Abstract
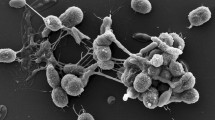
A novel Gram-positive actinobacterium, designated WT-2-1T, was isolated from a sample of petroleum-contaminated soil collected in Daqing, Heilongjiang province, China and characterised using a polyphasic taxonomic approach. The optimal growth for strain WT-2-1T was found to be at 25–35 °C and at pH 6.0–9.0 and with 0–4% (w/v) NaCl, forming blackish green-coloured colonies. Chemotaxonomic and molecular characteristics of the isolate match those described for members of the genus Geodermatophilus. The peptidoglycan was found to contain meso-diaminopimelic acid; galactose, glucose and xylose were detected as diagnostic sugars. The main phospholipids were identified as diphosphatidylglycerol, phosphatidylcholine, phosphatidylinositol, phosphatidylethanolamine and phosphatidylglycerol; MK-9(H4) was the dominant menaquinone present. The major cellular fatty acids were identified as iso-C16:0 and iso-C15:0. 16S rRNA gene sequence analysis showed that strain WT-2-1T is a member of the genus Geodermatophilus, with high sequence similarities to Geodermatophilus aquaeductus BMG801T (98.4%), Geodermatophilus saharensis CF5/5T (98.4%), Geodermatophilus bullaregiensis BMG841T (98.3%) and Geodermatophilus normandii CF5/3T (98.3%). Based on the phenotypic characteristics, phylogenetic data and DNA–DNA hybridization results, the isolate is concluded to represent a novel species of the genus Geodermatophilus, for which the name Geodermatophilus daqingensis sp. nov. is proposed. The type strain is WT-2-1T (=CGMCC 4.7381T = DSM 104001T).

Similar content being viewed by others
References
Bertazzo M, Montero-Calasanz MC, Martinez-Garcia M et al (2014) Geodermatophilus brasiliensis sp. nov., isolated from Brazilian soil. Int J Syst Evol Microbiol 64:2841–2848
Collins MD (1985) Isoprenoid quinone analyses in bacterial classification and identification. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic Press, London, pp 267–284
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–789
Fitch WM (1971) toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Gordon RE, Barnett DA, Handerhan JE, Pang CHN (1974) Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Bacteriol 24:54–63
Gregersen T (1978) Rapid method for distinction of gramnegative from gram-positive bacteria. Eur J Appl Microbiol Biotechnol 5:123–127
Hezbri K, Ghodhbane-Gtari F, Montero-Calasanz MC et al (2015a) Geodermatophilus aquaeductus sp. nov., isolated from the ruins of Hadrian’s aqueduct. Antonie Van Leeuwenhoek 108:41–50
Hezbri K, Ghodhbane-Gtari F, Montero-Calasanz MC et al (2015b) Description of Geodermatophilus bullaregiensis sp. nov. Antonie Van Leeuwenhoek 108:415–425
Huss VAR, Festl H, Schleifer KH (1983) Studies on the spectrometric determination of DNA hybridisation from renaturation rates. Syst Appl Microbiol 4:184–192
Ishiguro EE, Wolfe RS (1970) Control of Morphogenesis in Geodermatophilus: ultrastructural studies. J Bacteriol 104:566–580
Jin L, Lee H-G, Kim H-S et al (2013) Geodermatophilus soli sp. nov. and Geodermatophilus terrae sp. nov., two actinobacteria isolated from grass soil. Int J Syst Evol Microbiol 63:2625–2629
Kim SB, Brown R, Oldfield C, Gilbert SC, Iliarionov S, Goodfellow M (2000) Gordonia amicalis sp. nov., a novel dibenzothiophene-desulphurizing actinomycete. Int J Syst Evol Microbiol 50:2031–2036
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lechevalier MP, Lechevalier HA (1970) Chemical composition as a criterion in the classification of aerobic actinomycetes. Int J Syst Bacteriol 20:435–443
Lechevalier MP, Lechevalier HA (1980) The chemotaxonomy of actinomycetes. In Actinomycete Taxonomy (Special Publication No. 6). Society for Industrial Microbiology, Arlington, pp 277–284
Luedemann GM (1968) Geodermatophilus, a new genus of the Dermatophilaceae (Actinomycetales). J Bacteriol 96:1848–1858
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal K, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Montero-Calasanz MC, Göker M, Poẗter G et al (2012) Geodermatophilus arenarius sp. nov., a xerophilic actinomycete isolated from Saharan desert sand in Chad. Extremophiles 16:903–909
Montero-Calasanz MC, Göker M, Poẗter G et al (2013a) Geodermatophilus saharensis sp. nov., isolated from sand of the Saharan desert in Chad. Arch Microbiol 195:153–159
Montero-Calasanz MC, Göker M, Poẗter G et al (2013b) Geodermatophilus normandii sp. nov., isolated from Saharan desert sand. Int J Syst Evol Microbiol 63:3437–3443
Montero-Calasanz MC, Göker M, Poẗter G et al (2014a) Description of Geodermatophilus amargosae sp. nov., to accommodate the not validly named Geodermatophilus obscurus subsp. amargosae (Luedemann, 1968). Curr Microbiol 68:365–371
Montero-Calasanz MC, Hofner B, Göker M (2014b) Geodermatophilus poikilotrophi sp. nov.: a multitolerant actinomycete isolated from dolomitic marble. Biomed Res Int 11:914767
Nie G-X, Ming H, Li S et al (2012) Geodermatophilus nigrescens sp. nov., isolated from a dry-hot valley. Antonie Van Leeuwenhoek 101:811–817
Normand P (2006) Geodermatophilaceae fam. nov., a formal description. Int J Syst Evol Microbiol 56:2277–2278
Normand P, Orso S, Cournoyer B, Jeannin P, Chapelon C, Dawson J, Evtushenko L, Misra AK (1996) Molecular phylogeny of the genus Frankia and related genera and emendation of the family Frankiaceae. Int J Syst Bacteriol 46:1–9
Qu J-H, Hui M, Qu J-Y et al (2013) Geodermatophilus taihuensis sp. nov., isolated from the interfacial sediment of a eutrophic lake. Int J Syst Evol Microbiol 63:4108–4112
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC Newsl 20:16
Sen A, Daubin V, Abrouk D et al (2014) Phylogeny of the class Actinobacteria revisited in the light of complete genomes. The orders ‘‘Frankiales’’ and Micrococcales should be split into coherent entities: proposal of Frankiales ord. nov., Geodermatophilales ord. nov., Acidothermales ord. nov. an. Int J Syst Evol Microbiol 64:3821–3832
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int Syst Bacteriol 16:313–340
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, pp 607–654
Staneck JL, Roberts GD (1974) Simplified approach to identification of aerobic actinomycetes by 279 thin-layer chromatography. Appl Microbiol 28:226–231
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Wu C, Lu X, Qin M, Wang Y, Ruan J (1989) Analysis of menaquinone compound in microbial cells by HPLC. Microbiology 16:176–178 [English translation of Microbiology (Beijing)]
Yokota A, Tamura T, Hasegawa T, Huang LH (1993) Catenuloplanes japonicus gen. nov., sp. nov., nom. rev., a new genus of the order Actinomycetales. Int J Syst Bacteriol 43:805–812
Zhang Y-Q, Chen J, Liu H-Y et al (2011) Geodermatophilus ruber sp. nov., isolated from rhizosphere soil of a medicinal plant. Int J Syst Evol Microbiol 61:190–193
Zhi X-Y, Li W-J, Stackebrandt E (2009) An update of the structure and 16S rRNA gene sequence-based definition of higher ranks of the class Actinobacteria, with the proposal of two new suborders and four new families and emended descriptions of the existing higher taxa. Int J Syst Evol Microbiol 59:589–608
Acknowledgements
This work was supported by the National Natural Science Fund of China (31372351 and 31672469), the Youth Foundation of Daqing Normal University (14ZR17)and the National Natural Science Fund of Heilongjiang Province (QC2011C094).
Author information
Authors and Affiliations
Corresponding author
Additional information
Yang Wang and Liguo Zhang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, Y., Zhang, L., Zhang, X. et al. Geodermatophilus daqingensis sp. nov., isolated from petroleum-contaminated soil. Antonie van Leeuwenhoek 110, 803–809 (2017). https://doi.org/10.1007/s10482-017-0853-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-017-0853-3