Abstract
Main conclusion
Latexes in immature fruit, young petioles and lignified trunks of fig trees protect the plant using toxic proteins and metabolites in various organ-dependent ways.
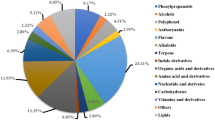
Latexes from plants contain high amounts of toxic proteins and metabolites, which attack microbes and herbivores after exudation at pest-induced wound sites. The protein and metabolite constituents of latexes are highly variable, depending on the plant species and organ. To determine the diversity of latex-based defense strategies in fig tree (Ficus carica) organs, we conducted comparative proteomic, transcriptomic and metabolomic analyses on latexes isolated from immature fruit, young petioles and lignified trunks of F. carica after constructing a unigene sequence library using RNA-seq data. Trypsin inhibitors were the most abundant proteins in petiole latex, while cysteine proteases (“ficins”) were the most abundant in immature fruit and trunk latexes. Galloylglycerol, a possible defense-related metabolite, appeared to be highly accumulated in all three latexes. The expression levels of pathogenesis-related proteins were highest in the latex of trunk, suggesting that this latex had adapted a defensive role against microbe attacks. Although young petioles and immature fruit are both unlignified soft organs, and potential food for herbivorous insects, unigenes for the sesquiterpenoid pathway, which likely produces defense-associated volatiles, and the phenylpropanoid pathway, which produces toxic furanocoumarins, were expressed less in immature fruit latex. This difference may indicate that while petioles and fruit protect the plant from attack by herbivores, the fruit must also attract insect pollinators at younger stages and animals after ripening. We also suggest possible candidate transcription factors and signal transduction proteins that are involved in the differential expression of the unigenes.








Similar content being viewed by others
Abbreviations
- DEG:
-
Differentially expressed (uni)gene
- GO:
-
Gene ontology
- PR:
-
Pathogenesis-related
- RPKM:
-
Read counts per kilobase of unigene per million mapped reads
References
Akimoto N, Ara T, Nakajima D, Suda K, Ikeda C, Takahashi S, Muneto R, Yamada M, Suzuki H, Shibata D, Sakurai N (2017) FlavonoidSearch: a system for comprehensive flavonoid annotation by mass spectrometry. Sci Rep UK 7:1243. https://doi.org/10.1038/s41598-017-01390-3
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402. https://doi.org/10.1093/nar/25.17.3389
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B Methodol 57:289–300
Birkett MA, Al Abassi S, Kröber T, Chamberlain K, Hooper AM, Guerin PM, Pettersson J, Pickett JA, Slade R, Wadhams LJ (2008) Antiectoparasitic activity of the gum resin, gum haggar, from the East African plant, Commiphora holtziana. Phytochemistry 69(8):1710–1715. https://doi.org/10.1016/j.phytochem.2008.02.017
Bruce TJ, Birkett MA, Blande J, Hooper AM, Martin JL, Khambay B, Prosser I, Smart LE, Wadhams LJ (2005) Response of economically important aphids to components of Hemizygia petiolata essential oil. Pest Manag Sci 61(11):1115–1121. https://doi.org/10.1002/ps.1102
Chanwitheesuk A, Teerawutgulrag A, Kilburn JD, Rakariyatham N (2007) Antimicrobial gallic acid from Caesalpinia mimosoides Lamk. Food Chem 100(3):1044–1048. https://doi.org/10.1016/j.foodchem.2005.11.008
Farag MA, Abdelfattah MS, Badr SE, Wessjohann LA (2014) Profiling the chemical content of Ficus lyrata extracts via UPLC–PDA–qTOF–MS and chemometrics. Nat Prod Res 28(19):1549–1556. https://doi.org/10.1080/14786419.2014.926353
Federici L, Di Matteo A, Fernandez-Recio J, Tsernoglou D, Cervone F (2006) Polygalacturonase inhibiting proteins: players in plant innate immunity? Trends Plant Sci 11(2):65–70. https://doi.org/10.1016/j.tplants.2005.12.005
Field D, Tiwari B, Booth T, Houten S, Swan D, Bertrand N, Thurston M (2006) Open software for biologists: from famine to feast. Nat Biotechnol 24(7):801–803. https://doi.org/10.1038/nbt0706-801
Fisher RA (1922) On the interpretation of χ2 from contingency tables, and the calculation of P. J R Stat Soc 85(1):87–94. https://doi.org/10.2307/2340521
Friedman M, Henika PR, Mandrell RE (2003) Antibacterial activities of phenolic benzaldehydes and benzoic acids against Campylobacter jejuni, Escherichia coli, Listeria monocytogenes, and Salmonella enterica. J Food Prot 66(10):1811–1821. https://doi.org/10.4315/0362-028X-66.10.1811
Gañan M, Martínez-Rodríguez AJ, Carrascosa AV (2009) Antimicrobial activity of phenolic compounds of wine against Campylobacter jejuni. Food Control 20(8):739–742. https://doi.org/10.1016/j.foodcont.2008.09.012
Gibernau M, Buser HR, Frey JE, Hossaert-McKey M (1997) Volatile compounds from extracts of figs of Ficus carica. Phytochemistry 46(2):241–244. https://doi.org/10.1016/S0031-9422(97)00292-6
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29(7):644–652. https://doi.org/10.1038/nbt.1883
Hagel JM, Yeung EC, Facchini PJ (2008) Got milk? The secret life of laticifers. Trends Plant Sci 13(12):631–639. https://doi.org/10.1016/j.tplants.2008.09.005
Hilder VA, Gatehouse AM, Sheerman SE, Barker RF, Boulter D (1987) A novel mechanism of insect resistance engineered into tobacco. Nature 330(6144):160–163. https://doi.org/10.1038/330160a0
Huang Y, Niu B, Gao Y, Fu L, Li W (2010) CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics 26(5):680–682. https://doi.org/10.1093/bioinformatics/btq003
Huynh QK, Borgmeyer JR, Zobel JF (1992) Isolation and characterization of a 22 kDa protein with antifungal properties from maize seeds. Biochem Biophys Res Commun 182(1):1–5. https://doi.org/10.1016/S0006-291X(05)80103-2
Jimenez CR, Huang L, Qiu Y, Burlingame AL (2003) In-gel digestion of proteins for MALDI-MS fingerprint mapping. In: Coligan JE, Dunn BM, Ploegh HL, Speicher DW, Wingfield PT (eds) Current protocols in protein science. Wiley, Hoboken, pp 16.4.1–16.4.5
Karamat F, Olry A, Munakata R, Koeduka T, Sugiyama A, Paris C, Hehn A, Bourgaud F, Yazaki K (2014) A coumarin-specific prenyltransferase catalyzes the crucial biosynthetic reaction for furanocoumarin formation in parsley. Plant J 77(4):627–638. https://doi.org/10.1111/tpj.12409
Karnchanatat A, Tiengburanatam N, Boonmee A, Puthong S, Sangvanich P (2011) Zingipain, a cysteine protease from Zingiber ottensii Valeton rhizomes with antiproliferative activities against fungi and human malignant cell lines. Prep Biochem Biotechnol 41(2):138–153. https://doi.org/10.1080/10826068.2011.547347
Kiran SR, Devi PS (2007) Evaluation of mosquitocidal activity of essential oil and sesquiterpenes from leaves of Chloroxylon swietenia DC. Parasitol Res 101(2):413–418. https://doi.org/10.1007/s00436-007-0485-z
Kitajima S, Kamei K, Taketani S, Yamaguchi M, Kawai F, Komatsu A, Inukai Y (2010) Two chitinase-like proteins abundantly accumulated in latex of mulberry show insecticidal activity. BMC Biochem 11(1):6. https://doi.org/10.1186/1471-2091-11-6
Kitajima S, Taira T, Oda K, Yamato KT, Inukai Y, Hori Y (2012) Comparative study of gene expression and major proteins’ function of laticifers in lignified and unlignified organs of mulberry. Planta 235(3):589–601. https://doi.org/10.1007/s00425-011-1533-6
Kitajima S, Yamamoto Y, Hirooka K, Taki C, Hibino S (2013) Laticifers in mulberry exclusively accumulate defense proteins related to biotic stresses. Plant Biotechnol 30(4):399–402. https://doi.org/10.5511/plantbiotechnology.13.0326a
Kitajima S, Miura K, Aoki W, Yamato KT, Taira T, Murakami R, Aburaya S (2016) Transcriptome and proteome analyses provide insight into laticifer’s defense of Euphorbia tirucalli against pests. Plant Physiol Biochem 108:434–446. https://doi.org/10.1016/j.plaphy.2016.08.008
Konno K (2011) Plant latex and other exudates as plant defense systems: roles of various defense chemicals and proteins contained therein. Phytochemistry 72(13):1510–1530. https://doi.org/10.1016/j.phytochem.2011.02.016
Konno K, Hirayama C, Nakamura M, Tateishi K, Tamura Y, Hattori M, Kohno K (2004) Papain protects papaya trees from herbivorous insects: role of cysteine proteases in latex. Plant J 37(3):370–378. https://doi.org/10.1046/j.1365-313X.2003.01968.x
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359. https://doi.org/10.1038/nmeth.1923
Lazreg-Aref H, Mars M, Fekih A, Aouni M, Said K (2012) Chemical composition and antibacterial activity of a hexane extract of Tunisian caprifig latex from the unripe fruit of Ficus carica. Pharm Biol 50(4):407–412. https://doi.org/10.3109/13880209.2011.608192
Lewinsohn TM (1991) The geographical distribution of plant latex. Chemoecology 2(1):64–68. https://doi.org/10.1007/BF01240668
Liu Y, Ahn JE, Datta S, Salzman RA, Moon J, Huyghues-Despointes B, Pittendrigh B, Murdock LL, Koiwa H, Zhu-Salzman K (2005) Arabidopsis vegetative storage protein is an anti-insect acid phosphatase. Plant Physiol 139(3):1545–1556. https://doi.org/10.1104/pp.105.066837
López-García B, Hernández M, Segundo BS (2012) Bromelain, a cysteine protease from pineapple (Ananas comosus) stem, is an inhibitor of fungal plant pathogens. Lett Appl Microbiol 55(1):62–67. https://doi.org/10.1111/j.1472-765X.2012.03258.x
Matsui K, Bae J, Esaka K, Morisaka H, Kuroda K, Ueda M (2013) Exoproteome profiles of Clostridium cellulovorans grown on various carbon sources. Appl Environ Microbiol 79(21):6576–6584. https://doi.org/10.1128/AEM.02137-13
Mawa S, Husain K, Jantan I (2013) Ficus carica L. (Moraceae): phytochemistry, traditional uses and biological activities. Evid Based Complement Altern Med 2013:974256. https://doi.org/10.1155/2013/974256
Munakata R, Olry A, Karamat F, Courdavault V, Sugiyama A, Krieger C, Silie P, Foureau E, Papon N, Grosjean J, Yazaki K (2016) Molecular evolution of parsnip (Pastinaca sativa) membrane-bound prenyltransferases for linear and/or angular furanocoumarin biosynthesis. New Phytol 211(1):332–344. https://doi.org/10.1111/nph.13899
Nohynek LJ, Alakomi HL, Kähkönen MP, Heinonen M, Helander IM, Oksman-Caldentey KM, Puupponen-Pimiä RH (2006) Berry phenolics: antimicrobial properties and mechanisms of action against severe human pathogens. Nutr Cancer 54(1):18–32. https://doi.org/10.1016/j.foodchem.2010.04.064
Oliveira AP, Silva LR, de Pinho PG, Gil-Izquierdo A, Valentão P, Silva BM, Pereira JA, Andrade PB (2010) Volatile profiling of Ficus carica varieties by HS-SPME and GC–IT–MS. Food Chem 123(2):548–557. https://doi.org/10.1016/j.foodchem.2010.04.064
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http://www.R-project.org/
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acid Res 43(7):e47–e47. https://doi.org/10.1093/nar/gkv007
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Sakurai N, Ara T, Enomoto M, Motegi T, Morishita Y, Kurabayashi A, Iijima Y, Ogata Y, Nakajima D, Suzuki H, Shibata D (2014) Tools and databases of the KOMICS web portal for preprocessing, mining, and dissemination of metabolomics data. Biomed Res Int 2014:194812. https://doi.org/10.1155/2014/194812
Shin R, Kim MJ, Paek KH (2003) The CaTin1 (Capsicum annuum TMV-induced clone 1) and CaTin1-2 genes are linked head-to-head and share a bidirectional promoter. Plant Cell Physiol 44(5):549–554. https://doi.org/10.1093/pcp/pcg069
Takahashi T, Okiura A, Saito K, Kohno M (2014) Identification of phenylpropanoids in fig (Ficus carica L.) leaves. J Agric Food Chem 62(41):10076–10083. https://doi.org/10.1021/jf5025938
Takahashi T, Okiura A, Kohno M (2017) Phenylpropanoid composition in fig (Ficus carica L.) leaves. J Nat Med 71(4):770–775. https://doi.org/10.1007/s11418-017-1093-6
Terras FR, Schoofs HM, Thevissen K, Osborn RW, Vanderleyden J, Cammue BP, Broekaert WF (1993) Synergistic enhancement of the antifungal activity of wheat and barley thionins by radish and oilseed rape 2S albumins and by barley trypsin inhibitors. Plant Physiol 103(4):1311–1319. https://doi.org/10.1104/pp.103.4.1311
Van Loon LC (1999) Occurrence and properties of plant pathogenesis-related proteins. In: Datta SK, Muthukrishnan S (eds) Pathogenesis-related proteins in plants. CRC Press LLC, Boca Raton, pp 1–20
Veberic R, Colaric M, Stampar F (2008) Phenolic acids and flavonoids of fig fruit (Ficus carica L.) in the northern Mediterranean region. Food Chem 106(1):153–157. https://doi.org/10.1016/j.foodchem.2007.05.061
Wang X, Shi M, Lu X, Ma R, Wu C, Guo A, Peng M, Tian W (2010) A method for protein extraction from different subcellular fractions of laticifer latex in Hevea brasiliensis compatible with 2-DE and MS. Proteome Sci 8(1):35. https://doi.org/10.1186/1477-5956-8-35
Wasano N, Konno K, Nakamura M, Hirayama C, Hattori M, Tateishi K (2009) A unique latex protein, MLX56, defends mulberry trees from insects. Phytochemistry 70(7):880–888. https://doi.org/10.1016/j.phytochem.2009.04.014
Acknowledgements
This study was supported by Grant-in-Aid for Scientific Research from the Ministry of Education, Science, Sports, Culture and Technology of Japan (to SK, No. 16K07641). The LC- Orbitrap analysis of proteome was technically supported by the Kyoto Integrated Science and Technology Bio-Analysis Center. We wish to thank Dr. Jun Wada for his generous aid in the proteome analysis.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kitajima, S., Aoki, W., Shibata, D. et al. Comparative multi-omics analysis reveals diverse latex-based defense strategies against pests among latex-producing organs of the fig tree (Ficus carica). Planta 247, 1423–1438 (2018). https://doi.org/10.1007/s00425-018-2880-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-018-2880-3