Abstract
Objectives
The purpose of this study is to screen bladder cancer-associated biomarkers by combining lncRNA, miRNA, and mRNA expression profile of bladder cancer, and to explore bladder cancer-associated tumor markers by constructing a ceRNA regulation network.
Methods
Bladder cancer mRNA and miRNA samples were downloaded from the TCGA database; the lncRNA and mRNA detected in the RNA-seq expression profile were identified using the HUGO Gene Nomenclature Committee database.
Results
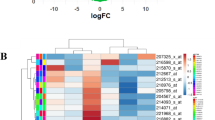
Screening for significant differentially expressed RNA resulted in 1693 mRNAs, 66 lncRNAs, and 130 miRNAs. Then, the significant differently expressed RNAs from the screening were subjected to annotation analysis of GO functional nodes and KEGG signaling pathways. A ceRNA regulation network consisting of lncRNA–miRNA–mRNA was constructed by synthesizing lncRNA–miRNA and miRNA–mRNA. Finally, a ceRNA regulation network consisting of two lncRNAs, one miRNA, and three mRNAs was obtained. KM remodeling curve analysis was performed for each factor.
Conclusions
In bladder cancer tumor samples, samples down-regulated by LINC01198, PPTRD-AS1, has-miR-216a, SEMA3D, EPHA5, and DCLK1 had a good overall survival prognosis, indicating that these several characteristic target molecules were found to be high-risk factors for bladder cancer tumors.










Similar content being viewed by others
Availability of data and material
The raw data were collected and analyzed by the authors, and are not ready to share their data, because the data have not been published.
References
Rudman SM, Malats DC (2017) Epidemiology of bladder cancer. In. Urologic Clinics of North America, pp 512–522
Parkin DM (2008) The global burden of urinary bladder cancer. Scand J Urol Nephrol Suppl 218(218):12–20
Zhang X, Han C, He J (2015) Research progress of oncogene and tumor suppressor gene in bladder cancer. Panminerva Med 57(4):191–200
Ye S, Yang L, Zhao X, Song W, Wang W, Zheng S (2014) Bioinformatics method to predict two regulation mechanism: TF–miRNA–mRNA and lncRNA–miRNA–mRNA in pancreatic cancer. J Cell Biochem Biophys 70(3):1849–1858
Liu H, Zhang Q, Lou Q, Zhang X, Cui Y, Wang P, Yang F, Wu F, Wang J, Fan T (2019) Differential analysis of lncRNA, miRNA and mRNA expression profiles and the prognostic value of lncRNA in esophageal cancer. Pathol Oncol Res 2019:1–11
Wang J, Liu X, Wu H, Ni P, Gu Z, Qiao Y, Chen N, Sun F, Fan Q (2010) CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res 38(16):5366–5383
Xiang-Hua L, Ming S, Feng-Qi N, Ying-Bin G, Er-Bao Z, Dan-Dan Y, Rong K, Rui X, Kai-Hua L, Jin-Hai L (2014) Lnc RNA HOTAIR functions as a competing endogenous RNA to regulate HER2 expression by sponging miR-331-3p in gastric cancer. Mol Cancer 13(1):92–117
Zhou X, Gao Q, Wang J, Zhang X, Liu K, Duan Z (2014) Linc-RNA-RoR acts as a “sponge” against mediation of the differentiation of endometrial cancer stem cells by microRNA-145. Gynecol Oncol 133(2):333–339
Yates B, Braschi B, Gray KA, Seal RL, Tweedie S, Bruford EA (2016) Genenames.org: the HGNC and VGNC resources in 2017. Nucleic Acids Res 41(21):9680–9687
Ritchie ME, Belinda P, Di W, Yifang H, Law CW, Wei S, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47
Wang L, Cao C, Ma Q, Zeng Q, Wang H, Cheng Z, Zhu G, Qi J, Ma H, Hai N (2014) RNA-seq analyses of multiple meristems of soybean: novel and alternative transcripts, evolutionary and functional implications. BMC Plant Biol 14(1):169
Szekely GJ, Rizzo ML (2005) Hierarchical clustering via joint between-within distances: extending ward's minimum variance method. J Classif 22(2):151–183
Press WH, Teukolsky SA, Vetterling WT, Flannery BP (2007) Numerical recipes source code CD-ROM 3rd edition: the art of scientiic computing, Section 16.4. Hierarchical clustering by phylogenetic trees. Cambridge University Press, New York
Da WH, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57
Wei HD, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13
Leonardo S, Laura P, Yvonne T, Lev K, Pier Paolo P (2011) A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146(3):353–358
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Bente F, Damgaard CK, Jargen K (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495(7441):384–388
Paraskevopoulou MD, Hatzigeorgiou AG (2016) Analyzing MiRNA–LncRNA Interactions. J Methods Mol Biol 1402(1):271–286
Paraskevopoulou MD, Vlachos IS, Karagkouni D, Georgakilas G, Kanellos I, Vergoulis T, Zagganas K, Tsanakas P, Floros E, Dalamagas T (2016) DIANA-LncBase v2: indexing microRNA targets on non-coding transcripts. Nucleic Acids Res 44:D231–D238
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Jun-Hao L, Shun L, Hui Z, Liang-Hu Q, Jian-Hua Y (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42:D92–D97
Chen T, Xie W, Xie L, Sun Y, Zhang Y, Shen Z, Sha N, Xu H, Wu Z, Hu H (2016) Expression of long noncoding RNA lncRNA-n336928 is correlated with tumor stage and grade and overall survival in bladder cancer. Biochem Biophys Res Commun 5(Suppl 1):666–670
Ratert N, Meyer HA, Jung M, Lioudmer P, Mollenkopf HJ, Wagner I, Miller K, Kilic E, Erbersdobler A, Weikert S (2013) miRNA profiling identifies candidate miRNAs for bladder cancer diagnosis and clinical outcome. J Mol Diagn 15(5):695–705
Pandolfi P, Pandolfi P (2012) 69 The ceRNA hypothesis and the non-coding revolution in cancer research and therapy. Eur J Cancer 48(Suppl 5):S16–S17
López JI, Angulo JC, Martín A, Sánchez-Chapado M, González-Corpas A, Colás B, Ropero S (2017) A DNA hypermethylation profile reveals new potential biomarkers for the evaluation of prognosis in urothelial bladder cancer. APMIS 125(9):787–796. https://doi.org/10.1111/apm.12719
Gao X, Zhang S, Chen Y, Wen X, Chen M, Wang S, Zhang Y (2019) Development of a novel six-long noncoding RNA signature predicting survival of patients with bladder urothelial carcinoma. J Cell Biochem Biophys 120(12):19796–19809. https://doi.org/10.1002/jcb.29285
Shafiei S, Kalantari E, Saeednejad Zanjani L, Abolhasani M, Asadi Lari MH, Madjd Z (2019) Increased expression of DCLK1, a novel putative CSC maker, is associated with tumor aggressiveness and worse disease-specific survival in patients with bladder carcinomas. J Exp Mol Pathol 108:164–172
Funding
None.
Author information
Authors and Affiliations
Contributions
YS and DYZ: project development, data collection, and manuscript writing. HJX and YH: data collection. YL: manuscript writing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, Y., Zhu, D., Xing, H. et al. Screening of characteristic biomolecules related to bladder cancer based on construction of ceRNA regulation network. World J Urol 38, 2835–2847 (2020). https://doi.org/10.1007/s00345-020-03086-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00345-020-03086-2