Abstract
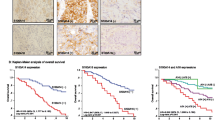
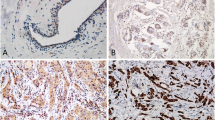
Tetraspanins have been implicated in multiple biological functions including protein networking and cell signaling. NET-6 (TSPAN 13) has been demonstrated to be a tumor suppressor gene in breast cancer, while CD151 is more likely to act as an oncogene. However, the biological function of both proteins is still inconclusive. Immunohistochemistry was used to analyze the expression of NET-6 and CD151 proteins in breast tumors and benign epithelial cells. The cellular expression of both markers was correlated with HER2, ER, and PR status as well as tumor grade, Ki-67 scores, invasion, and metastasis. Expression of NET-6 and CD151 was variable both in tumors and in benign epithelial cells. Expression of NET-6 and CD151 was stronger in tumors than in benign epithelial cells. The expression of NET-6 was also stronger in HER2-negative, low-grade, lymphovascular invasion-negative, and non-metastatic breast tumors. There was no correlation between NET-6 expression and ER, or PR, or triple-negative status. There was no correlation between CD151 expression and HER2, ER, PR, or triple-negative status, tumor grade, or Ki-67 scores, invasion, and metastasis. The expression of tetraspanins NET-6 and CD151 may indicate an alteration of their biological function during neoplastic transformation. NET-6 expression in tumors might be a potential marker indicating the outcome of breast cancer.


Similar content being viewed by others
References
Termini CM, Gillette JM. Tetraspanins function as regulators of cellular signaling. Front Cell Dev Biol. 2017;5:34. https://doi.org/10.3389/fcell.2017.00034.
Charrin S, Jouannet S, Boucheix C, Rubinstein E. Tetraspanins at a glance. J Cell Sci. 2014;127:3641–8. https://doi.org/10.1242/jcs.154906.
Yang YG, Sari IN, Zia MF, Lee SR, Song SJ, Kwon HY. Tetraspanins: spanning from solid tumors to hematologic malignancies. Exp Hematol. 2016;44:322–8. https://doi.org/10.1016/j.exphem.2016.02.006.
van Deventer SJ, Dunlock VE, van Spriel AB. Molecular interactions shaping the tetraspanin web. Biochem Soc Trans. 2017;45:741–50. https://doi.org/10.1042/BST20160284.
Khushman M, Bhardwaj A, Patel GK, et al. Exosomal markers (CD63 and CD9) expression pattern using immunohistochemistry in resected malignant and nonmalignant pancreatic specimens. Pancreas. 2017;46:782–8. https://doi.org/10.1097/MPA.0000000000000847.
Yu J, Lee CY, Changou CA, Cedano-Prieto DM, Takada YK, Takada Y. The CD9, CD81, and CD151 EC2 domains bind to the classical RGD-binding site of integrin alphavbeta3. Biochem J. 2017;474:589–96. https://doi.org/10.1042/BCJ20160998.
Medrano M, Communal L, Brown KR, et al. Interrogation of functional cell-surface markers identifies CD151 dependency in high-grade serous ovarian cancer. Cell Rep. 2017;18:2343–58. https://doi.org/10.1016/j.celrep.2017.02.028.
Kwon HJ, Choi JE, Kang SH, Son Y, Bae YK. Prognostic significance of CD9 expression differs between tumour cells and stromal immune cells, and depends on the molecular subtype of the invasive breast carcinoma. Histopathology. 2017;70:1155–65. https://doi.org/10.1111/his.13184.
Di Giacomo V, Tian TV, Mas A, et al. DeltaNp63alpha promotes adhesion of metastatic prostate cancer cells to the bone through regulation of CD82. Oncogene. 2017;36:4381–92. https://doi.org/10.1038/onc.2017.42.
Zhu Y, Ailane N, Sala-Valdes M, et al. Multi-factorial modulation of colorectal carcinoma cells motility—partial coordination by the tetraspanin Co-029/tspan8. Oncotarget. 2017;8:27454–70. https://doi.org/10.18632/oncotarget.16247.
Huang H, Sossey-Alaoui K, Beachy SH, Geradts J. The tetraspanin superfamily member NET-6 is a new tumor suppressor gene. J Cancer Res Clin Oncol. 2007;133:761–9. https://doi.org/10.1007/s00432-007-0221-1.
Huang H, Groth J, Sossey-Alaoui K, Hawthorn L, Beall S, Geradts J. Aberrant expression of novel and previously described cell membrane markers in human breast cancer cell lines and tumors. Clin Cancer Res. 2005;11:4357–64. https://doi.org/10.1158/1078-0432.CCR-04-2107.
Romanska HM, Potemski P, Kusinska R, Kopczynski J, Sadej R, Kordek R. Expression of CD151/Tspan24 and integrin alpha 3 complex in aid of prognostication of HER2-negative high-grade ductal carcinoma in situ. Int J Clin Exp Pathol. 2015;8:9471–8.
Romanska HM, Potemski P, Krakowska M, et al. Lack of CD151/integrin alpha3beta1 complex is predictive of poor outcome in node-negative lobular breast carcinoma: opposing roles of CD151 in invasive lobular and ductal breast cancers. Br J Cancer. 2015;113:1350–7. https://doi.org/10.1158/1078-0432.CCR-04-2107.
Tilghman J, Schiapparelli P, Lal B, et al. Regulation of glioblastoma tumor-propagating cells by the integrin partner tetraspanin CD151. Neoplasia. 2016;18:185–98. https://doi.org/10.1016/j.neo.2016.02.003.
Fisher OM, Levert-Mignon AJ, Lehane CW, et al. CD151 gene and protein expression provides independent prognostic information for patients with adenocarcinoma of the esophagus and gastroesophageal junction treated by esophagectomy. Ann Surg Oncol. 2016;23:746–54. https://doi.org/10.1245/s10434-016-5504-9.
Wadkin J, Patten DA, Kamarajah SK, et al. CD151 supports VCAM-1-mediated lymphocyte adhesion to liver endothelium and is upregulated in chronic liver disease and hepatocellular carcinoma. Am J Physiol Gastrointest Liver Physiol. 2017;313:G138–49. https://doi.org/10.1152/ajpgi.00411.2016.
Nienstedt JC, Grobe A, Lebok P, et al. CD151 expression is frequent but unrelated to clinical outcome in head and neck cancer. Clin Oral Investig. 2017;21:1503–8. https://doi.org/10.1007/s00784-016-1911-3.
Nishimura R, Okamoto N, Satou M, Kojima K, Tanaka S. HER 2 immunohistochemistry for breast cancer cell blocks can be used in the same way as that used for histological specimens. Diagn Cytopathol. 2016;44:274–9. https://doi.org/10.1002/dc.23433.
Iwai K, Ishii M, Ohshima S, Miyatake K, Saeki Y. Expression and function of transmembrane-4 superfamily (tetraspanin) proteins in osteoclasts: reciprocal roles of Tspan-5 and NET-6 during osteoclastogenesis. Allergol Int. 2007;56:457–63. https://doi.org/10.2332/allergolint.O-07-488.
Arencibia JM, Martin S, Perez-Rodriguez FJ, Bonnin A. Gene expression profiling reveals overexpression of TSPAN13 in prostate cancer. Int J Oncol. 2009;34:457–63.
Sossey-Alaoui K, Vieira L, David D, Boavida MG, Cowell JK. Molecular characterization of a 7p15-21 homozygous deletion in a Wilms tumor. Genes Chromosomes Cancer. 2003;36:1–6. https://doi.org/10.1002/gcc.10133.
Zevian SC, Johnson JL, Winterwood NE, et al. CD151 promotes alpha3beta1 integrin-dependent organization of carcinoma cell junctions and restrains collective cell invasion. Cancer Biol Ther. 2015;16:1626–40. https://doi.org/10.1080/15384047.2015.1095396.
Yue S, Mu W, Erb U, Zoller M. The tetraspanins CD151 and Tspan8 are essential exosome components for the crosstalk between cancer initiating cells and their surrounding. Oncotarget. 2015;6:2366–84. https://doi.org/10.18632/oncotarget.2958.
Acknowledgements
This study was funded by grants from the Guangxi Bureau of Science and Technology, No. 1598013-3 (PI: Huayi Huang) and No. 2017GXNSFAA198043 (PI: Huayi Huang) and a grant from The Health and Family Planning Commission of Guangxi Zhuang Autonomous Region, No. Z2015353 (PI: Huayi Huang).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The study was approved by the Institute Review Boards of Guangzhou Huayin Medical Laboratory and the People’s Hospital of Guangxi Zhuang Autonomous Region for the use of human materials.
Informed consent
This study was waived for the patients’ informed consent from the above-mentioned institutions because the study was a retrospective with pathology archived tissues.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jiang, L., Zhang, X., Geradts, J. et al. Expression of tetraspanins NET-6 and CD151 in breast cancer as a potential tumor biomarker. Clin Exp Med 19, 377–384 (2019). https://doi.org/10.1007/s10238-019-00554-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-019-00554-x