Abstract
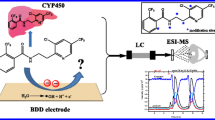
Novel, rapid and precise RP-HPLC–DAD method was developed, validated and successfully applied for determination of metabolic changes of ethyl 5-(4-bromophenyl)-1-(3-(2-(2-hydroxybenzylidene)hydrazinyl)-3-oxopropyl)-2-methyl-1H-pyrrole-3-carboxylate (12b) in isolated rat hepatocytes. The analytes were detected by a simple DAD detector at 279 nm wavelength. A single-step extraction method was implemented to enable fast purification and extraction from cellular culture, resulting in a complete recovery. Thereafter, the method was adequately transferred to a LC–MS system for identification of unknown products. Additionally, network metabolism evaluation was performed to predict the structures of major metabolites with their isotope mass through BioTransformer 3.0. The data from the LC–MS analysis and the online server were compared for comprehensive identification. The results indicated formation of four metabolic products, obtained through processes of hydrolysis (12 and b), hydroxylation in the structure 12b (M1) and O-dealkylation (M2).
Similar content being viewed by others
Introduction
A leading cause of death from a single infecting agent is commonly linked with tuberculosis (TB), outranking even HIV/AIDS. The current COVID-19 pandemic caused a decrease in the successful reduction of TB cases and some constraints in providing essential TB health services. Despite some divergence in global TB targets, some areas of the world have kept some success in this direction [1].
The past decades pointed a dramatic increase of the anti-tubercular therapy, regardless of its introduction in the 1950s [2] with the observed multiplication of Mycobacterium Avium Complex (MAC) infections, affecting mostly patients with AIDS worldwide [3, 4]. Another serious burden is the drug-resistant TB with increasing the day-by-day cases caused by Mycobacterium tuberculosis-resistant strains to one or more anti-tubercular drugs [5,6,7,8].
Varieties chemical moieties have been introduced in the urgent discovery of novel anti-tubercular agents, including hydrazones [9] and derivatives of pyrrole [10, 11]. In addition, pyrrole-based compounds are identified as potential chemotherapeutics applicable for the inhibition of M. tuberculosis and other atypical mycobacteria, including M. avium complex, which greatly contributes to the deaths of AIDS patients [10, 12].
Drug discovery is a time-consuming, expensive and unsuccessful for most of the candidate molecules process [13, 14]. Therefore, the preliminary evaluation of the pharmacokinetic including the molecules absorption, distribution, metabolism and excretion (ADME) is essential. Metabolism studies can identify active and/or toxic metabolites that can be utilized to further explore the metabolic liability [15, 16].
Considering the perspective anti-tubercular effects of previously synthesized pyrrole hydrazones and the necessity of toxicological profiling [17], the aim of the current study is focused on development and validation of an appropriate RP-HPLC–DAD method for investigation of biotransformation of such derivatives.
Materials and Methods
Chemicals and Reagents
All of the reagents used for preparation of mobile phase and solutions in the current work are purchased from Sigma Aldrich. The acetonitrile and methanol are obtained from Fisher Scientific (Seoul, Republic of Korea) and are of HPLC grade. Water used throughout the analyses is from a Millipore system.
Animals and Experimental Procedure
For this investigation, 200 g male Wistar rats, housed in plexiglass cages at room temperature, were used. The former were acquired from the National Breeding Center, Sofia, Bulgaria, and all conducted investigations were approved by the Institutional Animal Care Committee in accordance with European Union Guidelines for animal experimentation. The rat was anesthetized with sodium pentobarbital (0.2 mL/ 100 g). A modified method described by Fau et al. [18] was applied for the in situ isolation of the cells and the liver perfusion. 100 mL HEPES buffer (pH = 7.85) was utilized for the perfusion of the liver, containing 10 mM HEPES, 142 mM NaCl, 7 mM KCl, 5 mM glucose and 0.6 mM EDTA (pH = 7.85). Subsequently, 200 ml HEPES buffer (pH = 7.85), and a final addition of 200 ml HEPES buffer containing collagenase type IV (50 mg/200 mL) and 7 mM CaCl2 (pH = 7.85) was added.
The procedure for cell suspension extraction includes the excise and mince into small pieces of the liver and further dispersion of the hepatocytes in 60 mL Krebs–Ringer-bicarbonate (KRB) buffer (pH = 7.35), containing 1.2 mM KH2PO4, 1 mM CaCl2, 1.2 mM MgSO4, 5 mM KCl, 5 mM NaHCO3, 4.5 mM glucose and 1% bovine serum albumin. The hepatocytes were centrifuged at 500 g for 1 min after the initial filtration, and there were washed three times with a KRB buffer. Cells were counted under the microscope and the viability was assessed by Trypan blue exclusion (0.05%), which was calculated to be 89% in average [18]. Cells were diluted with a KRB, to make a suspension of about 3 × 106 hepatocytes/mL. Incubations were carried out in 25 mL Erlenmeyer flasks with each containing 3 mL of the cell suspension (i.e., 9 × 106 hepatocytes). The latter procedures were performed in a 5% CO2 + 95% O2 atmosphere.
Stock and Working Solutions Preparation
For preparation of stock solution, a 2.4 mg sample of the tested compound was accurately weighed and dissolved in a volumetric flask of 5 mL with dimethyl sulfoxide (DMSO) (1 mM). Portions (37.5–300 µL) of this solution were diluted to 10 mL with mobile phase and were used for calibration standards. For the investigation, a 210 µL of the stock solution was pipetted out and was incubated with isolated rat hepatocytes in final concentration 100 μM for 2 h.
Sample Preparation
The sample preparation included a rapid and simple protein precipitation. In an Eppendorf tube, 400 µL of incubating hepatocytes with hydrazone was deproteinized with 1000 µL methanol. After having vortex-mixed thoroughly for 2 min, the samples were centrifuged for 15 min at strictly 14 000 rpm. Prior to analysis, the supernatant was double filtered through PVDF sterile syringe filters (through 0.47 μm, and through 0.22 μm) and a volume of 20 µL was injected into the chromatographic system.
Chromatographic Conditions
The samples were initially analyzed by HPLC system that consisted of an UltiMateDionex 3000 equipped with an UltiMateDionex 3000 diode array detector, all from TermoFischer Scientific®, San Jose, CA, USA. The data were processed by Chromeleon® 7.2 software. The analytes were separated on a Purospher STAR RP-18 (4.6 × 250 mm, particles size 5 µm) and Purospher STAR RP-18 (4.6 × 125 mm, particles size 5 µm) columns, which were obtained from Merck Millipore, Germany. The formers were conditioned at 25 °C in a column oven. The mobile phase consisted of acetonitrile: phosphate buffer pH 3.5: methanol in ratio 35/35/30 (v/v/v). The phosphate buffer was prepared dissolving 5.04 g of disodium hydrogen phosphate and 3.01 g of potassium dihydrogen phosphate in dist. water and the volume was made up to 1000 mL with distilled water. For adjusting the pH to 3.5, orthophosphoric acid was used, as described in European Pharmacopoea. All of the solvents were filtered through a membrane filter (0.20 μm) using a Millipore glass filter holder. The working flow and injection volume were set at 1.0 mL/min and 20 µL, respectively. The signal from the DAD was set to 279 nm.
The appropriate wavelength for detection of the standard and working solutions was evaluated, where the samples were scanned at a wavelength range of 200—400 nm using UV–Vis Hewlett Packard 8452 Spectrophotometer. The maximum wavelength (λmax) was found to be 279 nm, determining the DAD detection to be set to this value.
For detecting the metabolites, a Q Exactive Plus mass spectrometer (ThermoFisher Scientific®) equipped with a heated electrospray ionization (HESI-II) probe (ThermoScientific) was used. The applied parameters were spray voltage 3.5 kV, sheath gas flow rate 38, auxiliary gas flow rate 12, spare gas flow rate 0, capillary temperature 320 °C, probe heater temperature 320 °C, and S-lens RF level 50. The final data acquisition and analysis were conducted on Xcalibur 4.1 software (Thermo Fisher Scientific®, San Jose, CA, USA).
The developed LC–DAD method was adequately transferred to the LC–MS system by acidifying the mobile phase with formic acid instead of an inorganic salt and finally the mobile phase consisted 0.1% formic acid in dist. water (solvent A) and 0.1% formic acid in acetonitrile (solvent B). The gradient program was as follows: 0–1.5 min, 5–10% B; 1.5–13 min solvent B linearly increased to 60% followed by 6 min isocratic elution; 19–23 min 60–95% B and finished with 95% solvent B an isocratic elution for 2 min with a flow rate of 3.0 mL/min. The equilibration time between each injection was 4 min and the total run time was 25 min.
Method Validation
The developed methods specificity, linearity, precision, accuracy and selectivity were determined, as required by the ICH guidelines Q2 (R1).
Linearity
The linearity was evaluated by analysis of 8 different linearly increasing concentrations 25, 50, 75, 100, 125, 150, 175 and 200% of selected hydrazone with calibration range 3.75–30 µM. Each solution was injected in triplicate and the obtained peak areas and concentrations were used to build a calibration curve (Fig. 1), with subsequent calculation of the linear equation parameters presented in Table 3.
Precision and Accuracy
The precision of the method was determined in terms of repeatability and intermediate precision and was assessed as a relative standard deviation (RSD) which was expected to be RSD ≤ 2.0%. Accuracy was evaluated by analysis of three different levels of the target concentration (50%, 100% and 150%). In order to prove the accuracy of the proposed method, percentage recovery and %RSD values should be ranging from 98.0% to 102.0%, and not more than 2.0%, respectively.
Specificity and Sensitivity
The specificity was evaluated through analysis of the blank solution, containing DMSO and isolated rat hepatocyte suspension.
The sensitivity was assessed by Limit of Detection (LOD) and Limit of Quantification (LOQ). The calculation was based on the standard deviation of the intercept (α) and a slope (S) of the curve, using the equations: LOD = (3 × α)/S and LOQ = (10 × α)/S.
Stability
The assessment tracked stability evaluation of the selected compound under conditions of bioanalytical tests. The experiment was conducted under 37 °C temperature for 24 h in phosphate buffer with pH 7.4.
Results and Discussion
Selection of a Representative Test Substance
Based on the perspective anti-tubercular effect of the previously synthesized by us ethyl 5-(4-bromophenyl)-1-(3-(2-(2-hydroxybenzylidene) hydrazinyl)-3-oxopropyl)-2-methyl-1H-pyrrole-3-carboxylate (12b) [17] this representative was selected for investigation of the biotransformation processes undergoing in isolated rat hepatocyte cultures.
Method Development
The studied N-pyrrolyl hydrazide-hydrozone is a hydrophobic molecule with log P – 5.38. Therefore, we selected a reversed-phase column for current analysis. Several C18 columns of different lengths 4.6 × 25 cm, 4.6 × 12.5 cm, with particle diameter of 5 µm were compared. The results indicated the shorter column as a more appropriate, delivering admissible run time of 17.203 min, symmetric peak shape and better resolution. The column is acids/base resistant, thus usable also for stability evaluation.
The selection of the mobile phase composition was based on our previous investigations with hydrazones, containing pyrrole ring [20]. The development of the appropriate mobile phase started with bi-component mixtures of methanol and acetonitrile in different ratios. The performed analysis indicated that increased amounts of methanol and acetonitrile resulted in deteriorated peak shapes or retention times of less than 5 min, respectively. This required introduction of additional acidifying solvent, attempting to improve the chromatographical behavior of the analyzed mixture. Thus, thereafter, phosphate buffer with pH 3.5 was added as a component of the mobile phase (Table 1).
To achieve symmetrical peak shapes and short retention times, the mobile phase consisting of acetonitrile: phosphate buffer pH 3.5: methanol in ratio 35/35/30 (v/v/v) was selected. The preliminary UV/VIS method developed by us identified 279 nm as the most adequate wavelength.
In order to determine the applicability of the method standard solutions of the evaluated hydrazone 12b and the expected hydrolysis products [21] of the corresponding hydrazide 12 and aldehyde b were chromatographed (Fig. 1).
The indicated conditions proved to be suitable to achieve a good separation between the expected degradation products, which determined this method as applicable for the further biotransformation evaluations of the initial ethyl 5-(4-bromophenyl)-1-(3-(2-(2-hydroxybenzylidene)hydrazinyl)-3-oxopropyl)-2-methyl-1H-pyrrole-3-carboxylate (12b).
Method Validation
Linearity
The linearity demonstrated the linear dependence of the analytical signal from the concentration of the analyte within the analytical area of the method [19]. For the calibration curve, the correlation coefficient (R2) must be above 0.99 to be acceptable.
Each solution as prepared according to the methodology explained in 2.6.1, was injected in triplicate and the obtained peak areas and concentrations were used to build a calibration curve (Fig. 2), with subsequent calculation of the linear equation y = 0.5908x-0.0189 and the correlation coefficient 0.9999 (Table 4).
The R2 coefficient corresponds to the requirement of a value not less than 0.99, which determines the method as linear in the evaluated range.
Precision and Accuracy
Repeatability and intermediate precision were determined from measurements performed on the same day (repeatability) and on different days (intermediate precision) of the 100% of the nominal concentration six-fold. Results are summarized in Table 2.
Accuracy was determined by injecting three different levels of the nominal concentration in triplicate. The data for their peak areas were used to calculate the average recovery and RSD, as presented in the following Table 3.
The calculated standard deviation of less than 2 and % recovery range of 99.09–101.29, indicate the developed method as accurate, precise and applicable for further analysis.
Specificity and Sensitivity
The chromatograms of the blanks and the spiked hydrazone sample at 30 µM presented the absence of interfering peaks in work-window retention time, which affirmed the specificity of the method.
In addition, no carryover was observed by injecting a blank solution of mobile phase and pre-treated hepatocyte suspension immediately following the highest calibration standard. The sensitivity of the method was also determined by calculating the Limit of Detection (LOD) and Limit of Quantification (LOQ) parameters applying the aforementioned equations.
As per ICH guidelines, depending on the instrument availability the limit of detection (LOD) and limit of quantitation (LOQ) may be determined by several approaches [19]. In the current research, the estimation was based on standard deviation of the response and the slope of the calibration curve of the analyte. The obtained values of the latter parameters are 740 nM and 2.47 µM, respectively, as given in Table 4.
Stability
In the current study, stability evaluation of the working standards in buffer with pH 7.4 was done. This assessment indicates the conditions under which the biological samples were stored, collected and processed during the analysis. Therefore, the solution of hydrazone (15 µM) was incubated for 24 h in buffer with physiological pH and temperature (37 °C). The applicability of the method and the stability of the molecules under the developed chromatographic conditions was confirmed by the lack of interfering peaks with retention times equal to or close to the ones of the analyzed substance.
Method Application
After development, validation and stability evaluation, the current method was successfully applied for metabolite determination. The selected compound was incubated for 2 h in isolated rat hepatocyte suspension. Thereafter, 20 µL of the pre-treated samples were injected into the HPLC system. The samples were collected in 60th (A) and 120th (B) minute.
The chromatogram on Fig. 3A represents the result at 60th minute of incubation indicating appearance of four new peaks two products of hydrolysis showing the release of salicylic aldehyde at retention time of 2.10 min and the corresponding initial N-pyrrolyl hydrazide at 4.21 min. The other two peaks lettered M1 and M2 with retention times 3.02 and 3.61 do not resemble with any of the known reference molecules.
The results (Fig. 3B) after 120 min of incubation demonstrate decrease in concentration of the tested 12b accompanied by an increase in the amount of the obtained metabolic products.
As an attempt to identify the two unknown molecules a multiple steps strategy was created starting with in silico prediction of the changes in the chemical structure of the evaluated hydrazone, followed by application of MS detection, as an approach for structural elucidation.
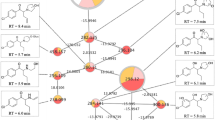
First, we applied a virtual preliminary metabolic online predictor BioTransformers 3.0. The obtained data established as the most probable to be the hydroxylation and O-dealkylation resulting in the corresponding compounds characterized with the chemical formulas and major isotopes of m/z—C24H24BrN3O5, 513.0899 and C22H20BrN3O4, 469.06372, respectively (Fig. 4).
A suitable LC–MS analysis was applied, with additional replacement of the used in the mobile phase inorganic phosphate buffer with the more MS acceptable formic acid acidifier [22]. This correction was pointed to avoid possible crystallization of the inorganic salt during the MS analysis. The subsequent evaluation of selectivity and analyte’s behavior determined the method as an appropriate and proved that altering this component of the mobile phase did not affect the analytical result (Fig. 5A). Therefore, we started with high-resolution full mass scans through LC–MS, which provides accurate mass measurements and precise assignment of molecular formulas. The results indicated the molecular masses of the obtained new products, identified in the chromatogram as M1 and M2 to be m/z of 513.0889 and 469.06372, respectively (Fig. 5B, C).
Finally, both methods for evaluation of the m/z values for the indicated possible metabolites were compared. The results allowed us to determine the most probable biotransformation pathway related to the proposed structures.
Conclusion
A rapid, sensitive and selective method for biotransformational evaluation of newly synthesized promising anti-tubercular pyrrole-based hydrazone is developed and validated. The analytical procedure is based on a C18 RP column, tempered at 25 °C and an isocratically eluted mobile phase of acetonitrile: phosphate buffer pH 3.5: methanol in ratio 35/35/30 (v/v/v) and DAD detection at 279 nm. In addition, in silico approach and LC–MS analysis for prediction and identification of the possible metabolites are conducted. The performed evaluations determined the formation of four metabolites two of which are the hydrolysis products 12 (N-pyrrolyl hydrazide) and b (salicylaldehyde), one is the hydroxylated compound M1 with retention time 3.01 and the last is the O-dealkylated 5-(4-bromophenyl)-1-(3-(2-(2-hydroxybenzylidene)hydrazineyl)-3-oxopropyl)-2-methyl-1H-pyrrole-3-carboxylic acid (M2) with retention time 3.61. The obtained results determined the suggested procedures applicable for future investigations of biotransforamtional changes of pyrrole-based hydrazide-hydrazones.
Data availability
All data used to support the findings of this study are available from the corresponding author upon request.
References
Global tuberculosis report 2021.(2021) Geneva: World Health Organization. Licence: CC BY-NC-SA 3.0 IGO.
Global Tuberculosis Control: Surveillance, Planning, Financing (2009) WHO Report, World Health Organization.
Masur H (1993) Recommendations on prophylaxis and therapy for disseminated mycobacterium avium complex disease in patients infected with the human immunodeficiency virus. N Engl J Med 329:898–904
Falkinham J (1996) Epidemiology of infection by nontuberculous mycobacteria. Clin Microbiol Rev 9:177
Hamilton CD (1999) Recent developments in epidemiology, treatment, and diagnosis of tuberculosis. Curr Infect Dis Reports 1:80–88
W. Fitzgerald D., Morse M., Pape J., Johnson J.W. (2000) Active tuberculosis in individuals infected with human immunodeficiency virus after isoniazid prophylaxis. Clin Infect Dis 31:1495–1497
Ahmad S, Alam O, Naim MJ, Shaquiquzzaman M, Alam MM, Iqbal M (2018) Pyrrole: An insight into recent pharmacological advances with structure activity relationship. Eur J Med Chem 157:527–561
Mateev E, Georgieva M, Zlatkov A (2022) Pyrrole as an important scaffold of anticancer drugs: recent advances. J Pharm Pharm Sci 25:24–40
Bijev A, Georgieva M (2010) Pyrrole-based hydrazones synthesized and evaluated in vitro as potential tuberculostatics. Lett Drug Des Discovery 7:430–437
Ragno R, Marshall GR, Santo R, Di CR, Massa S, Rompei R, Artico M (2000) Antimycobacterial pyrroles: synthesis, anti- Mycobacterium tuberculosis activity and QSAR studies. Bioorg Med Chem 8:1423–1432
Biava M, Porretta GC, Deidda D, Pompei R, Tafi A, Manetti F (2004) Antimycobacterial compounds. New pyrrole derivatives of BM212. Bioorg Med Chem 12:1453–1458
Semenya D, Touitou M, Ribeiro CM, Pavan FR, Pisano L, Singh V, Chibale K, Bano G, Toscani A, Manetti F, Gianibbi B, Castagnolo D (2021) Structural rigidification of N-Aryl-pyrroles into indoles active against intracellular and drug-resistant mycobacteria. ACS Med Chem Lett 13:63–69
Kiriiri GK, Njogu PM, Mwangi AN (2020) Exploring different approaches to improve the success of drug discovery and development projects: a review. Futur J Pharm Sci. https://doi.org/10.1186/s43094-020-00047-9
Gaudelet T, Day B, Jamasb AR, Soman J, Regep C, Liu G, Hayter JBR, Vickers R, Roberts C, Tang J, Roblin D, Blundell TL, Bronstein MM, Taylor-King JP (2021) Utilizing graph machine learning within drug discovery and development. Brief Bioinform 22:159
Abdel-Mohsen HT, Abood A, Flanagan KJ, Meindl A, Senge MO, El Diwani HI (2020) Synthesis, crystal structure, and ADME prediction studies of novel imidazopyrimidines as antibacterial and cytotoxic agents. Arch Pharm 353:1900271
Tsochatzis ED, Alberto LJ, Kappenstein O, Tietz T, Hoekstra EJ (2020) Quantification of PET cyclic and linear oligomers in teabags by a validated LC-MS method – In silico toxicity assessment and consumer’s exposure. Food Chem 317:126427
Georgieva M, Bijev A, Prodanova P (2010) Synthesis and comparative study of tuberculostatic activity of pyrrole-based hydrazones related to structural variations. Pharmacia 57:3–14
Fau D, Berson A, Eugene D, Fromenty B, Fisch C, Pessayre D (1992) Mechanism for the hepatotoxicity of the antiandrogen, nilutamide. Evidence suggesting that redox cycling of this nitroaromatic drug leads to oxidative stress in isolated hepatocytes. J Pharmacol Exp Ther 263:32
Chaursiya AC, Dumpala RL (2020) A review on revision of ICH Q2 (R1) and new ICH Q14 guidance. Global J Pharma Allied Sci 1:1–6
Mateeva A, Peikova L, Kondeva-Burdina M, Georgieva M (2022) Development of new HPLC method for identification of metabolic degradation of N-pyrrolylhydrazide hydrazones with determined MAO- B activity in cellular cultures. Pharmacia 69:15–20
Tzankova D, Peikova L, Vladimirova S, Georgieva M (2019) Development and validation of RP-HPLC method for stability evaluation of model hydrazone, containing a pyrrole ring. Pharmacia 66:127–134
Patyra E, Kwiatek K (2021) Comparison of HPLC–DAD and LC–MS techniques for the determination of tetracyclines in medicated feeds using one extraction protocol. Chromatographia 84:741–749
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Methodology: M. Kondeva-Burdina, L. Peikova; Formal analysis and investigation: A. Mateeva, M. Kondeva-Burdina, L. Peikova, P. Nedialkov; Writing—original draft preparation: A. Mateeva; Writing—review and editing: P. Nedialkov, M. Georgieva; Supervision: M. Georgieva. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Competing interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
The vivarium (certificate of registration of farm No. 0072/01.08.2007) was inspected by the Bulgarian Drug Agency in order to check the husbandry conditions (No. A-11–1081/03.11.2011). All procedures were carried out in strict compliance with the requirements of the Institutional Committee for Animal Welfare and the principles set out in the European Convention for the Protection of Vertebrate Animals Used for Experimental and Other Scientific Purposes (ETS 123) (Council of Europe, 1991), throughout the experiment.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mateeva, A., Kondeva-Burdina, M., Nedialkov, P. et al. Development of Hyphenated Techniques and Network Identification Approaches for Biotransformational Evaluation of Promising Antitubercular N-pyrrolyl hydrazide-hydrazone in Isolated Rat Hepatocytes. Chromatographia 86, 497–505 (2023). https://doi.org/10.1007/s10337-023-04260-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10337-023-04260-5