Abstract
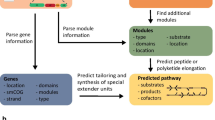
Increasing availability of new genomes and putative biosynthetic gene clusters (BGCs) has extended the opportunity to access novel chemical diversity for agriculture, medicine, environmental and industrial purposes. However, functional characterization of BGCs through heterologous expression is limited because expression may require complex regulatory mechanisms, specific folding or activation. We developed an integrated workflow for BGC characterization that integrates pathway identification, modular design, DNA synthesis, assembly and characterization. This workflow was applied to characterize multiple phenazine-modifying enzymes. Phenazine pathways are useful for this workflow because all phenazines are derived from a core scaffold for modification by diverse modifying enzymes (PhzM, PhzS, PhzH, and PhzO) that produce characterized compounds. We expressed refactored synthetic modules of previously uncharacterized phenazine BGCs heterologously in Escherichia coli and were able to identify metabolic intermediates they produced, including a previously unidentified metabolite. These results demonstrate how this approach can accelerate functional characterization of BGCs.









Similar content being viewed by others
References
Blankenfeldt W, Parsons JF (2014) The structural biology of phenazine biosynthesis. Curr Opin Struct Biol 29:26–33. https://doi.org/10.1016/j.sbi.2014.08.013
Chen et al. 2015. DIVA: more science, less DNA construction. 2015 Synth Biol Eng Evol Des (SEED) Proc Poster
Chin-A-Woeng TFC, Thomas-Oates JE, Lugtenberg BJJ, Bloemberg BJJ (2001) Introduction of the phzH gene of Pseudomonas chlororaphis PCL1391 extends the range of biocontrol ability of phenazine-1-carboxylic acid-producing Pseudomonas spp. strains. MPMI 14(8):1006–1015. https://doi.org/10.1094/mpmi.2001.14.8.1006
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97(12):6640–6645. https://doi.org/10.1073/pnas.120163297
Delaney SM, Mavrodi DV, Bonsall RF, Thomashow LS (2001) phzO, a gene for biosynthesis of 2-hydroxylated phenazine compounds in Pseudomonas aureofaciens 30-84. J Bacteriol 183(1):318–327. https://doi.org/10.1128/JB.183.1.318-327.2001
Kushwaha M, Salis HM (2015) A portable expression resource for engineering cross-species genetic circuits and pathways. Nat Commun 6:7832. https://doi.org/10.1038/ncomms8832
Hadjithomas M, Chen IMA, Chu K et al (2015) IMG-ABC: a knowledge base to fuel discovery of biosynthetic gene clusters and novel secondary metabolites. MBio 6(4):e00932-15. https://doi.org/10.1128/mBio.00932-15
Hillson JN, Rosengarten RD, Keasling JD (2012) j5 DNA assembly design automation software. ACS Synth Biol 1(1):14–21. https://doi.org/10.1021/sb2000116
Mavrodi DV, Ksenzenko VN, Bonsall RF, Cook RJ, Boronin AM, Thomashow LS (1998) A seven-gene locus for synthesis of phenazine-1-carboxylic acid by Pseudomonas fluorescens 2-79. J Bacteriol 180(9):2541–2548
Mavrodi DV, Bonsall RF, Delaney SM, Soule MJ, Phillips G, Thomashow LS (2001) Functional analysis of genes for biosynthesis of pyocyanin and phenazine-1-carboxamide from Pseudomonas aeruginosa PAO1. J Bacteriol 183:6454–6465. https://doi.org/10.1128/JB.183.21.6454-6465.2001
Mavrodi DV, Blankenfeldt W, Thomashow LS (2006) Phenazine compounds in fluorescent Pseudomonas spp. biosynthesis and regulation. Annu Rev Phytopathol 44:417–445. https://doi.org/10.1146/annurev.phyto.44.013106.145710
Mavrodi DV, Peever TL, Mavrodi OV et al (2010) Diversity and evolution of the phenazine biosynthesis pathway. Appl Environ Microbiol 76(3):866–879. https://doi.org/10.1128/AEM.02009-09
Oberortner E, Cheng JF, Hillson NJ, Deutsch S (2017) Streamlining the design-to-build transition with build-optimization software tools. ACS Synth Biol 6(3):485–496. https://doi.org/10.1021/acssynbio.6b00200
Owen JG, Reddy BVB, Ternei MA et al (2013) Mapping gene clusters within arrayed metagenomic libraries to expand the structural diversity of biomedically relevant natural products. Proc Natl Acad Sci 110(29):11797–11802. https://doi.org/10.1073/pnas.1222159110
Pierson LS, Thomashow LS (1992) Cloning and heterologous expression of the phenazine biosynthetic locus from Pseudomonas aureofaciens 30-84. Mol Plant Microbe Interact 5:330–339. https://doi.org/10.1094/MPMI-5-330
Pierson EA, Wang D, Pierson LS III (2013) Roles and regulation of phenazines in the biological control strain Pseudomonas chlororaphis 30-84. In: Chincholkar S, Thomashow LS (eds) Microbial phenazines. Springer, Berlin, pp 141–162
Glasser NR, Kern SE, Newmann DK (2007) Phenazine redox cycling enhances anaerobic survival in Pseudomonas aeruginosa by facilitating generation of ATP and a proton-motive force. Mol Microbiol 92(2):1365–2958. https://doi.org/10.1111/mmi.12566
Ren H, Wang B, Zhao H (2017) Breaking the silence: new strategies for discovering novel natural products. Curr Opin Biotechnol 48:21–27. https://doi.org/10.1016/j.copbio.2017.02.008
Rui Z, Ye M, Wang S et al (2012) Insights into a divergent phenazine biosynthetic pathway governed by a plasmid-born esmeraldin gene cluster. Chem Biol 19:1116–1125. https://doi.org/10.1016/j.chembiol.2012.07.025
Smanski MJ, Zhaou H, Claesen J, Shen B, Fischbach MA, Voigt CA (2016) Synthetic biology to access and expand nature’s chemical diversity. Nat Rev Microbiol 14(3):135–149. https://doi.org/10.1038/nrmicro.2015.24
Santos CNS, Yoshikuni Y (2014) Engineering complex biological systems in bacteria through recombinase-assisted genome engineering. Nat Protoc 9(6):1320–1336. https://doi.org/10.1038/nprot.2014.084
Thomashow LS, Bonsall RF, Weller DM (2002) Antibiotic production by soil and rhizosphere microbes in situ. In: Hurst CJ (ed) Manual of environmental microbiology. ASM Press, Washington, pp 638–647
Thomashow LS (2013) Phenazines in the environment: microbes, habitats, and ecological relevance. In: Chincholkar S, Thomashow LS (eds) Microbial phenazines. Springer, Berlin, pp 199–216
Cheng X, Zhang X, Pflugrath JW, Studier FW (1994) The structure of bacteriophage T7 lysozyme, a zinc amidase and an inhibitor of T7 RNA polymerase. Proc Natl Acad Sci USA 91(9):4034–4038
Ikeda RA (1992) The efficiency of promoter clearance distinguishes T7 class II and class III promoters. J Biol Chem 267(16):11322–11328
Gibson GG, Young L, Chuang RY, Venter JC, Hutchison CA III, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345. https://doi.org/10.1038/nmeth.1318
Kehmann F, Cherpillod F (1924) Neue Synthesen in der Gruppe der Chinon-imid-farbstoffe. V. Über Synthesen ausgehend von Oxybenzochinon. Helv Chim Acta 7(1):973–980. https://doi.org/10.1002/hlca.192400701122
Mcllwain H (1937) The phenazine series. Part VI. Reactions of alkyl phenazonium salts; the phenazyls. J Chem Soc 0:1704–1711. https://doi.org/10.1039/JR9370001704
Acknowledgements
The work conducted by the U.S. Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, operated by Lawrence Berkeley National Laboratory under Contract no. DE-AC02-05CH11231. We would like to thank Dr. Wayne Reeve of Murdoch University for kindly providing us with a live strain of Rhizobium leguminosarum bv. trifolii CC283b.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Coates, R.C., Bowen, B.P., Oberortner, E. et al. An integrated workflow for phenazine-modifying enzyme characterization. J Ind Microbiol Biotechnol 45, 567–577 (2018). https://doi.org/10.1007/s10295-018-2025-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2025-5