Abstract
Here, we report the discovery and molecular characterization of a novel mitovirus isolated from tissues of Lagenaria siceraria, named "Lagenaria siceraria associated mitovirus 1" (LsaMV1). Next-generation sequencing and adapter-ligation-mediated amplification were used to obtain the full-length genome sequence of LsaMV1. The genome of LsaMV1 is 3,098 nucleotides (nt) long and contains an opening reading frame (ORF) encoding a putative RNA-dependent RNA polymerase (RdRP). Homology searches and phylogenetic analysis of the 855-aa RdRP suggested that LsaMV1 is a member of a new species in the family Mitoviridae. This is the first report of a member of the family Mitoviridae associated with the important summer vegetable bottle gourd.


Similar content being viewed by others
Availability of data and material
The complete genome sequence of LsaMV1 has been deposited in the NCBI database with accession number MZ032029.
References
Khan MR, Khan MW (1998) Interactive effects of ozone and powdery mildew (Sphaerotheca fuliginea) on bottle gourd (Lagenaria siceraria). Agric Ecosyst Environ 70:109–118
Maddu H, Gasti V, Shantappa T, Mulge R, Shirol A, Mastiholi A, Kulkarni M (2012) Evaluation of bottle gourd genotypes [Lagenaria siceraria (Mol.) Standl.] for various horticultural characters*. Karnataka J Agric Sci 25:241–244
Polito L, Bortolotti M, Maiello S, Battelli MG, Bolognesi A (2016) Plants producing ribosome-inactivating proteins in traditional medicine. Molecules 21:1560
Liu H, Fu Y, Xie J, Cheng J, Ghabrial SA, Li G, Peng Y, Yi X, Jiang D (2012) Evolutionary genomics of mycovirus-related dsRNA viruses reveals cross-family horizontal gene transfer and evolution of diverse viral lineages. BMC Evol Biol 12:91
Pearson MN, Beever RE, Boine B, Arthur K (2009) Mycoviruses of filamentous fungi and their relevance to plant pathology. Mol Plant Pathol 10:115–128
Hillman BI, Cai G (2013) Chapter Six - The Family Narnaviridae: Simplest of RNA Viruses. In: Ghabrial SA (ed) Advances in Virus Research. Academic Press, pp 149–176
Osawa S, Jukes TH, Watanabe K, Muto A (1992) Recent evidence for evolution of the genetic code. Microbiol Rev 56:229–264
Jukes TH, Osawa S (1991) Recent evidence for evolution of the genetic code. In: Osawa S, Honjo T (Eds) Evolution of life: Fossils, Molecules, and Culture. Springer Japan, Tokyo, pp 79–95
Nibert ML (2017) Mitovirus UGA(Trp) codon usage parallels that of host mitochondria. Virology 507:96–100
Chiapello M, Rodriguez-Romero J, Ayllon MA, Turina M (2020) Analysis of the virome associated to grapevine downy mildew lesions reveals new mycovirus lineages. Virus Evol 6:veaa058
Chiba Y, Oiki S, Yaguchi T, Urayama SI, Hagiwara D (2021) Discovery of divided RdRP sequences and a hitherto unknown genomic complexity in fungal viruses. Virus Evol 7:veaa101
Katoh K, Standley D (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973
Darriba D, Taboada GL, Doallo R, Posada D (2011) ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27:1164–1165
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (eds) (2011) Virus taxonomy: ninth report of the international committee on taxonomy of viruses. Elsevier, Amsterdam , pp 1055–1060
Acknowledgements
This work was supported by the Wuhan Science and Technology Planning Project (Grant No. 2020020601012260).
Author information
Authors and Affiliations
Contributions
MZ and XC designed the study; XC, JT, SH, HZ, and HC performed research; DH and JL analyzed data; DH, JL, and MZ contributed new methods; and XC and DH wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Ioly Kotta-Loizou.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
705_2021_5235_MOESM1_ESM.tif
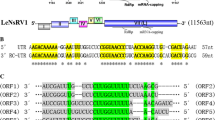
Supplementary file1 (TIF 2298 KB) Fig. S1 Schematic diagram of the genomic structure of LsaMV1. Gray boxes represent ORFs; single gray lines represent untranslated regions. The conserved sequences of the RdRP domains in the ORF are indicated
Rights and permissions
About this article
Cite this article
Chen, X., Hai, D., Li, J. et al. Complete genome sequence of a novel mitovirus associated with Lagenaria siceraria. Arch Virol 166, 3427–3431 (2021). https://doi.org/10.1007/s00705-021-05235-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05235-3