grula MM, Huang ML, Sewell G, Grula MM, Huang ML, Sewell G (1994) Interactions of certain polyacrylamides with soil bacteria. Soil Sci 158:291–300. https://doi.org/10.1097/00010694-199410000-00009
Haveroen ME, Mackinnon MD, Fedorak PM (2005) Polyacrylamide added as a nitrogen source stimulates methanogenesis in consortia from various wastewaters. Water Res 39:3333–3341
Imachi H, Sakai S, Nagai H, Yamaguchi T, Takai K (2009) Methanofollis ethanolicus sp. nov., an ethanol-utilizing methanogen isolated from a lotus field. Int J Syst Evolut Microbiol 59:800–805
Kaşgöz H, Özgümüş S, Orbay M (2001) Preparation of modified polyacrylamide hydrogels and application in removal of Cu(II) ion. Polymer 42:7497–7502
Kay-Shoemake JL, Watwood ME, Lentz RD, Sojka RE (1998a) Polyacrylamide as an organic nitrogen source for soil microorganisms with potential effects on inorganic soil nitrogen in agricultural soil. Soil Biol Biochem 30:1045–1052
Kay-Shoemake JL, Watwood ME, Sojka RE, Lentz RD (1998b) Polyacrylamide as a substrate for microbial amidase in culture and soil. Soil Biol Biochem 30:1647–1654
Kim DY, Rhee YH (2003) Biodegradation of microbial and synthetic polyesters by fungi. Appl Microbiol Biotechnol 61:300–308
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Li CY, Zhang D, Li XX et al (2015) The biofilm property and its correlationship with high-molecular-weight polyacrylamide degradation in a water injection pipeline of Daqing oilfield. J Hazard Mater 304:388–399
Liu L, Wang Z, Lin K, Cai W (2012) Microbial degradation of polyacrylamide by aerobic granules. Environ Technol 33:1049–1054
Liu J, Ren J, Xu R, Yu B, Wang J (2016) Biodegradation of partially hydrolyzed polyacrylamide by immobilized bacteria isolated from HPAM-containing wastewater. Environ Progr Sustain Energy 35:1344–1352. https://doi.org/10.1002/ep.12354
Ma F, Wei L, Wang L, Chang CC (2008) Isolation and identification of the sulphate-reducing bacteria strain H1 and its function for hydrolysed polyacrylamide degradation. Int J Biotechnol 10:55–63
Mcginnis S, Madden TL (2004) BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucl Acids Res 32:20–25
Mckew BA, Dumbrell AJ, Taylor JD, Mcgenity TJ, Underwood GJ (2013) Differences between aerobic and anaerobic degradation of microphytobenthic biofilm-derived organic matter within intertidal sediments. FEMS Microbiol Ecol 84:495–509
Mnif S, Bru-Adan V, Godon JJ, Sayadi S, Chamkha M (2013) Characterization of the microbial diversity in production waters of mesothermic and geothermic Tunisian oilfields. J Basic Microbiol 53:45–61
Nakamiya K, Kinoshita S (1995) Isolation of polyacrylamide-degrading bacteria. J Ferment Bioeng 80:418–420
Pham VD et al (2009) Characterizing microbial diversity in production water from an Alaskan mesothermic petroleum reservoir with two independent molecular methods. Environ Microbiol 11:176–187
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sang G, Pi Y, Bao M, Li Y, Lu J (2015) Biodegradation for hydrolyzed polyacrylamide in the anaerobic baffled reactor combined aeration tank. Ecol Eng 84:121–127
Savage KN et al (2010) Biodegradation of low-molecular-weight alkanes under mesophilic, sulfate-reducing conditions: metabolic intermediates and community patterns. FEMS Microbiol Ecol 72:485–495
Sewell GW (1987) Interactions of polyacrylamides used for enhanced oil recovery and reservoir isolates of the sulfate-reducing bacterium Desulfovibrio. Oklahoma State University, Stillwater
Silva TR, Verde LCL, Neto EVS, Oliveira VM (2013) Diversity analyses of microbial communities in petroleum samples from Brazilian oil fields. Int Biodeterior Biodegrad 81:57–70
Song W, Zhang Y, Gao Y et al (2017) Cleavage of the main carbon chain backbone of high molecular weight polyacrylamide by aerobic and anaerobic biological treatment. Chemosphere 189:277–283
Strous M et al (1999) Missing lithotroph identified as new planctomycete. Nature 400:446–449
Suzuki J, Hukushima K, Suzuki S (1978) Effect of ozone treatment upon biodegradability of water-soluble polymers. Environ Sci Technol 12:1180–1183
Takai K, Horikoshi K (2000) Rapid detection and quantification of members of the archaeal community by quantitative PCR using fluorogenic probes. Appl Environ Microbiol 66:5066–5072
Tyl RW, Friedman MA (2003) Effects of acrylamide on rodent reproductive performance. Reprod Toxicol 17:1–13
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang LY, Gao CX, Mbadinga SM, Zhou L, Liu JF, Gu JD, Mu BZ (2011) Characterization of an alkane-degrading methanogenic enrichment culture from production water of an oil reservoir after 274 days of incubation. Int Biodeterior Biodegrad 65:444–450
Wen Q, Chen Z, Zhao Y, Zhang H, Feng Y (2010) Biodegradation of polyacrylamide by bacteria isolated from activated sludge and oil-contaminated soil. J Hazard Mater 175:955–959
Yu Y, Breitbart M, Mcnairnie P, Rohwer F (2006) FastGroupII: a web-based bioinformatics platform for analyses of large 16S rDNA libraries. BMC Bioinform 7:213–219
Yu F, Fu R, Xie Y, Chen W (2015) Isolation and characterization of polyacrylamide-degrading bacteria from dewatered sludge. Int J Environ Res Public Health 12:4214–4230
Zhao X, Liu L, Wang Y, Dai H, Wang D, Cai H (2008) Influences of partially hydrolyzed polyacrylamide (HPAM) residue on the flocculation behavior of oily wastewater produced from polymer flooding. Sep Purif Technol 62:199–204
Zhao F, Ma F, Shi R, Zhang J, Han S, Zhang Y (2015) Production of rhamnolipids by Pseudomonas aeruginosa is inhibited by H2S but resumes in a co-culture with P. stutzeri: applications for microbial enhanced oil recovery. Biotechnol Lett 37:1–6
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grants No. 41673084, 41530318, 41273084), NSFC/RGC Joint Research Fund (Grant No. 41161160560) and the Research Foundation of Shanghai (No. 15JC1401400).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, H., Liu, JF., Li, CY. et al. Anaerobic biodegradation of partially hydrolyzed polyacrylamide in long-term methanogenic enrichment cultures from production water of oil reservoirs. Biodegradation 29, 233–243 (2018). https://doi.org/10.1007/s10532-018-9825-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-018-9825-1