Abstract
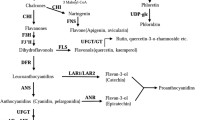
Chinese bayberry (Myrica rubra) is a fruit crop with cultivars producing fruit ranging from white (Shuijing, SJ) to red (Dongkui, DK) and dark red-purple (Biqi, BQ), as a result of different levels of anthocyanin accumulation. Genes encoding the anthocyanin biosynthesis enzymes chalcone synthase, chalcone isomerase, flavanone 3-hydroxylase (F3H), flavonoid 3′-hydroxylase (F3′H), dihydroflavonol 4-reductase (DFR), anthocyanidin synthase (ANS) and UDP-glucose:flavonoid 3-O-glucosyltransferase (UFGT), as well as MrMYB1, a R2R3 MYB transcription factor homologous to known activators of anthocyanin biosynthesis, were isolated from ripe fruit of BQ. Differences in mRNA abundance of MrF3H, MrF3′H, MrDFR1, MrANS and MrUFGT were highly correlated with differential accumulation of anthocyanins between cultivars, suggesting coordinated regulation by transcription factors. The transcript level of MrMYB1 was strongly associated with the anthocyanin content in ripe fruit of the three cultivars, as well as different anthocyanin containing tissues of BQ fruit. Fruit bagging strongly inhibited anthocyanin accumulation in fruit as well as the expression of all anthocyanin biosynthetic genes and MrMYB1. Over-expression of MrMYB1 stimulated both anthocyanin accumulation and activated an Arabidopsis-DFR promoter in tobacco (Nicotiana tabacum). MrMYB1d, an allele with a 1 bp deletion at nucleotide 30 of coding sequence, was observed in SJ and DK fruit, suggesting that a nonsense mutation of the MYB1 protein may be responsible for no or low expression of MYB1 in the white and red fruit. These results show that coordinated expression of multiple biosynthetic genes is involved in anthocyanin accumulation in Chinese bayberry fruit, and this is regulated by MrMYB1.







Similar content being viewed by others
Abbreviations
- ANS:
-
Anthocyanidin synthase
- bHLH:
-
Basic-helix-loop-helix
- BQ:
-
Biqi
- CDS:
-
Coding sequence
- CHI:
-
Chalcone isomerase
- CHS:
-
Chalcone synthase
- DAFB:
-
Days after full bloom
- DFR:
-
Dihydroflavonol 4-reductase
- DK:
-
Dongkui
- EBGs:
-
Early biosynthetic genes
- F3H:
-
Flavanone 3-hydroxylase
- F3′H:
-
Flavonoid 3′-hydroxylase
- LBGs:
-
Late biosynthetic genes
- LUC:
-
Firefly luciferase
- Q-PCR:
-
Real-time quantitative PCR
- REN:
-
Renilla luciferase
- SJ:
-
Shuijing
- TFs:
-
Transcription factors
- UFGT:
-
UDP-glucose:flavonoid 3-O-glucosyltransferase
References
Aharoni A, De Vos CHR, Wein M, Sun ZK, Greco R, Kroon A, Mol JNM, O’Connell AP (2001) The strawberry FaMYB1 transcription factor suppresses anthocyanin and flavonol accumulation in transgenic tobacco. Plant J 28:319–332
Ahmed N, Maekawa M, Noda K (2009) Anthocyanin accumulation and expression pattern of anthocyanin biosynthesis genes in developing wheat coleoptiles. Biol Plant 53:223–228
Albert NW, Lewis DH, Zhang H-B, Irving LJ, Jameson PE, Davies KM (2009) Light-induced vegetative anthocyanin pigmentation in Petunia. J Exp Bot 60:2191–2202
Allan AC, Hellens RP, Laing WA (2008) MYB transcription factors that colour our fruit. Trends Plant Sci 13:99–102
Ban Y, Honda C, Hatsuyama Y, Igarashi M, Bessho H, Moriguchi T (2007) Isolation and functional analysis of a MYB transcription factor gene that is a key regulator for the development of red coloration in apple skin. Plant Cell Physiol 48:958–970
Borevitz JO, Xia Y, Blount J, Dixon RA, Lamb C (2000) Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12:2383–2393
Boss PK, Davies C, Robinson SP (1996) Analysis of the expression of anthocyanin pathway genes in developing Vitis vinifera L cv Shiraz grape berries and the implications for pathway regulation. Plant Physiol 111:1059–1066
Broun P (2005) Transcriptional control of flavonoid biosynthesis: a complex network of conserved regulators involved in multiple aspects of differentiation in Arabidopsis. Curr Opin Plant Biol 8:272–279
Chagne D, Carlisle CM, Blond C, Volz RK, Whitworth CJ, Oraguzie NC, Crowhurst RN, Allan AC, Espley RV, Hellens RP, Gardiner SE (2007) Mapping a candidate gene (MdMYB10) for red flesh and foliage colour in apple. BMC Genomics 8:212
Chen KS, Xu CJ, Zhang B, Ferguson IB (2004) Red bayberry: botany and horticulture. Hortic Rev 30:83–114
Cominelli E, Gusmaroli G, Allegra D, Galbiati M, Wade HK, Jenkins GI, Tonelli C (2008) Expression analyssis of anthocyanin regulatory genes in response to different light qualities in Arabidopsis thaliana. Plant Physiol 165:886–894
Davies KM, Schwinn KE (2003) Transcriptional regulation of secondary metabolism. Funct Plant Biol 30:913–925
Dubos C, Le Gourrierec J, Baudry A, Huep G, Lanet E, Debeaujon I, Routaboul JM, Alboresi A, Weisshaar B, Lepiniec L (2008) MYBL2 is a new regulator of flavonoid biosynthesis in Arabidopsis thaliana. Plant J 55:940–953
Espley RV, Hellens RP, Putterill J, Stevenson DE, Kutty-Amma S, Allan AC (2007) Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 49:414–427
Espley RV, Brendolise C, Chagné D, Kutty-Amma S, Green S, Volz R, Putterill J, Schouten HJ, Gardiner SE, Hellens RP, Allan AC (2009) Multiple repeats of a promoter segment causes transcription factor autoregulation in red apples. Plant Cell 21:168–183
Fang ZX, Zhang YH, Lü Y, Ma GP, Chen JC, Liu DH, Ye XQ (2009) Phenolic compounds and antioxidant capacities of bayberry juices. Food Chem 113:884–888
Galego L, Almeida J (2007) Molecular genetic basis of flower colour variegation in Linaria. Genet Res 89:129–134
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53:814–827
Grotewold E (2006) The genetics and biochemistry of floral pigments. Annu Rev Plant Biol 57:761–780
Hellens R, Allan A, Friel E, Bolitho K, Grafton K, Templeton M, Karunairetnam S, Gleave A, Laing W (2005) Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1:13
Huang C-H, Yu B, Teng Y-W, Su J, Shu Q, Cheng Z-Q, Zeng L-Q (2009) Effects of fruit bagging on coloring and related physiology, and qualities of red Chinese sand pears during fruit maturation. Sci Hortic 121:149–158
Jia H-J, Araki A, Okamoto G (2004) Influence of fruit bagging on aroma volatiles and skin coloration of ‘Hakuho’ peach (Prunus persica Batsch). Postharvest Biol Technol 35:61–68
Ju Z-G, Liu C-L, Yuan Y-B (1995) Activities of chalcone synthase and UDPGal: flavonoid-3-o-glycosyltransferase in relation to anthocyanin synthesis in apple. Sci Hortic 63:175–185
Kim S, Binzel ML, Park S, Yoo K-S, Pike LM (2004) Inactivation of DFR (dihydroflavonol 4-reductase) gene transcription results in blockage of anthocyanin production in yellow onions (Allium cepa). Mol Breed 14:253–263
Kobayashi S, Ishimaru M, Hiraoka K, Honda C (2002) Myb-related genes of the Kyoho grape (Vitis labruscana) regulate anthocyanin biosynthesis. Planta 215:924–933
Kobayashi S, Goto-Yamamoto N, Hirochika H (2004) Retrotransposon-induced mutations in grape skin color. Science 304:982
Koes R, Verweij W, Quattrocchio F (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10:236–242
Kumar S, Tamura K, Nei M (2004) MEGA3:integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Lo Piero AR, Consoli A, Puglisi I, Orestano G, Recupero GR, Petrone G (2005) Anthocyaninless cultivars of sweet orange lack to express the UDP-glucose flavonoid 3-O-glucosyl transferase. J Plant Biochem Biotechnol 14:9–14
Matsui K, Umemura Y, Ohme-Takagi M (2008) AtMYBL2, a protein with a single MYB domain, acts as a negative regulator of anthocyanin biosynthesis in Arabidopsis. Plant J 55:954–967
Matus JT, Aquea F, Arce-Johnson P (2008) Analysis of the grape MYB R2R3 subfamily reveals expanded wine quality-related clades and conserved gene structure organization across Vitis and Arabidopsis genomes. BMC Plant Biol 8:83
Matus JT, Loyola R, Vega A, Peña-Neira A, Bordeu E, Arce-Johnson P, Alcalde JA (2009) Post-veraison sunlight exposure induces MYB-mediated transcriptional regulation of anthocyanin and flavonol synthesis in berry skins of Vitis vinifera. J Exp Bot 60:853–867
Nakatsuka T, Haruta KS, Pitaksutheepong C, Abe Y, Kakizaki Y, Yamamoto K, Shimada N, Yamamura S, Nishihara M (2008) Identification and characterization of R2R3-MYB and bHLH transcription factors regulating anthocyanin biosynthesis in gentian flowers. Plant Cell Physiol 49:1818–1829
Palapol Y, Ketsa S, Lin-Wang K, Ferguson IB, Allan AC (2009) A MYB transcription factor regulates anthocyanin biosynthesis in mangosteen (Garcinia mangostana L.) fruit during ripening. Planta 229:1323–1334
Park J-S, Kim J-B, Cho K-J, Cheon C-I, Sung M-K, Choung M-G, Roh K-H (2008) Arabidopsis R2R3-MYB transcription factor AtMYB60 functions as a transcriptional repressor of anthocyanin biosynthesis in lettuce (Lactuca sativa). Plant Cell Rep 27:985–994
Paz-Ares J, Ghosal D, Wienand U, Peterson PA, Saedler H (1987) The regulatory c1 locus of Zea mays encodes a protein with homology to myb proto-oncogene products and with structural similarities to transcriptional activators. EMBO J 6:3553–3558
Romero I, Sanchez-Ballesta MT, Maldonado R, Escribano MI, Merodio C (2008) Anthocyanin, antioxidant activity and stress-induced gene expression in high CO2-treated table grapes stored at low temperature. Plant Physiol 165:522–530
Rowen DD, Cao MS, Lin-Wang K, Cooney JM, Jensen DJ, Austin PT, Hunt MB, Norling C, Hellens RP, Schaffer RJ, Allan AC (2009) Environmental regulation of leaf colour in red 35S:PAP1 Arabidopsis thaliana. New Phytol 182:102–115
Saitoh K, Onishi K, Mikami I, Thidar K, Sano Y (2004) Allelic diversification at the C (OsC1) locus of wild and cultivated rice: nucleotide changes associated with phenotypes. Genetics 168:997–1007
Schwinn K, Venail J, Shang Y, Mackay S, Alm V, Butelli E, Oyama R, Bailey P, Davies K, Martin C (2006) A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum. Plant Cell 18:831–851
Shan LL, Li X, Wang P, Cai C, Zhang B, Sun CD, Zhang WS, Xu CJ, Ferguson IB, Chen KS (2008) Characterization of cDNAs associated with lignification and their expression profiles in loquat fruit with different lignin accumulation. Planta 227:1243–1254
Spelt C, Quattrocchio F, Mol JN, Koes R (2000) anthocyanin1 of petunia encodes a basic helix-loop-helix protein that directly activates transcription of structural anthocyanin genes. Plant Cell 12:1619–1632
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4:447–456
Streisfeld MA, Rausher MD (2009) Genetic changes contributing to the parallel evolution of red floral pigmentation among Ipomoea species. New Phytol 183:751–763
Takos AM, Jaffé FW, Jacob SR, Bogs J, Robinson SP, Walker AR (2006) Light-induced expression of a MYB gene regulates anthocyanin biosynthesis in red apples. Plant Physiol 142:1216–1232
Ubi BE, Honda C, Bessho H, Kondo S, Wada M, Kobayashi S, Moriguchi T (2006) Expression analysis of anthocyanin biosynthetic genes in apple skin: effect of UV-B and temperature. Plant Sci 170:571–578
Walker AR, Lee E, Bogs J, McDavid DAJ, Thomas MR, Robinson SP (2007) White grapes arose through the mutation of two similar and adjacent regulatory genes. Plant J 49:772–785
Wrolstand RE, Culbertson JD, Cornwell CJ, Mattick L (1982) Detection of adulteration in blackberry juice concentrates and wines. J Assoc Off Anal Chem 6:1417–1423
Yin XR, Chen KS, Allan AC, Wu RM, Zhang B, Lallu N, Ferguson IB (2008) Ethylene-induced modulation of genes associated with the ethylene signalling pathway in ripening kiwifruit. J Exp Bot 59:2097–2108
Yuan Y, Chiu L-W, Li L (2009) Transcriptional regulation of anthocyanin biosynthesis in red cabbage. Planta 230:1141–1153
Zhang WS, Li X, Zheng JT, Wang GY, Sun CD, Ferguson IB, Chen KS (2008) Bioactive components and antioxidant capacity of Chinese bayberry (Myrica rubra Sieb. and Zucc.) fruit in relation to fruit maturity and postharvest storage. Eur Food Res Technol 227:1091–1097
Zhang SM, Gao ZS, Xu CJ, Chen KS, Wang GY, Zheng JT, Lu T (2009) Genetic diversity of chinese bayberry (Myrica rubra Sieb. et Zucc.) accessions revealed by amplified fragment length polymorphism. HortScience 44:487–491
Zimmermann IM, Heim MA, Weisshaar B, Uhrig JF (2004) Comprehensive identification of Arabidopsis thaliana MYB transcription factors interacting with R/B-like BHLH proteins. Plant J 40:22–34
Acknowledgments
This research was supported by the Special Scientific Research Fund of Agricultural Public Welfare Profession of China (200903044), the National Research and Development Project of Transgenic Crops of China (2009ZX08009-122B), the 948 project, the Science and Technology Project of Zhejiang Province (2006C14016, 2009C14023), the Specialized Research Fund for the Doctoral Program of Higher Education of China (20090101110099), the Science Foundation of Chinese University, and the 111 project (No. B06014). It is part of a collaborative programme between Zhejiang University, China, and the New Zealand Institute for Plant and Food Research, NZ.
Author information
Authors and Affiliations
Corresponding author
Additional information
S.-S. Niu and C.-J. Xu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Niu, SS., Xu, CJ., Zhang, WS. et al. Coordinated regulation of anthocyanin biosynthesis in Chinese bayberry (Myrica rubra) fruit by a R2R3 MYB transcription factor. Planta 231, 887–899 (2010). https://doi.org/10.1007/s00425-009-1095-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-009-1095-z