Abstract
Prp31 is one of the key tri-snRNP components essential for pre-mRNA splicing although its exact molecular function is not well studied. In a previous study, suppressor mutations were identified in the PRP31 ortholog in two spontaneous suppressors of Fgprp4 mutant deleted of the only kinase of the spliceosome in Fusarium graminearum. To further characterize the function of FgPrp31 and its relationship with FgPrp4 kinase, in this study we identified additional suppressor mutations in FgPrp31 and determined the suppressive effects of selected mutations. In total, 28 of the 35 suppressors had missense or nonsense mutations in the C terminus 465–594 aa (CT130) region of FgPrp31. The other 7 had missense or deletion mutations in the 7–64 aa region. The nonsense mutation at R464 in FgPRP31 resulted in the truncation of CT130 that contains all the putative Prp4 kinase-phosphorylation sites reported in humans, and partially rescued intron splicing defects of Fgprp4. The CT130 of FgPrp31 displayed self-inhibitory interaction with the N-terminal 1-463 (N463) region, which was reduced or abolished by the L532P, D534G, or G529D mutation in yeast two-hybrid assays. The N463 region, but not full-length FgPrp31, interacted with the N-terminal region of FgBrr2, one main U5 snRNP protein. The L532P mutation in FgPrp31 increased its interaction with FgBrr2. In contrast, suppressor mutations in FgPrp31 reduced its interaction with FgPrp6, another key component of tri-snRNP. Furthermore, we showed that FgPrp31 was phosphorylated by FgPrp4 in vivo. Site-directed mutagenesis analysis showed that phosphorylation at multiple sites in FgPrp31 is necessary to suppress Fgprp4, and S520 and S521 are important FgPrp4-phosphorylation sites. Overall, these results indicated that phosphorylation by FgPrp4 at multiple sites may release the self-inhibitory binding of FgPrp31 and affect its interaction with other components of tri-snRNP during spliceosome activation.








Similar content being viewed by others
References
Absmeier E, Wollenhaupt J, Mozaffari-Jovin S, Becke C, Lee CT, Preussner M, Heyd F, Urlaub H, Lührmann R, Santos KF, Wahl MC (2015) The large N-terminal region of the Brr2 RNA helicase guides productive spliceosome activation. Genes Dev 29:2576–2587
Agafonov DE, Kastner B, Dybkov O, Hofele RV, Liu WT, Urlaub H, Lührmann R, Stark H (2016) Molecular architecture of the human U4/U6.U5 tri-snRNP. Science 351:1416–1420
Bertram K, Agafonov DE, Dybkov O, Haselbach D, Leelaram MN, Will CL, Urlaub H, Kastner B, Lührmann R, Stark H (2017) Cryo-EM structure of a pre-catalytic human spliceosome primed for activation. Cell 170:701–713 e711. https://doi.org/10.1016/j.cell.2017.07.011
Boesler C, Rigo N, Anokhina MM, Tauchert MJ, Agafonov DE, Kastner B, Urlaub H, Ficner R, Will CL, Lührmann R (2016) A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity. Nat Commun 7:11997
Bottner CA, Schmidt H, Vogel S, Michele M, Kaufer NF (2005) Multiple genetic and biochemical interactions of Brr2, Prp8, Prp31, Prp1 and Prp4 kinase suggest a function in the control of the activation of spliceosomes in Schizosaccharomyces pombe. Curr Genet 48:151–161. https://doi.org/10.1007/s00294-005-0013-6
Bourett TM, Sweigard JA, Czymmek KJ, Carroll A, Howard RJ (2002) Reef coral fluorescent proteins for visualizing fungal pathogens. Fungal Genet Biol 37:211–220
Bruno KS, Tenjo F, Li L, Hamer JE, Xu JR (2004) Cellular localization and role of kinase activity of PMK1 in Magnaporthe grisea. Eukaryot Cell 3:1525–1532. https://doi.org/10.1128/EC.3.6.1525-1532.2004
Chen D, Wang Y, Zhou X, Wang Y, Xu JR (2014) The Sch9 kinase regulates conidium size, stress responses, and pathogenesis in Fusarium graminearum. PLoS One 9:e105811. https://doi.org/10.1371/journal.pone.0105811
Church M, Smith KC, Alhussain MM, Pennings S, Fleming AB (2017) Sas3 and Ada2(Gcn5)-dependent histone H3 acetylation is required for transcription elongation at the de-repressed FLO1 gene. Nucleic Acids Res 45:4413–4430. https://doi.org/10.1093/nar/gkx028
Cuomo CA, Güldener U, Xu JR et al (2007) The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 317:1400–1402. https://doi.org/10.1126/science.1143708
Fair FJ, Pleiss JA (2017) The power of fission: yeast as a tool for understanding complex splicing. Curr Genet 63:375–380
Figueroa M, Hammond-Kosack KE, Solomon PS (2017) A review of wheat diseases—a field perspective. Mol Plant Pathol. https://doi.org/10.1111/mpp.12618
Galej WP, Oubridge C, Newman AJ, Nagai K (2013) Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493:638–643. https://doi.org/10.1038/nature11843
Gao X, Jin Q, Jiang C, Li Y, Li C, Liu H, Kang Z, Xu JR (2016) FgPrp4 kinase is important for spliceosome B-complex activation and splicing efficiency in Fusarium graminearum. PLoS Genet 12:e1005973. https://doi.org/10.1371/journal.pgen.1005973
Goswami RS, Xu JR, Trail F, Hilburn K, Kistler HC (2006) Genomic analysis of host-pathogen interaction between Fusarium graminearum and wheat during early stages of disease development. Microbiology 6:1877–1890
Hendler A, Medina EM, Buchler NE, de Bruin RAM, Aharoni A (2017) The evolution of a G1/S transcriptional network in yeasts. Curr Genet. https://doi.org/10.1007/s00294-017-0726-3
Hou Z, Xue C, Peng Y, Katan T, Kistler HC, Xu JR (2002) A mitogen-activated protein kinase gene (MGV1) in Fusarium graminearum is required for female fertility, heterokaryon formation, and plant infection. Mol Plant Microbe Interact 15:1119–1127. https://doi.org/10.1094/MPMI.2002.15.11.1119
Iliuk A, Martinez JS, Hall MC, Tao WA (2011) Phosphorylation assay based on multifunctionalized soluble nanopolymer. Anal Chem 83:2767–2774. https://doi.org/10.1021/ac2000708
Janke C, Ortiz J, Lechner J, Shevchenko A, Shevchenko A, Magiera MM, Schramm C, Schiebel E (2001) The budding yeast proteins Spc24p and Spc25p interact with Ndc80p and Nuf2p at the kinetochore and are important for kinetochore clustering and checkpoint control. EMBO J 20:777–791. https://doi.org/10.1093/emboj/20.4.777
Jiang C, Xu JR, Li HQ (2016) Distinct cell cycle regulation during saprophytic and pathogenic growth in fungal pathogens. Curr Genet 62:185–189
Kazi T, Islam PB, Fakhoury AM (2017) FvSNF1, the sucrose non-fermenting protein kinase gene of Fusarium virguliforme, is required for cell-wall-degrading enzymes expression and sudden death syndrome development. Curr Genet 63:723–738
Kuhn AN, Kaufer NF (2003) Pre-mRNA splicing in Schizosaccharomyces pombe: regulatory role of a kinase conserved from fission yeast to mammals. Curr Genet 42:241–251
Kuhn AN, Reichl EM, Brow DA (2002) Distinct domains of splicing factor Prp8 mediate different aspects of spliceosome activation. Proc Natl Acad Sci USA 99:9145–9149
Li Y, Zhang X, Hu S, Liu H, Xu JR (2017) PKA activity is essential for relieving the suppression of hyphal growth and appressorium formation by MoSfl1 in Magnaporthe oryzae. PLoS Genet 13:e1006954. https://doi.org/10.1371/journal.pgen.1006954
Liu S, Rauhut R, Vornlocher HP, Luhrmann R (2006) The network of protein–protein interactions within the human U4/U6.U5 tri-snRNP. RNA 12:1418–1430. https://doi.org/10.1261/rna.55406
Liu S, Li P, Dybkov O, Nottrott S, Hartmuth K, Lührmann R, Carlomagno T, Wahl MC (2007) Binding of the human Prp31 Nop domain to a composite RNA-protein platform in U4 snRNP. Science 316:115–120. https://doi.org/10.1126/science.1137924
Liu W, Zhou X, Li G, Li L, Kong L, Wang C, Zhang H, Xu JR (2011) Multiple plant surface signals are sensed by different mechanisms in the rice blast fungus for appressorium formation. PLoS Pathog 7:e1001261. https://doi.org/10.1371/journal.ppat.1001261
Liu H, Zhang S, Ma J, Dai Y, Li C, Lyu X, Wang C, Xu JR (2015) Two Cdc2 Kinase genes with distinct functions in vegetative and infectious hyphae in Fusarium graminearum. PLoS Pathog 11:e1004913. https://doi.org/10.1371/journal.ppat.1004913
Makarova OV, Makarov EM, Liu S, Vornlocher HP, Luhrmann R (2002) Protein 61K, encoded by a gene (PRPF31) linked to autosomal dominant retinitis pigmentosa, is required for U4/U6⋅U5 tri-snRNP formation and pre-mRNA splicing. EMBO J 21:1148–1157. https://doi.org/10.1093/emboj/21.5.1148
McKay SL, Johnson TL (2010) A bird’s-eye view of post-translational modifications in the spliceosome and their roles in spliceosome dynamics. Mol Biosyst 6:2093–2102. https://doi.org/10.1039/c002828b
Mozaffari-Jovin S, Santos KF, Hsiao HH, Will CL, Urlaub H, Wahl MC, Luhrmann R (2012) The Prp8 RNase H-like domain inhibits Brr2-mediated U4/U6 snRNA unwinding by blocking Brr2 loading onto the U4 snRNA. Genes Dev 26:2422–2434. https://doi.org/10.1101/gad.200949.112
Mozaffari-Jovin S, Wandersleben T, Santos KF, Will CL, Luhrmann R, Wahl MC (2013) Inhibition of RNA helicase Brr2 by the C-terminal tail of the spliceosomal protein Prp. 8 Science 341:80–84. https://doi.org/10.1126/science.1237515
Nguyen TH, Galej WP, Bai XC, Savva CG, Newman AJ, Scheres SH, Nagai K (2015) The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 523:47–52. https://doi.org/10.1038/nature14548
Schneider M, Hsiao HH, Will CL, Giet R, Urlaub H, Luhrmann R (2010) Human PRP4 kinase is required for stable tri-snRNP association during spliceosomal B complex formation. Nat Struct Mol Biol 17:216–221. https://doi.org/10.1038/nsmb.1718
Theuser M, Hobartner C, Wahl MC, Santos KF (2016) Substrate-assisted mechanism of RNP disruption by the spliceosomal Brr2 RNA helicase. Proc Natl Acad Sci USA 113:7798–7803. https://doi.org/10.1073/pnas.1524616113
Vithana EN, Abu-Safieh L, Allen MJ et al (2001) A human homolog of yeast pre-mRNA splicing gene, PRP31, underlies autosomal dominant retinitis pigmentosa on chromosome 19q13.4 (RP11). Mol Cell 8:375–381
Wan R, Yan C, Bai R, Huang G, Shi Y (2016a) Structure of a yeast catalytic step I spliceosome at 3.4 A resolution. Science 353:895–904. https://doi.org/10.1126/science.aag2235
Wan R, Yan C, Bai R, Wang L, Huang M, Wong CC, Shi Y (2016b) The 3.8 A structure of the U4/U6.U5 tri-snRNP: insights into spliceosome assembly and catalysis. Science 351:466–475. https://doi.org/10.1126/science.aad6466
Wang C, Zhang S, Hou R et al (2011) Functional analysis of the kinome of the wheat scab fungus Fusarium graminearum. PLoS Pathog 7:e1002460
Weidenhammer EM, Singh M, Ruiz-Noriega M, Woolford JL (1996) The PRP31 gene encodes a novel protein required for pre-mRNA splicing in Saccharomyces cerevisiae. Nucleic Acids Res 24:1164–1170
Weidenhammer EM, Ruiz-Noriega M, Woolford JL (1997) Prp31p promotes the association of the U4/U6 × U5 tri-snRNP with prespliceosomes to form spliceosomes in Saccharomyces cerevisiae. Mol Cell Biol 17:3580–3588
Will CL, Luhrmann R (2011) Spliceosome structure and function. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a003707
Xie F, Sun S, Xu A, Zheng S, Xue M, Wu P, Zeng JH, Bai L (2014) Advanced oxidation protein products induce intestine epithelial cell death through a redox-dependent, c-jun N-terminal kinase and poly (ADP-ribose) polymerase-1-mediated pathway. Cell Death Dis 5:e1006. https://doi.org/10.1038/cddis.2013.542
Yang C, Liu H, Li G, Liu M, Yun Y, Wang C, Ma Z, Xu JR (2015) The MADS-box transcription factor FgMcm1 regulates cell identity and fungal development in Fusarium graminearum. Environ Microbiol 17:2762–2776. https://doi.org/10.1111/1462-2920.12747
Zhao X, Xu JR (2007) A highly conserved MAPK-docking site in Mst7 is essential for Pmk1 activation in Magnaporthe grisea. Mol Microbiol 63:881–894
Zhou X, Li G, Xu JR (2011a) Efficient approaches for generating GFP fusion and epitope-tagging constructs in filamentous fungi. Methods Mol Biol 722:199–212
Zhou X, Liu W, Wang C, Xu Q, Wang Y, Ding S, Xu JR (2011b) A MADS-box transcription factor MoMcm1 is required for male fertility, microconidium production and virulence in Magnaporthe oryzae. Mol Microbiol 80:33–53
Acknowledgements
We thank Drs. Chenfang Wang, Cong Jiang, Huiquan Liu, and Anton Iliuk for fruitful discussions. We also thank Dr. Yimei Zhang and Ms. Xiaoping Li for assistance with the yeast two-hybrid and co-IP assays. This work was supported by Grants from the Nature Science Foundation of China (31600117), US Wheat and Barley Scab Initiative (106616), and Fundamental Research Funds for the Central Universities (2452016019).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: XG, QJ, JRX. Performed the experiments: XG, CS, JZ, KY, JW. Analyzed the data: XG, QJ, JRX. Contributed reagents/materials/analysis tools: XG, QJ. Wrote the paper: XG, QJ, JRX.
Corresponding authors
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.

294_2018_838_MOESM2_ESM.jpg
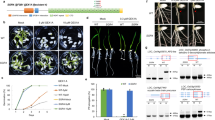
Fig S1 Deletion of C-terminal 130 aa in FgPRP31 (a) The FgPRP31 gene, FgPRP31ΔCT130 replacement construct, and primers used. (b) Three-day-old PDA cultures (Top), two-week-old mating cultures (Middle), and plant infection assays on wheat coleoptile (Bottom) of the wild type (WT) and the FgPRP31ΔCT130 mutant strains (JPG 2882 KB)

294_2018_838_MOESM3_ESM.jpg
Fig S2 Yeast two-hybrid assays for the FgPrp31-FgBrr2 and FgPrp31-FgPrp6 interactions. Yeast transformants expressing the marked bait and prey constructs were assayed for growth on SD-Trp-Leu-His plates and LacZ activities. The positive and negative controls are from the Matchmaker kit (JPG 3328 KB)
Rights and permissions
About this article
Cite this article
Gao, X., Zhang, J., Song, C. et al. Phosphorylation by Prp4 kinase releases the self-inhibition of FgPrp31 in Fusarium graminearum. Curr Genet 64, 1261–1274 (2018). https://doi.org/10.1007/s00294-018-0838-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-018-0838-4