Abstract
Paracetamol is a relatively safe analgesia/antipyretic drug without the risks of addiction, dependence, tolerance, and withdrawal when used alone. However, when administrated in an opioid/paracetamol combination product, which often contains a large quantity of paracetamol, it can be potentially dangerous due to the risk of hepatotoxicity. Paracetamol is known to be metabolized into N-(4-hydroxyphenyl)-arachidonamide (AM404) via fatty acid amide hydrolase (FAAH) and into N-acetyl-p-benzoquinone imine (NAPQI) via cytochrome P450 (CYP) enzymes. However, the underlying mechanism of paracetamol is still unclear. In addition, paracetamol has the potential to interact with other drugs that are also involved with CYP family enzymes (inducer/inhibitor/substrate), an example being illicit drugs. In our present work, we looked into the relationship between paracetamol and its metabolites (AM404 and NAPQI) using molecular docking and molecular dynamics (MD) simulations. We first carried out a series of molecular docking studies between paracetamol/AM404/NAQPI and their reported targets, including CYP 2E1, FAAH, TRPA1, CB1, and TRPV1. Subsequently, we performed MD simulations and energy decomposition for CB1-AM404, TRPV1-AM404, and TRPV1-NAPQI for further investigation of the dynamics interactions. Finally, we summarized and discussed the reported drug–drug interactions between paracetamol and central nervous system drugs, especially illicit drugs. Overall, we are able to provide new insights into the structural and functional roles of paracetamol and its metabolites that can inform the potential prevention and treatment of paracetamol overdose.

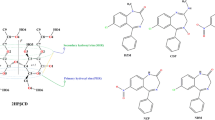
Paracetamol and its metabolites






Similar content being viewed by others
References
Ramadan WH et al (2016) Trends of acetaminophen overuse among ambulatory patients in Lebanon: an observational study. Int J Res Pharm Sci 6: 41–45
Mallet C, Eschalier A, Daulhac L (2017) Paracetamol: update on its analgesic mechanism of action. In: Maldonado C (ed) Pain relief. From analgesics to alternative therapies. InTech. Chapter 10. https://doi.org/10.5772/63264
Hylands-White N, Duarte RV, Raphael JH (2017) An overview of treatment approaches for chronic pain management. Rheumatol Int 37(1):29–42
Stephan BC, Parsa FD (2016) Avoiding opioids and their harmful side effects in the postoperative patient: exogenous opioids, endogenous endorphins, wellness, mood, and their relation to postoperative pain. Hawai’i J Med Public Health 75(3):63
Chan HS et al (2017) Designing safer analgesics via μ-opioid receptor pathways. Trends Pharmacol Sci 38(11):1016–1037
Dart RC, Green JL (2016) The prescription paradox of acetaminophen safety. Pharmacoepidemiol Drug Saf 25(5):599–601
Blieden M et al (2014) A perspective on the epidemiology of acetaminophen exposure and toxicity in the United States. Expert Rev Clin Pharmacol 7(3):341–348
McCarthy DM et al (2014) Patient recall of health care provider counseling for opioid-acetaminophen prescriptions. Pain Med 15(10):1750–1756
Vosler PS et al (2014) Clinical and pathologic characteristics of intranasal abuse of combined opioid-acetaminophen medications. Int Forum Allergy Rhinol 4(10):839–844
Rojas KM, Li H (2017) Adverse events and over-the-counter (OTC) drugs: is inappropriate labeling the problem?—The case of acetaminophen. In: Proceedings of the Human Factors and Ergonomics Society Annual Meeting. Sage, Los Angeles, pp 676–680
Stueber T et al (2018) Activation of the capsaicin-receptor TRPV1 by the acetaminophen metabolite N-arachidonoylaminophenol results in cytotoxicity. Life Sci 194:67–74
Klinger-Gratz PP et al (2018) Acetaminophen relieves inflammatory pain through CB1 cannabinoid receptors in the rostral ventromedial medulla. J Neurosci 38(2):322–334
Mallet C et al (2010) TRPV1 in brain is involved in acetaminophen-induced antinociception. PLoS One 5(9):e12748
Gentry C, Andersson DA, Bevan S (2015) TRPA1 mediates the hypothermic action of acetaminophen. Sci Rep 5:12771
Tóth A, Blumberg PM, Boczán J (2009) Anandamide and the vanilloid receptor (TRPV1). Vitam Horm 81:389–419
Sharma CV et al (2017) First evidence of the conversion of paracetamol to AM404 in human cerebrospinal fluid. J Pain Res 10:2703
Alexander S, Kendall D (2007) The complications of promiscuity: endocannabinoid action and metabolism. Br J Pharmacol 152(5):602–623
Busquets-Garcia A, Bains J, Marsicano G (2018) CB 1 receptor signaling in the brain: extracting specificity from ubiquity. Neuropsychopharmacology 43(1):4
Howlett A (2005) Cannabinoid receptor signaling. In: Pertwee RG (ed) Cannabinoids. Springer, Berlin, pp 53–79
Lu D, Potter D (2017) Cannabinoids and the cannabinoid receptors: an overview. In: Preedy VR (ed) Handbook of cannabis and related pathologies: biology, pharmacology, diagnosis, and treatment. Elsevier, Amsterdam, pp 553–563
Seltzman HH et al (2016) Peripherally selective cannabinoid 1 receptor (CB1R) agonists for the treatment of neuropathic pain. J Med Chem 59(16):7525–7543
Iring A, Hricisák L, Benyó Z (2017) CB1 receptor-mediated respiratory depression by endocannabinoids. Respir Physiol Neurobiol 240:48–52
Barrière DA et al (2013) Fatty acid amide hydrolase-dependent generation of antinociceptive drug metabolites acting on TRPV1 in the brain. PLoS One 8(8):e70690
Mallet C et al (2008) Endocannabinoid and serotonergic systems are needed for acetaminophen-induced analgesia. Pain 139(1):190–200
Hama AT, Sagen J (2010) Cannabinoid receptor-mediated antinociception with acetaminophen drug combinations in rats with neuropathic spinal cord injury pain. Neuropharmacology 58(4–5):758–766
Evans R, Scott R, Ross R (2007) Chronic exposure of sensory neurones to increased levels of nerve growth factor modulates CB1/TRPV1 receptor crosstalk. Br J Pharmacol 152(3):404–413
Weinhold P et al (2010) TRPA1 receptor induced relaxation of the human urethra involves TRPV1 and cannabinoid receptor mediated signals, and cyclooxygenase activation. J Urol 183(5):2070–2076
Eberhardt MJ et al (2017) Reactive metabolites of acetaminophen activate and sensitize the capsaicin receptor TRPV1. Sci Rep 7(1):12775
Rosenbaum T, A Jara-Oseguera (2012) TRPV1 in cell signaling: molecular mechanisms of function and modulation. In: Kamkin A, Lozinsky I (eds) Mechanically gated channels and their regulation. Springer, Berlin, pp 69–102
Zhang X et al (2014) Nitro-oleic acid desensitizes TRPA1 and TRPV1 agonist responses in adult rat DRG neurons. Exp Neurol 251:12–21
Akopian AN et al (2007) Transient receptor potential TRPA1 channel desensitization in sensory neurons is agonist dependent and regulated by TRPV1-directed internalization. J Physiol 583(1):175–193
Libert F et al (2004) Acetaminophen: a central analgesic drug that involves a spinal tropisetron-sensitive, non–5-HT3 receptor-mediated effect. Mol Pharmacol 66(3):728–734
Park J-Y, Harris D (2003) Construction and assessment of models of CYP2E1: predictions of metabolism from docking, molecular dynamics, and density functional theoretical calculations. J Med Chem 46(9):1645–1660
Cao E et al (2013) TRPV1 structures in distinct conformations reveal activation mechanisms. Nature 504(7478):113
Feng Z et al (2015) Structural insight into tetrameric hTRPV1 from homology modeling, molecular docking, molecular dynamics simulation, virtual screening and bioassay validations. J Chem Inf Model 54(9):2483–2499
Feng Z et al (2016) Multi-functional diarylurea small molecule inhibitors of TRPV1 with therapeutic potential for neuroinflammation. AAPS J 18(4):898–913
Hua T et al (2017) Crystal structures of agonist-bound human cannabinoid receptor CB 1. Nature 547(7664):468
Marti-Renom MA et al (2000) Comparative protein structure modeling of genes and genomes. Annu Rev Biophys Biomol Struct 29(1):291–325
Jain AN (1996) Scoring noncovalent protein-ligand interactions: a continuous differentiable function tuned to compute binding affinities. J Comput Aided-Mol Des 10(5):427–440
Feng Z et al (2015) Structural insight into tetrameric hTRPV1 from homology modeling, molecular docking, molecular dynamics simulation, virtual screening and bioassay validations. J Chem Inf Model 55(3):572–588
Chen J-Z, Wang J, Xie X-Q (2007) GPCR structure-based virtual screening approach for CB2 antagonist search. J Chem Inf Model 47(4):1626–1637
Feng Z et al (2014) Modeling, molecular dynamics simulation, and mutation validation for structure of cannabinoid receptor 2 based on known crystal structures of GPCRs. J Chem Inf Model 54(9):2483–2499
Feng Z et al (2015) Design and activity of AP endonuclease-1 inhibitors. J Chem Biol 8(3):79–93
Jo S et al (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29(11):1859–1865
Wu EL et al (2014) CHARMM-GUI membrane builder toward realistic biological membrane simulations. J Comput Chem 35(27):1997–2004
Maier JA et al (2015) ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J Chem Theory Comput 11(8):3696–3713
Dickson CJ et al (2014) Lipid14: the amber lipid force field. J Chem Theory Comput 10(2):865–879
Jorgensen WL et al (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935
Jakalian A, Jack DB, Bayly CI (2002) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J Comput Chem 23(16):1623–1641
Wang J et al (2004) Development and testing of a general amber force field. J Comput Chem 25(9):1157–1174
Wang J et al (2006) Automatic atom type and bond type perception in molecular mechanical calculations. J Mol Graph Model 25(2):247–260
Götz AW et al (2012) Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born. J Chem Theory Comput 8(5):1542–1555
Salomon-Ferrer R et al (2013) Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh Ewald. J Chem Theory Comput 9(9):3878–3888
Case D et al (2016) AMBER 2016, University of California, San Francisco
Loncharich RJ, Brooks BR, Pastor RW (1992) Langevin dynamics of peptides: the frictional dependence of isomerization rates of N-acetylalanyl-N′-methylamide. Biopolymers 32(5):523–535
Izaguirre JA et al (2001) Langevin stabilization of molecular dynamics. J Chem Phys 114(5):2090–2098
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N· log (N) method for Ewald sums in large systems. J Chem Phys 98(12):10089–10092
Essmann U et al (1995) A smooth particle mesh Ewald method. J Chem Phys 103(19):8577–8593
Ryckaert J-P, Ciccotti G, Berendsen HJ (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341
Wang J, Hou T (2012) Develop and test a solvent accessible surface area-based model in conformational entropy calculations. J Chem Inf Model 52(5):1199–1212
Hawkins GD, Cramer CJ, Truhlar DG (1996) Parametrized models of aqueous free energies of solvation based on pairwise descreening of solute atomic charges from a dielectric medium. J Phys Chem 100(51):19824–19839
Kollman PA et al (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33(12):889–897
Cruz JN et al (2018) Molecular dynamics simulation and binding free energy studies of novel leads belonging to the benzofuran class inhibitors of Mycobacterium tuberculosis polyketide synthase 13. J Biomol Struct Dynamics. https://doi.org/10.1080/07391102.2018.1462734
Tsui V, Case DA (2000) Theory and applications of the generalized born solvation model in macromolecular simulations. Biopolymers 56(4):275–291
Bashford D, Case DA (2000) Generalized born models of macromolecular solvation effects. Annu Rev Phys Chem 51(1):129–152
Sitkoff D, Sharp KA, Honig B (1994) Accurate calculation of hydration free energies using macroscopic solvent models. J Phys Chem 98(7):1978–1988
Still WC et al (1990) Semianalytical treatment of solvation for molecular mechanics and dynamics. J Am Chem Soc 112(16):6127–6129
Weiser J, Shenkin PS, Still WC (1999) Approximate atomic surfaces from linear combinations of pairwise overlaps (LCPO). J Comput Chem 20(2):217–230
Hu J et al (2016) Difference and influence of inactive and active states of cannabinoid receptor subtype CB2: from conformation to drug discovery. J Chem Inf Model 56(6):1152–1163
Phipps MJ et al (2015) Energy decomposition analysis approaches and their evaluation on prototypical protein–drug interaction patterns. Chem Soc Rev 44(10):3177–3211
McDaniel JG, Schmidt J (2016) Next-generation force fields from symmetry-adapted perturbation theory. Annu Rev Phys Chem 67:467–488
DeVore NM et al (2012) Structural comparison of cytochromes P450 2A6, 2A13, and 2E1 with pilocarpine. FEBS J 279(9):1621–1631
Hartman JH et al (2015) Cooperativity in CYP2E1 metabolism of acetaminophen and styrene mixtures. Biochem Pharmacol 97(3):341–349
Bertolacci L et al (2013) A binding site for nonsteroidal anti-inflammatory drugs in fatty acid amide hydrolase. J Am Chem Soc 135(1):22–25
Andersson DA et al (2011) TRPA1 mediates spinal antinociception induced by acetaminophen and the cannabinoid Δ 9-tetrahydrocannabiorcol. Nat Commun 2:551
Paulsen CE et al (2015) Structure of the TRPA1 ion channel suggests regulatory mechanisms. Nature 520(7548):511
Sinning C et al (2008) New analgesics synthetically derived from the paracetamol metabolite N-(4-hydroxyphenyl)-(5 Z, 8 Z, 11 Z, 14 Z)-icosatetra-5, 8, 11, 14-enamide. J Med Chem 51(24):7800–7805
Hua T et al (2016) Crystal structure of the human cannabinoid receptor CB 1. Cell 167(3):750–762.e14
Shao Z et al (2016) High-resolution crystal structure of the human CB1 cannabinoid receptor. Nature 540(7634):602
Acknowledgments
The authors would like to acknowledge funding support to the X.-Q. Xie laboratory from the NIH NIDA (P30 DA035778A1), the National Institutes of Health (NIH) (R01 DA025612), and the Department of Defense (W81XWH-16-1-0490), funding support to the J.M. Wang laboratory from the NIH of the United States (R01-GM079383, R21-GM097617), and funding to Y.Q. Wang from the National Natural Science Foundation of China (31400667), Chongqing Municipal Education Commission Science and Technology Research Project (KJ1500902, KJ1600908) and Chongqing Research Program of Basic Research and Frontier Technology (cstc2014jcyjA10044, cstc2018jcyjAX0683). Computational support from the Center for Research Computing of University of Pittsburgh, and the Extreme Science and Engineering Discovery Environment (CHE090098), are acknowledged.
Author information
Authors and Affiliations
Contributions
X.-Q. X. and Z.F. designed the research. Y.W., W.L. and N.W. performed the research, carried out the analysis and wrote the manuscript. X.H. and J.W. performed MD simulation and carried out the analysis.
Corresponding authors
Rights and permissions
About this article
Cite this article
Wang, Y., Lin, W., Wu, N. et al. An insight into paracetamol and its metabolites using molecular docking and molecular dynamics simulation. J Mol Model 24, 243 (2018). https://doi.org/10.1007/s00894-018-3790-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3790-9