Abstract
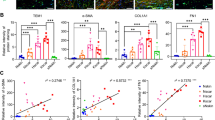
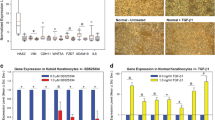
Among raised dermal scar types, keloid (KS) and hypertrophic scars (HS) are considered to present clinical similarities, but there are no known specific biomarkers that allow both scar types to be easily distinguished. Development and progression of raised dermal scars comprises the activation of several molecular pathways and cell defence mechanisms leading to elevated extracellular matrix component synthesis, delayed apoptosis, altered migration and differentiation. Therefore, the aim here was to identify biomarkers that may differentiate between KS and HS compared to normal skin (NS). To achieve this aim, NS (n = 14), KS (n = 14) and HS (n = 14) biopsies were evaluated using histology by H&E staining. Tissue biopsies and primary fibroblasts (passages 0–4) were employed to assess the gene expression levels of 21 biomarkers selected from our previous microarray studies using qRT-PCR. Finally, protein expression was evaluated using In-Cell Western Blotting in primary fibroblasts (p 0–4). Our results demonstrated that out of the 21 biomarkers screened at mRNA and protein levels, α2β1-integrin, Hsp27, PAI-2, MMP-19 and CGRP showed significantly higher expression (p < 0.05) in KS compared to NS and HS. Additionally, these five key biomarkers were found to be significantly higher (p < 0.05) at mRNA level in KS taken from the sternum, a region known to be subjected to high mechanical forces in the body during the performance of daily movements. In conclusion, our findings offer potential molecular targets in raised dermal scars differentiation. Future targeted research may allow provision of diagnostic and prognostic markers in keloid versus hypertrophic scars.















Similar content being viewed by others
References
Akaishi S, Akimoto M, Ogawa R, Hyakusoku H (2008) The relationship between keloid growth pattern and stretching tension: visual analysis using the finite element method. Ann Plast Surg 604:445–451
Akaishi S, Ogawa R, Hyakusoku H (2008) Keloid and hypertrophic scar: neurogenic inflammation hypotheses. Med Hypotheses 711:32–38
Altun V, Hakvoort TE, Van Zuijlen PPM, Van Der Kwast TH, Prens EP (2001) Nerve outgrowth and neuropeptide expression during the remodeling of human burn wound scars: a 7-month follow-up study of 22 patients. Burns 277:717–722
Atiyeh BS, Costagliola M, Hayek SN (2005) Keloid or hypertrophic scar: the controversy: review of the literature. Ann Plast Surg 546:676–680
Bayat A, Arscott G, Ollier WE, McGrouther DA, Ferguson MW (2005) Keloid disease: clinical relevance of single versus multiple site scars. Br J Plast Surg 581:28–37
Bayat A, Arscott G, Ollier WER, Ferguson MWJ, Mc Grouther DA (2004) Description of site-specific morphology of keloid phenotypes in an Afrocaribbean population. Br J Plast Surg 572:122–133
Bayat A, Bock O, Mrowietz U, Ollier WE, Ferguson MW (2003) Genetic susceptibility to keloid disease and hypertrophic scarring: transforming growth factor beta1 common polymorphisms and plasma levels. Plast Reconstr Surg 1112:535–543
Bayat A, McGrouther DA, Ferguson MWJ (2003) Skin scarring. Br Med J 3267380:88–92
Berry DP, Harding KG, Stanton MR, Jasani B, Ehrlich HP (1998) Human wound contraction: collagen organization, fibroblasts, and myofibroblasts. Plast Reconstr Surg 1021:124–131
Bignotti E, Tassi RA, Calza S, Ravaggi A, Romani C, Rossi E, Falchetti M, Odicino FE, Pecorelli S, Santin AD (2006) Differential gene expression profiles between tumor biopsies and short-term primary cultures of ovarian serous carcinomas: identification of novel molecular biomarkers for early diagnosis and therapy. Gynecol Oncol 1032:405–416
Bitar KN, Kaminski MS, Hailat N, Cease KB, Strahler JR (1991) Hsp27 is a mediator of sustained smooth muscle contraction in response to bombesin. Biochem Biophys Res Commun 1813:1192–1200
Blake MJ, Gershon D, Fargnoli J, Holbrook NJ (1990) Discordant expression of heat shock protein mRNAs in tissues of heat-stressed rats. J Biol Chem 26525:15275–15279
Brissett AE, Sherris DA (2001) Scar contractures, hypertrophic scars, and keloids. Facial Plast Surg 174:263–271
Brown JJ, Bayat A (2009) Genetic susceptibility to raised dermal scarring. Br J Dermatol 1611:8–18
Butler PD, Longaker MT, Yang GP (2008) Current progress in keloid research and treatment. J Am Coll Surg 2064:731–741
Charette SJ, Lavoie JN, Lambert H, Landry J (2000) Inhibition of Daxx-mediated apoptosis by heat shock protein 27. Mol Cell Biol 2020:7602–7612
Chen G, Gharib TG, Huang CC, Taylor JM, Misek DE, Kardia SL, Giordano TJ, Iannettoni MD, Orringer MB, Hanash SM, Beer DG (2002) Discordant protein and mRNA expression in lung adenocarcinomas. Mol Cell Proteomics 14:304–313
Chen J-J, Jin P-S, Zhao S, Cen Y, Liu Y, Xu X-W, Duan W-Q, Wang H-S (2011) Effect of heat shock protein 47 on collagen synthesis of keloid in vivo. ANZ J Surg 816:425–430
Chen J, Wang JH, Zhuang HX (2006) Influence of substance P on the proliferation and apoptosis of fibroblasts of pathological scars. Chin J Burns 224:277–280
Chen W, Fu X, Sun X, Sun T, Zhao Z, Sheng Z (2003) Analysis of differentially expressed genes in keloids and normal skin with cDNA microarray. J Surg Res 1132:208–216
Chike-Obi CJ, Cole PD, Brissett AE (2009) Keloids: pathogenesis, clinical features, and management. Semin Plast Surg 233:178–184
Croucher DR, Saunders DN, Lobov S, Ranson M (2008) Revisiting the biological roles of PAI2 (SERPINB2) in cancer. Nat Rev Cancer 87:535–545
Crowe R, Parkhouse N, McGrouther D, Burnstock G (1994) Neuropeptide-containing nerves in painful hypertrophic human scar tissue. Br J Dermatol 1304:444–452
Dangles V, Lazar V, Validire P, Richon S, Wertheimer M, Laville V, Janneau JL, Barrois M, Bovin C, Poynard T, Vallancien G, Bellet D (2002) Gene expression profiles of bladder cancers: evidence for a striking effect of in vitro cell models on gene patterns. Br J Cancer 868:1283–1289
Dasu MR, Hawkins HK, Barrow RE, Xue H, Herndon DN (2004) Gene expression profiles from hypertrophic scar fibroblasts before and after IL-6 stimulation. J Pathol 2024:476–485
Davidson S, Aziz N, Rashid RM, Khachemoune A (2009) A primary care perspective on keloids. Med Gen Med Medscape Gen Med 11:18
Dipietro LA, Reintjes MG, Low QEH, Levi B, Gamelli RL (2001) Modulation of macrophage recruitment into wounds by monocyte chemoattractant protein-1. Wound Repair Regen 91:28–33
Gauglitz GG, Korting HC, Pavicic T, Ruzicka T, Jeschke MG (2011) Hypertrophic scarring and keloids: pathomechanisms and current and emerging treatment strategies. Mol Med 171–2:113–125
Gu LH, Coulombe PA (2007) Keratin function in skin epithelia: a broadening palette with surprising shades. Curr Opin Cell Biol 191:13–23
Hu ZF, Gao JH, Li W, Song YB, Li CL (2006) Differential gene expression profile of keloids: a study with cDNA microarray. Nan Fang Yi Ke Da Xue Xue Bao 263:308–312
Huang C, Akaishi S, Ogawa R (2012) Mechanosignaling pathways in cutaneous scarring. Arch Dermatol Res 3048:589–597
Huang C, Nie F, Qin Z, Li B, Zhao X (2013) A snapshot of gene expression signatures generated using microarray datasets associated with excessive scarring. Am J Dermatopathol 351:64–73
Huang H, Kamm RD, Lee RT (2004) Cell mechanics and mechanotransduction: pathways, probes, and physiology. Am J Physiol Cell Physiol 2871:C1–C11
Ingber D (1991) Integrins as mechanochemical transducers. Curr Opin Cell Biol 35:841–848
Kazeem AA (1988) The immunological aspects of keloid tumor formation. J Surg Oncol 381:16–18
Ketchum LD, Cohen IK, Masters FW (1974) Hypertrophic scars and keloids. A collective review. Plast Reconstr Surg 532:140–154
Kuwano K, Araya J, Hara H (2007) Molecular targets in idiopathic pulmonary fibrosis. Respir Circ 5810:979–985
Laplante AF, Moulin V, Auger FA, Landry J, Li H, Morrow G, Tanguay RM, Germain L (1998) Expression of heat shock proteins in mouse skin during wound healing. J Histochem Cytochem 4611:1291–1301
Lee JY, Yang CC, Chao SC, Wong TW (2004) Histopathological differential diagnosis of keloid and hypertrophic scar. Am J Dermatopathol 265:379–384
Low SQ, Moy RL (1992) Scar wars strategies: target collagen. J Dermatol Surg Oncol 1811:981–986
Luger TA, Lotti T (1998) Neuropeptides: role in inflammatory skin diseases. J Eur Acad Dermatol Venereol 103:207–211
Luo S, Benathan M, Raffoul W, Panizzon RG, Egloff DV (2001) Abnormal balance between proliferation and apoptotic cell death in fibroblasts derived from keloid lesions. Plast Reconstr Surg 1071:87–96
Marneros AG, Krieg T (2004) Keloids—clinical diagnosis, pathogenesis, and treatment options. J Dtsch Dermatol Ges 211:905–913
Morris SD (2002) Heat shock proteins and the skin. Clin Exp Dermatol 273:220–224
Naitoh M, Kubota H, Ikeda M, Tanaka T, Shirane H, Suzuki S, Nagata K (2005) Gene expression in human keloids is altered from dermal to chondrocytic and osteogenic lineage. Genes Cells 1011:1081–1091
Ogawa R (2008) Keloid and hypertrophic scarring may result from a mechanoreceptor or mechanosensitive nociceptor disorder. Med Hypotheses 714:493–500
Ogawa R (2011) Mechanobiology of scarring. Wound Repair Regen 19:s2–s9
Ogawa R, Akaishi S, Huang C, Dohi T, Aoki M, Omori Y, Koike S, Kobe K, Akimoto M, Hyakusoku H (2011) Clinical applications of basic research that shows reducing skin tension could prevent and treat abnormal scarring: the importance of fascial/subcutaneous tensile reduction sutures and flap surgery for keloid and hypertrophic scar reconstruction. J Nippon Med School 782:68–76
Ogawa R, Hsu C-K (2013) Mechanobiological dysregulation of the epidermis and dermis in skin disorders and in degeneration. J Cell Mol Med 177:817–822
Ogawa R, Okai K, Tokumura F, Mori K, Ohmori Y, Huang C, Hyakusoku H, Akaishi S (2012) The relationship between skin stretching/contraction and pathologic scarring: the important role of mechanical forces in keloid generation. Wound Repair Regen 202:149–157
Paddock HN, Schultz GS, Baker HV, Varela JC, Beierle EA, Moldawer LL, Mozingo DW (2003) Analysis of gene expression patterns in human postburn hypertrophic scars. J Burn Care Rehabil 246:371–377
Parcellier A, Gurbuxani S, Schmitt E, Solary E, Garrido C (2003) Heat shock proteins, cellular chaperones that modulate mitochondrial cell death pathways. Biochem Biophys Res Commun 3043:505–512
Rockwell WB (1997) Elevated levels of testosterone receptors in keloid tissue: an experimental investigation. Plast Reconstr Surg 1002:396
Satish L, Lyons-Weiler J, Hebda PA, Wells A (2006) Gene expression patterns in isolated keloid fibroblasts. Wound Repair Regen 144:463–470
Schierle HP, Scholz D, Lemperle G (1997) Elevated levels of testosterone receptors in keloid tissue: an experimental investigation. Plast Reconstr Surg 1002:390–395
Scholzen T, Armstrong CA, Bunnett NW, Luger TA, Olerud JC, Ansel JE (1998) Neuropeptides in the skin: interactions between the neuroendocrine and the skin immune systems. Exp Dermatol 72–73:81–96
Seifert O, Bayat A, Geffers R, Dienus K, Buer J, Löfgren S, Matussek A (2008) Identification of unique gene expression patterns within different lesional sites of keloids. Wound Repair Regen 162:254–265
Seifert O, Mrowietz U (2009) Keloid scarring: bench and bedside. Arch Dermatol Res 3014:259–272
Shih B, Bayat A (2010) Genetics of keloid scarring. Arch Dermatol Res 3025:319–339
Shih B, McGrouther DA, Bayat A (2010) Identification of novel keloid biomarkers through profiling of tissue biopsies versus cell cultures in keloid margin specimens compared to adjacent normal skin. Eplasty 7(10):e24
Slemp AE, Kirschner RE (2006) Keloids and scars: a review of keloids and scars, their pathogenesis, risk factors, and management. Curr Opin Pediatr 184:396–402
Smith JC, Boone BE, Opalenik SR, Williams SM, Russell SB (2007) Gene profiling of keloid fibroblasts shows altered expression in multiple fibrosis-associated pathways. J Invest Dermatol 1285:1298–1310
Suarez E, Syed F, Alonso-Rasgado T, Mandal P, Bayat A (2013) Up-regulation of tension-related proteins in keloids: knockdown of hsp27, α2β1-integrin, and pai-2 shows convincing reduction of extracellular matrix production. Plast Reconstr Surg 1312:158e–173e
Swamy SMK, Tan P, Zhu YZ, Lu J, Achuth HN, Moochhala S (2004) Role of phenytoin in wound healing: microarray analysis of early transcriptional responses in human dermal fibroblasts. Biochem Biophys Res Commun 3143:661–666
Syed F, Ahmadi E, Iqbal SA, Singh S, McGrouther DA, Bayat A (2011) Fibroblasts from the growing margin of keloid scars produce higher levels of collagen I and III compared with intralesional and extralesional sites: clinical implications for lesional site-directed therapy. Br J Dermatol 1641:83–96
Tomasek JJ, Gabbiani G, Hinz B, Chaponnier C, Brown RA (2002) Myofibroblasts and mechano: regulation of connective tissue remodelling. Nat Rev Mol Cell Biol 35:349–363
Tsou R, Cole JK, Nathens AB, Isik FF, Heimbach DM, Engrav LH, Gibran NS (2000) Analysis of hypertrophic and normal scar gene expression with cDNA microarrays. J Burn Care Rehabil 216:541–550
Verhaegen PD, van Zuijlen PP, Pennings NM, van Marle J, Niessen FB, van der Horst CM, Middelkoop E (2009) Differences in collagen architecture between keloid, hypertrophic scar, normotrophic scar, and normal skin: an objective histopathological analysis. Wound Repair Regen 175:649–656
Wong TW, Chiu HC, Chen JS, Lin LJ, Chang CC (1995) Symptomatic keloids in two children: dramatic improvement with silicone cream occlusive dressing. Arch Dermatol 1317:775–777
Wong VW, Akaishi S, Longaker MT, Gurtner GC (2011) Pushing back: wound mechanotransduction in repair and regeneration. J Invest Dermatol 13111:2186–2196
Wu J, Ma B, Yi S, Wang Z, He W, Luo G, Chen X, Wang X, Chen A, Barisoni D (2004) Gene expression of early hypertrophic scar tissue screened by means of cDNA microarrays. J Trauma 576:1276–1286
Yamaoka J, Di ZH, Sun W, Kawana S (2007) Erratum to “Changes in cutaneous sensory nerve fibers induced by skin-scratching in mice” [J. Dermatol. Sci. 46 (2007) 41–51]. J Dermatol Sci 472:172–182
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Suarez, E., Syed, F., Alonso-Rasgado, T. et al. Identification of biomarkers involved in differential profiling of hypertrophic and keloid scars versus normal skin. Arch Dermatol Res 307, 115–133 (2015). https://doi.org/10.1007/s00403-014-1512-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00403-014-1512-4