Abstract
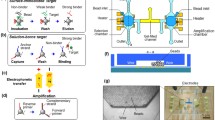
Aptamers are synthetic single-stranded nucleic acid molecules that bind to biochemical targets with high affinity and specificity. The method of systematic evolution of ligands by exponential enrichment (SELEX) is widely used to isolate aptamers from randomized oligonucleotides. Recently, microfluidic technology has been applied to improve the efficiency and reduce the cost in SELEX processes. In this work, we present an approach that exploits surface acoustic waves to improve the affinity selection process in microfluidic SELEX. Acoustic streaming is used to enhance the interactions of the solution-based oligonucleotide molecules with microbead-immobilized target molecules, allowing the identification of high-affinity aptamer candidates in a more efficient manner. For demonstration, a DNA aptamer is isolated within three rounds of selection in 5 h to specifically bind to immunoglobulin E, a representative target protein, with an equilibrium dissociation constant of approximately 22.6 nM.








Similar content being viewed by others
References
Afzal M, Park J, Jeon JS, Akmal M, Yoon T-S, Sung HJ (2021) Acoustofluidic separation of proteins using aptamer-functionalized microparticles. Anal Chem 93(23):8309–8317
Ahmed H, Park J, Destgeer G, Afzal M, Sung HJ (2019) Surface acoustic wave-based micromixing enhancement using a single interdigital transducer. Appl Phys Lett 114(4):043702
Ahn JY, Lee S, Jo M, Kang J, Kim E, Jeong OC, Laurell T, Kim S (2012) Sol-gel derived nanoporous compositions for entrapping small molecules and their outlook toward aptamer screening. Anal Chem 84(6):2647–2653
Bachman H, Chen CY, Rufo J, Zhao SG, Yang SJ, Tian ZH, Nama N, Huang PH, Huang TJ (2020) An acoustofluidic device for efficient mixing over a wide range of flow rates. Lab Chip 20(7):1238–1248
Ch MH, Amani J, Sedighian H, Amin M, Salimian J, Halabian R, Fooladi AAI (2016) Isolation of a new ssDNA aptamer against staphylococcal enterotoxin B based on CNBr-activated sepharose-4B affinity chromatography. J Mol Recognit 29(9):436–445
Chakravarthy M, AlShamaileh H, Huang H, Tannenberg RK, Chen S, Worrall S, Dodd PR, Veedu RN (2018) Development of DNA aptamers targeting low-molecular-weight amyloid-β peptide aggregates in vitro. Chem Commun (camb) 54(36):4593–4596
Collins DJ, Ma Z, Han J, Ai Y (2017) Continuous micro-vortex-based nanoparticle manipulation via focused surface acoustic waves. Lab Chip 17(1):91–103
Dembowski SK, Bowser MT (2018) Microfluidic methods for aptamer selection and characterization. Analyst 143(1):21–32
Destgeer G, Sung HJ (2015) Recent advances in microfluidic actuation and micro-object manipulation via surface acoustic waves. Lab Chip 15(13):2722–2738
Destgeer G, Ha BH, Jung JH, Sung HJ (2014) Submicron separation of microspheres via travelling surface acoustic waves. Lab Chip 14(24):4665–4672
Dong LL, Tan QW, Ye W, Liu DL, Chen HF, Hu HW, Wen D, Liu Y, Cao Y, Kang JW, Fan J, Guo W, Wu WZ (2015) Screening and identifying a novel ssDNA aptamer against alpha-fetoprotein using CE-SELEX. Sci Rep. https://doi.org/10.1038/srep15552
Hoinka J, Backofen R, Przytycka TM (2018) AptaSUITE: a full-featured bioinformatics framework for the comprehensive analysis of aptamers from HT-SELEX experiments. Mol Ther Nucleic Acids 11:515–517
Hong SL, Wan YT, Tang M, Pang DW, Zhang ZL (2017) Multifunctional screening platform for the highly efficient discovery of aptamers with high affinity and specificity. Anal Chem 89(12):6535–6542
Kim HR, Song MY, Chan Kim B (2020) Rapid isolation of bacteria-specific aptamers with a non-SELEX-based method. Anal Biochem 591:113542
Li PX, Ma ZC, Zhou YN, Collins DJ, Wang ZF, Ai Y (2019) Detachable acoustophoretic system for fluorescence-activated sorting at the single-droplet level. Anal Chem 91(15):9970–9977
Lisi S, Fiore E, Scarano S, Pascale E, Boehman Y, Duconge F, Chierici S, Minunni M, Peyrin E, Ravelet C (2018) Non-SELEX isolation of DNA aptamers for the homogeneous-phase fluorescence anisotropy sensing of tau proteins. Anal Chim Acta 1038:173–181
Ma YY, Li XL, Li W, Liu Z (2018) Glycan-imprinted magnetic nanoparticle-based SELEX for efficient screening of glycoprotein-binding aptamers. ACS Appl Mater Interfaces 10(47):40918–40926
Macdonald J, Houghton P, Xiang DX, Duan W, Shigdar S (2016) Truncation and mutation of a transferrin receptor aptamer enhances binding affinity. Nucleic Acid Ther 26(6):348–354
Minagawa H, Kataoka Y, Kuwahara M, Horii K, Shiratori I, Waga I (2020) A high affinity modified DNA aptamer containing base-appended bases for human β-defensin. Anal Biochem 594:113627
Muller PB, Barnkob R, Jensen MJH, Bruus H (2012) A numerical study of microparticle acoustophoresis driven by acoustic radiation forces and streaming-induced drag forces. Lab Chip 12(22):4617–4627
Nakatsuka N, Yang K-A, Abendroth JM, Cheung KM, Xu X, Yang H, Zhao C, Zhu B, Rim YS, Yang Y, Weiss PS, Stojanović MN, Andrews AM (2018) Aptamer–field-effect transistors overcome Debye length limitations for small-molecule sensing. Science 362(6412):319
Narayan C, Kwon J, Kim C, Kim S-J, Jang SK (2020) Virus-based SELEX (viro-SELEX) allows development of aptamers targeting knotty proteins. Analyst 145(4):1473–1482
Ni S, Zhuo Z, Pan Y, Yu Y, Li F, Liu J, Wang L, Wu X, Li D, Wan Y, Zhang L, Yang Z, Zhang B-T, Lu A, Zhang G (2021) Recent progress in aptamer discoveries and modifications for therapeutic applications. ACS Appl Mater Interfaces 13(8):9500–9519
Olsen T, Zhu J, Kim J, Pei R, Stojanovic MN, Lin Q (2017) An integrated microfluidic SELEX approach using combined electrokinetic and hydrodynamic manipulation. SLAS Technol 22(1):63–72
Park JW, Lee SJ, Ren S, Lee S, Kim S, Laurell T (2016) Acousto-microfluidics for screening of ssDNA aptamer. Sci Rep. https://doi.org/10.1038/srep27121
Qu H, Csordas AT, Wang J, Oh SS, Eisenstein MS, Soh HT (2016) Rapid and label-free strategy to isolate aptamers for metal ions. ACS Nano 10(8):7558–7565
Sinha A, Gopinathan P, Chung YD, Lin HY, Li KH, Ma HP, Huang PC, Shiesh SC, Lee GB (2018) An integrated microfluidic platform to perform uninterrupted SELEX cycles to screen affinity reagents specific to cardiovascular biomarkers. Biosens Bioelectron 122:104–112
Song KM, Lee S, Ban C (2012) Aptamers and their biological applications. Sensors 12(1):612–631
Thevendran R, Navien TN, Meng X, Wen K, Lin Q, Sarah S, Tang T-H, Citartan M (2020) Mathematical approaches in estimating aptamer-target binding affinity. Anal Biochem 600:113742
Tominaga J, Saini R, Hoe C, Tang T, Tan S, Citartan M, Gopinath SCB (2012) Asymmetric PCR for good quality ssDNA generation towards DNA aptamer production. Songklanakarin J Sci Technol 34:125–131
Wang G, Liu J, Chen K, Xu Y, Liu B, Liao J, Zhu L, Hu X, Li J, Pu Y, Zhong W, Fu T, Liu H, Tan W (2017) Selection and characterization of DNA aptamer against glucagon receptor by cell-SELEX. Sci Rep 7(1):7179
Wiegand TW, Williams PB, Dreskin SC, Jouvin MH, Kinet JP, Tasset D (1996) High-affinity oligonucleotide ligands to human IgE inhibit binding to Fc epsilon receptor I. J Immunol 157(1):221–230
Yan JH, Xiong HJ, Cai SD, Wen NC, He QY, Liu YF, Peng DM, Liu ZB (2019) Advances in aptamer screening technologies. Talanta 200:124–144
Yüce M, Ullah N, Budak H (2015) Trends in aptamer selection methods and applications. Analyst 140(16):5379–5399
Zhang Y, Lai BS, Juhas M (2019) Recent advances in aptamer discovery and applications. Molecules 24(5):941
Zheng J, Yang RH, Shi ML, Wu CC, Fang XH, Li YH, Li JH, Tan WH (2015) Rationally designed molecular beacons for bioanalytical and biomedical applications. Chem Soc Rev 44(10):3036–3055
Zhou J, Rossi J (2017) Aptamers as targeted therapeutics: current potential and challenges. Nat Rev Drug Discov 16(3):181–202
Zhuo ZJ, Yu YY, Wang ML, Li J, Zhang ZK, Liu J, Wu XH, Lu AP, Zhang G, Zhang BT (2017) Recent advances in SELEX technology and aptamer applications in biomedicine. Int J Mol Sci 18(10):2142
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31(13):3406–3415
Acknowledgements
We gratefully acknowledge financial support from the National Institutes of Health (grant number 1R33CA196470-01A1) and a pilot grant from the Opportunity Funds Management Core of the Centers for Medical Countermeasures against Radiation, National Institute of Allergy and Infectious Diseases (grant number U19AI067773). C. Bai also gratefully acknowledges a National Scholarship (award number 201806280166) from the China Scholarship Council and financial support from the National Natural Science Foundation of China (grant number 52175545).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bai, C., Meng, X., Wen, K. et al. Surface acoustic wave-assisted microfluidic isolation of aptamers. Microfluid Nanofluid 26, 43 (2022). https://doi.org/10.1007/s10404-022-02548-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10404-022-02548-w