Abstract
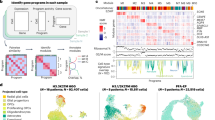
Histone H3 K27M mutation is the defining molecular feature of the devastating pediatric brain tumor, diffuse intrinsic pontine glioma (DIPG). The prevalence of histone H3 K27M mutations indicates a critical role in DIPGs, but the contribution of the mutation to disease pathogenesis remains unclear. We show that knockdown of this mutation in DIPG xenografts restores K27M-dependent loss of H3K27me3 and delays tumor growth. Comparisons of matched DIPG xenografts with and without K27M knockdown allowed identification of mutation-specific effects on the transcriptome and epigenome. The resulting transcriptional changes recapitulate expression signatures from K27M primary DIPG tumors and are strongly enriched for genes associated with nervous system development. Integrated analysis of ChIP-seq and expression data showed that genes upregulated by the mutation are overrepresented in apparently bivalent promoters. Many of these targets are associated with more immature differentiation states. Expression profiles indicate K27M knockdown decreases proliferation and increases differentiation within lineages represented in DIPG. These data suggest that K27M-mediated loss of H3K27me3 directly regulates a subset of genes by releasing poised promoters, and contributes to tumor phenotype and growth by limiting differentiation. The delayed tumor growth associated with knockdown of H3 K27M provides evidence that this highly recurrent mutation is a relevant therapeutic target.







Similar content being viewed by others
Change history
11 April 2019
The original article can be found online.
References
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM et al (2000) Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat Genet 25:25–29. https://doi.org/10.1038/75556
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF et al (2009) Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462:108–112. https://doi.org/10.1038/nature08460
Beguelin W, Popovic R, Teater M, Jiang Y, Bunting KL, Rosen M et al (2013) EZH2 is required for germinal center formation and somatic EZH2 mutations promote lymphoid transformation. Cancer Cell 23:677–692. https://doi.org/10.1016/j.ccr.2013.04.011
Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A et al (2008) An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet 40:499–507. https://doi.org/10.1038/ng.127
Bender S, Tang Y, Lindroth AM, Hovestadt V, Jones DT, Kool M et al (2013) Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell 24:660–672. https://doi.org/10.1016/j.ccr.2013.10.006
Buczkowicz P, Hoeman C, Rakopoulos P, Pajovic S, Letourneau L, Dzamba M et al (2014) Genomic analysis of diffuse intrinsic pontine gliomas identifies three molecular subgroups and recurrent activating ACVR1 mutations. Nat Genet 46:451–456. https://doi.org/10.1038/ng.2936
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D et al (2018) DNA methylation-based classification of central nervous system tumours. Nature 555:469–474. https://doi.org/10.1038/nature26000
Chan KM, Fang D, Gan H, Hashizume R, Yu C, Schroeder M et al (2013) The histone H3.3K27M mutation in pediatric glioma reprograms H3K27Methylation and gene expression. Genes Dev 27:985–990. https://doi.org/10.1101/gad.217778.113
Conway E, Healy E, Bracken AP (2015) PRC2 mediated H3K27Methylations in cellular identity and cancer. Curr Opin Cell Biol 37:42–48. https://doi.org/10.1016/j.ceb.2015.10.003
Cordero FJ, Huang Z, Grenier C, He X, Hu G, McLendon RE, Murphy SK, Hashizume R, Becher OJ (2017) Histone H3.3K27M represses to accelerate gliomagenesis in a murine model of DIPG. Mol Cancer Res 15:1243–1254. https://doi.org/10.1158/1541-7786.MCR-16-0389
Crooke ST, Witztum JL, Bennett CF, Baker BF (2018) RNA-targeted therapeutics. Cell Metab 27:714–739. https://doi.org/10.1016/j.cmet.2018.03.004
Endersby R, Zhu X, Hay N, Ellison DW, Baker SJ (2011) Nonredundant functions for Akt isoforms in astrocyte growth and gliomagenesis in an orthotopic transplantation model. Cancer Res 71:4106–4116. https://doi.org/10.1158/0008-5472.CAN-10-3597
Ernst T, Chase AJ, Score J, Hidalgo-Curtis CE, Bryant C, Jones AV, Waghorn K et al (2010) Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet 42:722–726. https://doi.org/10.1038/ng.621
Feinberg AP, Koldobskiy MA, Gondor A (2016) Epigenetic modulators, modifiers and mediators in cancer aetiology and progression. Nat Rev Genet 17:284–299. https://doi.org/10.1038/nrg.2016.13
Fellmann C, Hoffmann T, Sridhar V, Hopfgartner B, Muhar M, Roth M et al (2013) An optimized microRNA backbone for effective single-copy RNAi. Cell Rep 5:1704–1713. https://doi.org/10.1016/j.celrep.2013.11.020
Filbin MG, Tirosh I, Hovestadt V, Shaw ML, Escalante LE, Mathewson ND et al (2018) Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360:331–335. https://doi.org/10.1126/science.aao4750
Fontebasso AM, Papillon-Cavanagh S, Schwartzentruber J, Nikbakht H, Gerges N, Fiset PO et al (2014) Recurrent somatic mutations in ACVR1 in pediatric midline high-grade astrocytoma. Nat Genet 46:462–466. https://doi.org/10.1038/ng.2950
Funato K, Major T, Lewis PW, Allis CD, Tabar V (2014) Use of human embryonic stem cells to model pediatric gliomas with H3.3K27M histone mutation. Science 346:1529–1533. https://doi.org/10.1126/science.1253799
Grasso CS, Tang Y, Truffaux N, Berlow NE, Liu L, Debily MA et al (2015) Functionally defined therapeutic targets in diffuse intrinsic pontine glioma. Nat Med 21:827. https://doi.org/10.1038/nm0715-827a
Hashizume R, Andor N, Ihara Y, Lerner R, Gan H, Chen X, Fang D, Huang X et al (2014) Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma. Nat Med 20:1394–1396. https://doi.org/10.1038/nm.3716
Hirabayashi Y, Suzki N, Tsuboi M, Endo TA, Toyoda T, Shinga J et al (2009) Polycomb limits the neurogenic competence of neural precursor cells to promote astrogenic fate transition. Neuron 63:600–613. https://doi.org/10.1016/j.neuron.2009.08.021
Jones C, Baker SJ (2014) Unique genetic and epigenetic mechanisms driving paediatric diffuse high-grade glioma. Nat Rev Cancer. https://doi.org/10.1038/nrc3811
La Manno G, Gyllborg D, Codeluppi S, Nishimura K, Salto C, Zeisel A et al (2016) Molecular diversity of midbrain development in mouse, human, and stem cells. Cell 167(566–580):e519. https://doi.org/10.1016/j.cell.2016.09.027
Larson JD, Kasper LH, Paugh BS, Jin H, Wu G, Kwon CH et al (2019) Histone H3.3 K27M accelerates spontaneous brainstem glioma and drives restricted changes in bivalent gene expression. Cancer Cell 35:1–16. https://doi.org/10.1016/j.ccell.2018.11.015
Laugesen A, Helin K (2014) Chromatin repressive complexes in stem cells, development, and cancer. Cell Stem Cell 14:735–751. https://doi.org/10.1016/j.stem.2014.05.006
Le S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18
Lewis PW, Muller MM, Koletsky MS, Cordero F, Lin S, Banaszynski LA, Garcia BA et al (2013) Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340:857–861. https://doi.org/10.1126/science.1232245
Lindquist RA, Guinto CD, Rodas-Rodriguez JL, Fuentealba LC, Tate MC, Rowitch DH et al (2016) Identification of proliferative progenitors associated with prominent postnatal growth of the pons. Nat Commun 7:11628. https://doi.org/10.1038/ncomms11628
Lu TT, Heyne S, Dror E, Casas E, Leonhardt L, Boenke T et al (2018) The polycomb-dependent epigenome controls beta cell dysfunction, dedifferentiation, and diabetes. Cell Metab 27(1294–1308):e1297. https://doi.org/10.1016/j.cmet.2018.04.013
Mackay A, Burford A, Carvalho D, Izquierdo E, Fazal-Salom J, Taylor KR et al (2017) Integrated molecular meta-analysis of 1,000 pediatric high-grade and diffuse intrinsic pontine glioma. Cancer Cell 32(520–537):e525. https://doi.org/10.1016/j.ccell.2017.08.017
Mi H, Muruganujan A, Casagrande JT, Thomas PD (2013) Large-scale gene function analysis with the PANTHER classification system. Nat Protoc 8:1551–1566. https://doi.org/10.1038/nprot.2013.092
Mohammad F, Weissmann S, Leblanc B, Pandey DP, Hojfeldt JW, Comet I et al (2017) EZH2 is a potential therapeutic target for H3K27M-mutant pediatric gliomas. Nat Med 23:483–492. https://doi.org/10.1038/nm.4293
Murphy BL, Obad S, Bihannic L, Ayrault O, Zindy F, Kauppinen S et al (2013) Silencing of the miR-17 ~ 92 cluster family inhibits medulloblastoma progression. Cancer Res 73:7068–7078. https://doi.org/10.1158/0008-5472.CAN-13-0927
Nagaraja S, Vitanza NA, Woo PJ, Taylor KR, Liu F, Zhang L et al (2017) Transcriptional dependencies in diffuse intrinsic pontine glioma. Cancer Cell 31(635–652):e636. https://doi.org/10.1016/j.ccell.2017.03.011
Nikoloski G, Langemeijer SM, Kuiper RP, Knops R, Massop M et al (2010) Somatic mutations of the histone methyltransferase gene EZH2 in myelodysplastic syndromes. Nat Genet 42:665–667. https://doi.org/10.1038/ng.620
Ntziachristos P, Tsirigos A, Van Vlierberghe P, Nedjic J, Trimarchi T, Flaherty MS et al (2012) Genetic inactivation of the polycomb repressive complex 2 in T cell acute lymphoblastic leukemia. Nat Med 18:298–301. https://doi.org/10.1038/nm.2651
Orlando DA, Chen MW, Brown VE, Solanki S, Choi YJ, Olson ER et al (2014) Quantitative ChIP-Seq normalization reveals global modulation of the epigenome. Cell Rep 9:1163–1170. https://doi.org/10.1016/j.celrep.2014.10.018
Paugh BS, Broniscer A, Qu C, Miller CP, Zhang J, Tatevossian RG et al (2011) Genome-wide analyses identify recurrent amplifications of receptor tyrosine kinases and cell-cycle regulatory genes in diffuse intrinsic pontine glioma. J Clin Oncol 29:3999–4006. https://doi.org/10.1200/JCO.2011.35.5677
Pereira JD, Sansom SN, Smith J, Dobenecker MW, Tarakhovsky A, Livesey FJ (2010) Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc Natl Acad Sci USA 107:15957–15962. https://doi.org/10.1073/pnas.1002530107
Piunti A, Hashizume R, Morgan MA, Bartom ET, Horbinski CM, Marshall SA, Rendleman EJ, Ma Q et al (2017) Therapeutic targeting of polycomb and BET bromodomain proteins in diffuse intrinsic pontine gliomas. Nat Med 23:493–500. https://doi.org/10.1038/nm.4296
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K et al (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482:226–231. https://doi.org/10.1038/nature10833
Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A et al (2011) The mutational landscape of head and neck squamous cell carcinoma. Science 333:1157–1160. https://doi.org/10.1126/science.1208130
Sturm D, Bender S, Jones DT, Lichter P, Grill J, Becher O, Hawkins C et al (2014) Paediatric and adult glioblastoma: multiform (epi)genomic culprits emerge. Nat Rev Cancer 14:92–107. https://doi.org/10.1038/nrc3655
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Taylor KR, Mackay A, Truffaux N, Butterfield Y, Morozova O, Philippe C et al (2014) Recurrent activating ACVR1 mutations in diffuse intrinsic pontine glioma. Nat Genet 46:457–461. https://doi.org/10.1038/ng.2925
Venneti S, Garimella MT, Sullivan LM, Martinez D, Huse JT, Heguy A et al (2013) Evaluation of histone 3 lysine 27 trimethylation (H3K27me3) and enhancer of Zest 2 (EZH2) in pediatric glial and glioneuronal tumors shows decreased H3K27me3 in H3F3A K27M mutant glioblastomas. Brain Pathol 23:558–564. https://doi.org/10.1111/bpa.12042
Voigt P, Tee WW, Reinberg D (2013) A double take on bivalent promoters. Genes Dev 27:1318–1338. https://doi.org/10.1101/gad.219626.113
Warren KE (2012) Diffuse intrinsic pontine glioma: poised for progress. Front Oncol 2:205. https://doi.org/10.3389/fonc.2012.00205
Wu G, Barnhill RL, Lee S, Li Y, Shao Y, Easton J et al (2016) The landscape of fusion transcripts in spitzoid melanoma and biologically indeterminate spitzoid tumors by RNA sequencing. Mod Pathol 29:359–369. https://doi.org/10.1038/modpathol.2016.37
Wu G, Broniscer A, McEachron TA, Lu C, Paugh BS, Becksfort J et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44:251–253. https://doi.org/10.1038/ng.1102
Wu G, Diaz AK, Paugh BS, Rankin SL, Ju B, Li Y et al (2014) The genomic landscape of diffuse intrinsic pontine glioma and pediatric non-brainstem high-grade glioma. Nat Genet 46:444–450. https://doi.org/10.1038/ng.2938
Zeisel A, Munoz-Manchado AB, Codeluppi S, Lonnerberg P, La Manno G, Jureus A et al (2015) Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347:1138–1142. https://doi.org/10.1126/science.aaa1934
Zhang J, Ding L, Holmfeldt L, Wu G, Heatley SL, Payne-Turner D et al (2012) The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481:157–163. https://doi.org/10.1038/nature10725
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O’Keeffe S et al (2014) An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci 34:11929–11947. https://doi.org/10.1523/JNEUROSCI.1860-14.2014
Zhang Y, Sloan SA, Clarke LE, Caneda C, Plaza CA, Blumenthal PD et al (2016) Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89:37–53. https://doi.org/10.1016/j.neuron.2015.11.013
Acknowledgements
We thank Michelle Monje for sharing SUDIPG-VI cells. Surgeries and preclinical imaging were performed by the Center for In Vivo Imaging and Therapeutics which is supported by SJCRH and NCI grants P30CA021765 and R50CA211481. This work was supported by NIH grants CA096832 and CA188516 (SJB), R25CA23944 (RMN), the NCI Cancer Center Support Grant CA21765, the St Jude Children’s Research Hospital-Washington University Pediatric Cancer Genome Project, and ALSAC.
Author information
Authors and Affiliations
Contributions
Conceptualization: ABS, LHK, and SJB. Methodology: ABS, LHK, XZ, JDL, and SJB. Software: MCR and LD. Formal analysis: ABS, LHK, YF, HJ, GW, TIS, DF, BX, and SBP. Investigation and validation: ABS, LHK, XZ, JDL, JE, YS, DAY, CR, KB, JZ, RMN, and BLR. Resources: AB and CW. Data curation: ABS, LHK, YF, HJ, and GW. Writing—original draft: ABS, LHK, and SJB. Writing—review and editing: ABS, LHK, SJB, GW, HJ, JE, and DWE. Visualization: ABS, LHK, HJ, GW, TIS, and DF. Supervision: SJB, JE, SBP, DWE, and JZ. Funding acquisition: SJB and JZ.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Silveira, A.B., Kasper, L.H., Fan, Y. et al. H3.3 K27M depletion increases differentiation and extends latency of diffuse intrinsic pontine glioma growth in vivo. Acta Neuropathol 137, 637–655 (2019). https://doi.org/10.1007/s00401-019-01975-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-019-01975-4