Abstract
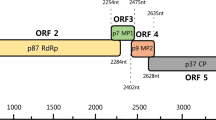
The complete genome sequence of a new caulimovirus in Pueraria montana was determined using high-throughput sequencing. The 7,572 nucleotide genome of pueraria virus A (PVA) contains genes that encode a movement protein, an aphid transmission factor, a virion-associated protein, a coat protein, a protease + reverse transcriptase + ribonuclease H, and a transactivator/viroplasmin protein, as well as two intergenic regions, which are all common features of members of the genus Caulimovirus. A sequence alignment revealed that the complete genome of PVA shares 66.82% nucleotide sequence identity with strawberry vein banding virus (GenBank accession no. KX249738.1). The results of phylogenetic analysis and the observation that the nucleotide sequence of the polymerase coding region differed by more than 20% indicated that PVA is a member of a new species the genus Caulimovirus, family Caulimoviridae.


Similar content being viewed by others
References
Teycheney P-Y, Geering ADW, Dasgupta I et al (2020) ICTV virus taxonomy profile: caulimoviridae. J Gen Virol 101:1025–1026. https://doi.org/10.1099/jgv.0.001497
Mart K, Jonas B, Coffin MJ et al (2021) Ortervirales: new virus order unifying five families of reverse-transcribing viruses. J Virol 92:e00515-e518. https://doi.org/10.1128/JVI.00515-18
Wang S, Zhang S, Wang S et al (2020) A comprehensive review on Pueraria: insights on its chemistry and medicinal value. Biomed Pharmacother 131:110734. https://doi.org/10.1016/J.BIOPHA.2020.110734
Zhang J, Wu ZJ (2012) First Report of Kudzu mosaic virus on Pueraria montana (Kudzu) in China. Plant Dis 97:148. https://doi.org/10.1094/PDIS-07-12-0671-PDN
Khankhum S, Bollich P, Valverde RA (2013) First report of Tobacco ringspot virus infecting kudzu (Pueraria montana) in Louisiana. Plant Dis 97:561. https://doi.org/10.1094/PDIS-10-12-0933-PDN
Zhou J, Aboughanem-Sabanadzovic N, Sabanadzovic S, Tzanetakis IE (2018) First report of soybean vein necrosis virus infecting Kudzu (Pueraria montana) in the United States of America. Plant Dis 102:1674. https://doi.org/10.1094/PDIS-01-18-0042-PDN
Liu H, Zhao F, Qiao Q et al (2021) Complete genome sequence of a divergent sweet potato chlorotic stunt virus isolate infecting Calystegia hederacea in China. Arch Virol 166:2037–2040. https://doi.org/10.1007/s00705-021-05076-0
Lim S, Baek D, Igori D, Moon JS (2017) Complete genome sequence of a putative new caulimovirus which exists as endogenous pararetroviral sequences in Angelica dahurica. Arch Virol 162:3837–3842. https://doi.org/10.1007/s00705-017-3517-8
Lim S, Igori D, Zhao F et al (2015) Complete genome sequence of a tentative new caulimovirus from the medicinal plant Atractylodes macrocephala. Arch Virol 160:3127–3131. https://doi.org/10.1007/s00705-015-2576-y
Pooggin MM, Ryabova LA (2018) Ribosome shunting, polycistronic translation, and evasion of antiviral defenses in plant pararetroviruses and beyond. Front Microbiol 9:644. https://doi.org/10.3389/fmicb.2018.00644
Menéndez-Arias L, Sebastián-Martín A, Álvarez M (2017) Viral reverse transcriptases. Virus Res 234:153–176. https://doi.org/10.1016/j.virusres.2016.12.019
Finn RD, Bateman A, Clements J et al (2014) Pfam: the protein families database. Nucleic Acids Res 42:D222–D230. https://doi.org/10.1093/nar/gkt1223
Petrzik K, Beneš V, Mráz I et al (1998) Strawberry vein banding virus—definitive member of the genus Caulimovirus. Virus Genes 16:303–305. https://doi.org/10.1023/A:1008039024963
Richins RD, Scholthof HB, Shepherd RJ (1987) Sequence of figwort mosaic virus DNA (caulimovirus group). Nucleic Acids Res. https://doi.org/10.1093/nar/15.20.8451
Leclerc D, Stavolone L, Meier E et al (2001) The product of ORF III in cauliflower mosaic virus interacts with the viral coat protein through its C-terminal proline rich domain. Virus Genes 22:159–165. https://doi.org/10.1023/A:1008121228637
Pahalawatta V, Druffel KL, Wyatt SD et al (2008) Genome structure and organization of a member of a novel and distinct species of the genus Caulimovirus associated with dahlia mosaic. Arch Virol 153:733–738. https://doi.org/10.1007/s00705-008-0043-8
Eid S, Almeyda CV, Saar DE et al (2011) Genomic characterization of pararetroviral sequences in wild Dahlia spp. in natural habitats. Arch Virol 156:2079. https://doi.org/10.1007/s00705-011-1076-y
Pappu HR, Druffel KL (2009) Use of conserved genomic regions and degenerate primers in a PCR-based assay for the detection of members of the genus Caulimovirus. J Virol Methods 157:102–104. https://doi.org/10.1016/J.JVIROMET.2008.11.014
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547
Acknowledgements
This work was supported by IPET (Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries; Project No. AGC1762111), Ministry of Agriculture, Food and Rural Affairs, Republic of Korea. Also, this study was supported by the Animal and Plant Quarantine Agency (No. FDM0012211) Republic of Korea. We thank Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies of involving human participants or animals.
Additional information
Handling Editor: Elvira Fiallo-Olivé.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gudeta, W.F., Igori, D., Belete, M. . et al. Complete genome sequence of pueraria virus A, a new member of the genus Caulimovirus. Arch Virol 167, 1481–1485 (2022). https://doi.org/10.1007/s00705-022-05431-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-022-05431-9