Abstract
Cancer metastasis risk increases in older individuals, but the mechanisms for this risk increase are unclear. Many peritoneal cancers, including ovarian cancer, preferentially metastasize to peritoneal fat depots. However, there is a dearth of studies exploring aged peritoneal adipose tissue in the context of cancer. Because adipose tissue produces signals which influence several diseases including cancer, proteomics of adipose tissue in aged and young mice may provide insight into metastatic mechanisms. We analyzed mesenteric, omental, and uterine adipose tissue groups from the peritoneal cavities of young and aged C57BL/6J mouse cohorts with a low-fraction SDS-PAGE gelLC-MS/MS method. We identified 2308 protein groups and quantified 2167 groups, among which several protein groups showed twofold or greater abundance differences between the aged and young cohorts. Cancer-related gene products previously identified as significant in another age-related study were found altered in this study. Several gene products known to suppress proliferation and cellular invasion were found downregulated in the aged cohort, including R-Ras, Arid1a, and heat shock protein β1. In addition, multiple protein groups were identified within single cohorts, including the proteins Cd11a, Stat3, and Ptk2b. These data suggest that adipose tissue is a strong candidate for analysis to identify possible contributors to cancer metastasis in older subjects. The results of this study, the first of its kind using uterine adipose tissue, contribute to the understanding of the role of adipose tissue in age-related alteration of oncogenic pathways, which may help elucidate the mechanisms of increased metastatic tumor burden in the aged.

We analyzed mesenteric, omental, and uterine adipose tissue groups from the peritoneal cavities of young and aged C57BL/6J mouse cohorts with a low-fraction SDS-PAGE gelLC-MS/MS method. These fat depots are preferential sites for many peritoneal cancers. The results of this study, the first of its kind using uterine adipose tissue, contribute to the understanding of the role of adipose tissue in age-related alteration of oncogenic pathways, which may help elucidate the mechanisms of increased metastatic tumor burden in the aged.




Similar content being viewed by others
References
Kim EY, Kim WK, K-J O, Han BS, Lee SC, Bae K-H. Recent advances in proteomic studies of adipose tissues and adipocytes. Int J Mol Sci. 2015;16(3):4581–99. https://doi.org/10.3390/ijms16034581.
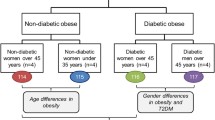
Gómez-Serrano M, Camafeita E, García-Santos E, López JA, Rubio MA, Sánchez-Pernaute A, et al. Proteome-wide alterations on adipose tissue from obese patients as age-, diabetes- and gender-specific hallmarks. Sci Rep. 2016;6 https://doi.org/10.1038/srep25756.
Galic S, Oakhill JS, Steinberg GR. Adipose tissue as an endocrine organ. Mol Cell Endocrinol. 2010;316(2):129–39. https://doi.org/10.1016/j.mce.2009.08.018.
Kershaw EE, Flier JS. Adipose tissue as an endocrine organ. J Clin Endocrinol Metab. 2004;89(6):2548–56. https://doi.org/10.1210/jc.2004-0395.
Coelho M, Oliveira T, Fernandes R. Biochemistry of adipose tissue: an endocrine organ. Arch Med Sci. 2013;9(2):191–200. https://doi.org/10.5114/aoms.2013.33181.
Boden G. Effects of free fatty acids (FFA) on glucose metabolism: significance for insulin resistance and type 2 diabetes. Exp Clin Endocrinol Diabetes. 2003;111(3):121–4. https://doi.org/10.1055/s-2003-39781.
Duvnjak L, Duvnjak M. The metabolic syndrome - an ongoing story. J Physiol Pharmacol. 2009;60(Suppl 7):19–24.
Matsuzawa Y. Therapy insight: adipocytokines in metabolic syndrome and related cardiovascular disease. Nat Clin Pract Cardiovasc Med. 2006;3(1):35–42. https://doi.org/10.1038/ncpcardio0380.
Fontana L, Eagon JC, Trujillo ME, Scherer PE, Klein S. Visceral fat adipokine secretion is associated with systemic inflammation in obese humans. Diabetes. 2007;56(4):1010–3. https://doi.org/10.2337/db06-1656.
Frayn KN. Visceral fat and insulin resistance--causative or correlative? Br J Nutr. 2000;83(Suppl 1):S71–7.
Gastaldelli A, Miyazaki Y, Pettiti M, Matsuda M, Mahankali S, Santini E, et al. Metabolic effects of visceral fat accumulation in type 2 diabetes. J Clin Endocrinol Metab. 2002;87(11):5098–103. https://doi.org/10.1210/jc.2002-020696.
Lehr S, Hartwig S, Lamers D, Famulla S, Müller S, Hanisch F-G, et al. Identification and validation of novel adipokines released from primary human adipocytes. Mol Cell Proteomics. 2012;11(1):M111.010504. https://doi.org/10.1074/mcp.M111.010504.
Hajer GR, van Haeften TW, Visseren FLJ. Adipose tissue dysfunction in obesity, diabetes, and vascular diseases. Eur Heart J. 2008;29(24):2959–71. https://doi.org/10.1093/eurheartj/ehn387.
Bjørndal B, Burri L, Staalesen V, Skorve J, Berge RK. Different adipose depots: their role in the development of metabolic syndrome and mitochondrial response to hypolipidemic agents. J Obes. 2011;2011:490650. https://doi.org/10.1155/2011/490650.
Peral B, Camafeita E, Fernández-Real J-M, López JA. Tackling the human adipose tissue proteome to gain insight into obesity and related pathologies. Expert Rev Proteomics. 2009;6(4):353–61. https://doi.org/10.1586/epr.09.53.
Guerre-Millo M. Adipose tissue hormones. J Endocrinol Investig. 2002;25(10):855–61. https://doi.org/10.1007/BF03344048.
Kim S-J, Chae S, Kim H, Mun D-G, Back S, Choi HY, et al. A protein profile of visceral adipose tissues linked to early pathogenesis of type 2 diabetes mellitus. Mol Cell Proteomics. 2014;13(3):811–22. https://doi.org/10.1074/mcp.M113.035501.
Ma S, Jing F, Xu C, Zhou L, Song Y, Yu C, et al. Thyrotropin and obesity: increased adipose triglyceride content through Glycerol-3-phosphate acyltransferase 3. Sci Rep. 2015;5 https://doi.org/10.1038/srep07633.
Brockman D, Chen X. Proteomics in the characterization of adipose dysfunction in obesity. Adipocyte. 2012;1(1):25–37. https://doi.org/10.4161/adip.19129.
Fang L, Kojima K, Zhou L, Crossman DK, Mobley JA, Grams J. Analysis of the human proteome in subcutaneous and visceral fat depots in diabetic and non-diabetic patients with morbid obesity. J Proteomics Bioinform. 2015;8(6):133–41. https://doi.org/10.4172/jpb.1000361.
Desruisseaux MS, Nagajyothi TME, Tanowitz HB, Scherer PE. Adipocyte, adipose tissue, and infectious disease. Infect Immun. 2007;75(3):1066–78. https://doi.org/10.1128/IAI.01455-06.
Thomas LW. The chemical composition of adipose tissue of man and mice. Exp Physiol. 1962;47(2):179–88. https://doi.org/10.1113/expphysiol.1962.sp001589.
Liu Y, Metzinger MN, Lewellen KA, Cripps SN, Carey KD, Harper EI, et al. Obesity contributes to ovarian cancer metastatic success through increased lipogenesis, enhanced vascularity, and decreased infiltration of M1 macrophages. Cancer Res. 2015;75(23):5046–57. https://doi.org/10.1158/0008-5472.CAN-15-0706.
Booth A, Magnuson A, Fouts J, Foster M. Adipose tissue, obesity and adipokines: role in cancer promotion. Horm Mol Biol Clin Invest. 2015;21(1):57–74. https://doi.org/10.1515/hmbci-2014-0037.
Rytka JM, Wueest S, Schoenle EJ, Konrad D. The portal theory supported by venous drainage-selective fat transplantation. Diabetes. 2011;60(1):56–63. https://doi.org/10.2337/db10-0697.
Ozcelik F, Yuksel C, Arslan E, Genc S, Omer B, Serdar MA. Relationship between visceral adipose tissue and adiponectin, inflammatory markers and thyroid hormones in obese males with hepatosteatosis and insulin resistance. Arch Med Res. 2013;44(4):273–80. https://doi.org/10.1016/j.arcmed.2013.04.001.
Lengyel E. Ovarian cancer development and metastasis. Am J Pathol. 2010;177(3):1053–64. https://doi.org/10.2353/ajpath.2010.100105.
Nieman KM, Kenny HA, Penicka CV, Ladanyi A, Buell-Gutbrod R, Zillhardt MR, et al. Adipocytes promote ovarian cancer metastasis and provide energy for rapid tumor growth. Nat Med. 2011;17(11):1498–503. https://doi.org/10.1038/nm.2492.
Clark R, Krishnan V, Schoof M, Rodriguez I, Theriault B, Chekmareva M, et al. Milky spots promote ovarian cancer metastatic colonization of peritoneal adipose in experimental models. Am J Pathol. 2013;183(2):576–91. https://doi.org/10.1016/j.ajpath.2013.04.023.
Khan SM, Funk HM, Thiolloy S, Lotan TL, Hickson J, Prins GS, et al. In vitro metastatic colonization of human ovarian cancer cells to the omentum. Clin Exp Metastasis. 2010;27(3):185–96. https://doi.org/10.1007/s10585-010-9317-0.
Pradeep S, Kim SW, SY W, Nishimura M, Chaluvally-Raghavan P, Miyake T, et al. Hematogenous metastasis of ovarian cancer: rethinking mode of spread. Cancer Cell. 2014;26(1):77–91. https://doi.org/10.1016/j.ccr.2014.05.002.
Cancer of the Ovary—Cancer Stat Facts. NIH National Cancer Institute. https://seer.cancer.gov/statfacts/html/ovary.html. Accessed 10/20/2017 2017.
Leitzmann MF, Koebnick C, Danforth KN, Brinton LA, Moore SC, Hollenbeck AR, et al. Body mass index and risk of ovarian cancer. Cancer. 2009;115(4):812–22. https://doi.org/10.1002/cncr.24086.
Nagarsheth N, Wicha MS, Zou W. Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat Rev Immunol. 2017;17(9):559–72. https://doi.org/10.1038/nri.2017.49.
Pasing Y, Colnoe S, Hansen T. Proteomics of hydrophobic samples: fast, robust and low-cost workflows for clinical approaches. Proteomics. 2016; https://doi.org/10.1002/pmic.201500462.
Forner F, Kumar C, Luber CA, Fromme T, Klingenspor M, Mann M. Proteome differences between brown and white fat mitochondria reveal specialized metabolic functions. Cell Metab. 2009;10(4):324–35. https://doi.org/10.1016/j.cmet.2009.08.014.
Okita N, Hayashida Y, Kojima Y, Fukushima M, Yuguchi K, Mikami K, et al. Differential responses of white adipose tissue and brown adipose tissue to caloric restriction in rats. Mech Ageing Dev. 2012;133(5):255–66. https://doi.org/10.1016/j.mad.2012.02.003.
Li J, Zhao W-G, Shen Z-F, Yuan T, Liu S-N, Liu Q, et al. Comparative proteome analysis of brown adipose tissue in obese C57BL/6J mice using iTRAQ-coupled 2D LC-MS/MS. PLoS One. 2015;10(3) https://doi.org/10.1371/journal.pone.0119350.
Müller S, Balaz M, Stefanicka P, Varga L, Amri E-Z, Ukropec J, et al. Proteomic analysis of human brown adipose tissue reveals utilization of coupled and uncoupled energy expenditure pathways. Sci Rep. 2016;6:30030. https://doi.org/10.1038/srep30030.
Riordan NH, Ichim TE, Min W-P, Wang H, Solano F, Lara F, et al. Non-expanded adipose stromal vascular fraction cell therapy for multiple sclerosis. J Transl Med. 2009;7:29. https://doi.org/10.1186/1479-5876-7-29.
Peinado JR, Jimenez-Gomez Y, Pulido MR, Ortega-Bellido M, Diaz-Lopez C, Padillo FJ, et al. The stromal-vascular fraction of adipose tissue contributes to major differences between subcutaneous and visceral fat depots. Proteomics. 2010;10(18):3356–66. https://doi.org/10.1002/pmic.201000350.
Lee MJ, Kim J, Kim MY, Bae Y-S, Ryu SH, Lee TG, et al. Proteomic analysis of tumor necrosis factor-α-induced secretome of human adipose tissue-derived mesenchymal stem cells. J Proteome Res. 2010;9(4):1754–62. https://doi.org/10.1021/pr900898n.
Alvarez-Llamas G, Szalowska E, MPd V, Weening D, Landman K, Hoek A, et al. Characterization of the human visceral adipose tissue secretome. Mol Cell Proteomics. 2007;6(4):589–600. https://doi.org/10.1074/mcp.M600265-MCP200.
Life span as a biomarker. https://www.jax.org/research-and-faculty/research-labs/the-harrison-lab/gerontology/life-span-as-a-biomarkerfiles/3/life-span-as-a-biomarker.html. Accessed 10/20/2017 2017.
Walther DM, Mann M. Accurate quantification of more than 4000 mouse tissue proteins reveals minimal proteome changes during aging. Mol Cell Proteomics. 2011;10(2):M110.004523. https://doi.org/10.1074/mcp.M110.004523.
Stauch KL, Purnell PR, Villeneuve LM, Fox HS. Data for mitochondrial proteomic alterations in the aging mouse brain. Data Brief. 2015;4:127–9. https://doi.org/10.1016/j.dib.2015.05.004.
Thakur D, Rejtar T, Wang D, Bones J, Cha S, Clodfelder-Miller B, et al. Microproteomic analysis of 10,000 laser captured microdissected breast tumor cells using short-range sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and porous layer open tubular (PLOT) LC-MS/MS. J Chromatogr A. 2011;1218(45):8168–74. https://doi.org/10.1016/j.chroma.2011.09.022.
Shevchenko A, Tomas H, Havli J, Olsen JV, Mann M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc. 2007;1(6):2856–60. https://doi.org/10.1038/nprot.2006.468.
Tyanova S, Temu T, Cox J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc. 2016;11(12):2301–19. https://doi.org/10.1038/nprot.2016.136.
Cox J, Hein MY, Luber CA, Paron I, Nagaraj N, Mann M. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol Cell Proteomics. 2014;13(9):2513–26. https://doi.org/10.1074/mcp.M113.031591.
Wessel D, Flügge UI. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal Biochem. 1984;138(1):141–3.
Di Girolamo F, Ponzi M, Crescenzi M, Alessandroni J, Guadagni F. A simple and effective method to analyze membrane proteins by SDS-PAGE and MALDI mass spectrometry. Anticancer Res. 2010;30(4):1121–9.
Vertommen A, Panis B, Swennen R, Carpentier SC. Evaluation of chloroform/methanol extraction to facilitate the study of membrane proteins of non-model plants. Planta. 2010;231(5):1113–25. https://doi.org/10.1007/s00425-010-1121-1.
Adachi J, Kumar C, Zhang Y, Mann M. In-depth analysis of the adipocyte proteome by mass spectrometry and bioinformatics. Mol Cell Proteomics. 2007;6(7):1257–73. https://doi.org/10.1074/mcp.M600476-MCP200.
Xie X, Yi Z, Bowen B, Wolf C, Flynn CR, Sinha S, et al. Characterization of the human adipocyte proteome and reproducibility of protein abundance by one-dimensional gel electrophoresis and HPLC-ESI-MS/MS. J Proteome Res. 2010;9(9):4521–34. https://doi.org/10.1021/pr100268f.
Weston LA, Bauer KM, Hummon AB. Comparison of bottom-up proteomic approaches for LC-MS analysis of complex proteomes. Anal Methods. 2013;5(18) https://doi.org/10.1039/C3AY40853A.
Piersma SR, Warmoes MO, de Wit M, de Reus I, Knol JC, Jiménez CR. Whole gel processing procedure for GeLC-MS/MS based proteomics. Proteome Sci. 2013;11:17. https://doi.org/10.1186/1477-5956-11-17.
Warmoes M, Jaspers JE, Pham TV, Piersma SR, Oudgenoeg G, Massink MPG, et al. Proteomics of mouse BRCA1-deficient mammary tumors identifies DNA repair proteins with potential diagnostic and prognostic value in human breast cancer. Mol Cell Proteomics. 2012;11(7) https://doi.org/10.1074/mcp.M111.013334.
Ma Y, Gao J, Yin J, Gu L, Liu X, Chen S, et al. Identification of a novel function of adipocyte plasma membrane-associated protein (APMAP) in gestational diabetes mellitus by proteomic analysis of omental adipose tissue. J Proteome Res. 2016;15(2):628–37. https://doi.org/10.1021/acs.jproteome.5b01030.
Ke M, Wu H, Zhu Z, Zhang C, Zhang Y, Deng Y. Differential proteomic analysis of white adipose tissues from T2D KKAy mice by LC-ESI-QTOF. Proteomics. 2017;17(5) https://doi.org/10.1002/pmic.201600219.
Tomé-Carneiro J, Crespo MC, García-Calvo E, Luque-García JL, Dávalos A, Visioli F. Proteomic evaluation of mouse adipose tissue and liver following hydroxytyrosol supplementation. Food Chem Toxicol. 2017;107(Part A):329–38. https://doi.org/10.1016/j.fct.2017.07.009.
Meierhofer D, Weidner C, Sauer S (2014) Integrative analysis of transcriptomics, proteomics, and metabolomics data of white adipose and liver tissue of high-fat diet and rosiglitazone-treated insulin-resistant mice identified pathway alterations and molecular hubs. J Proteome Res 13(12):5592–5602
Oliva K, Barker G, Rice GE, Bailey MJ, Lappas M. 2D-DIGE to identify proteins associated with gestational diabetes in omental adipose tissue. J Endocrinol. 2013;218(2):165–78. https://doi.org/10.1530/JOE-13-0010.
Thomas PD, Campbell MJ, Kejariwal A, Mi H, Karlak B, Daverman R, et al. PANTHER: a library of protein families and subfamilies indexed by function. Genome Res. 2003;13(9):2129–41. https://doi.org/10.1101/gr.772403.
Thomas PD, Kejariwal A, Guo N, Mi H, Campbell MJ, Muruganujan A, et al. Applications for protein sequence–function evolution data: mRNA/protein expression analysis and coding SNP scoring tools. Nucleic Acids Res. 2006;34(suppl_2):W645–W50. https://doi.org/10.1093/nar/gkl229.
Song J, Zheng B, Bu X, Fei Y, Shi S. Negative association of R-Ras activation and breast cancer development. Oncol Rep. 2014;31(6):2776–84. https://doi.org/10.3892/or.2014.3121.
He F, Li J, Xu J, Zhang S, Xu Y, Zhao W, et al. Decreased expression of ARID1A associates with poor prognosis and promotes metastases of hepatocellular carcinoma. J Exp Clin Cancer Res. 2015;34(1) https://doi.org/10.1186/s13046-015-0164-3.
Vargas-Roig LM, Fanelli MA, López LA, Gago FE, Tello O, Aznar JC, et al. Heat shock proteins and cell proliferation in human breast cancer biopsy samples. Cancer Detect Prev. 1997;21(5):441–51.
Jiang E, Perrard XD, Yang D, Khan IM, Perrard JL, Smith CW, et al. Essential role of CD11a in CD8+ T-cell accumulation and activation in adipose tissue. Arterioscler Thromb Vasc Biol. 2014;34(1):34–43. https://doi.org/10.1161/ATVBAHA.113.302077.
Nguyen AV, Wu Y-Y, Liu Q, Wang D, Nguyen S, Loh R, et al. STAT3 in epithelial cells regulates inflammation and tumor progression to malignant state in colon. Neoplasia. 2013;15(9):998–1008.
Di Cioccio V, Strippoli R, Bizzarri C, Troiani G, Cervellera MN, Gloaguen I, et al. Key role of proline-rich tyrosine kinase 2 in interleukin-8 (CXCL8/IL-8)-mediated human neutrophil chemotaxis. Immunology. 2004;111(4):407–15. https://doi.org/10.1111/j.1365-2567.2004.01822.x.
Ouderkirk JL, Krendel M. Non-muscle myosins in tumor progression, cancer cell invasion and metastasis. Cytoskeleton (Hoboken). 2014;71(8):447–63. https://doi.org/10.1002/cm.21187.
Calle EE, Kaaks R. Overweight, obesity and cancer: epidemiological evidence and proposed mechanisms. Nat Rev Cancer. 2004;4(8):579–91. https://doi.org/10.1038/nrc1408.
Tripathi S, Flobak Å, Chawla K, Baudot A, Bruland T, Thommesen L, et al. The gastrin and cholecystokinin receptors mediated signaling network: a scaffold for data analysis and new hypotheses on regulatory mechanisms. BMC Syst Biol. 2015;9:40. https://doi.org/10.1186/s12918-015-0181-z.
Acknowledgements
We gratefully acknowledge the assistance of the Notre Dame Mass Spectrometry and Proteomics Facility (MSPF).
Funding
PF was supported by an Arthur J. Schmitt Presidential Fellowship. ABH was supported by the National Institutes of Health (R01GM110406) and the National Science Foundation (CAREER Award, CHE-1351595). EAL was supported by a National Science Foundation Graduate Research Fellowship Program grant DGE-1313583. MSS is supported by the National Institutes of Health (RO1CA109545) and the Leo and Anne Albert Charitable Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This research includes animal research. All mouse procedures were carried out according to the regulations of the University of Notre Dame Animal Care and Use Committee (IACUC 14-02-1577) and with approval of the Notre Dame Institutional Review Board.
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Feist, P.E., Loughran, E.A., Stack, M.S. et al. Quantitative proteomic analysis of murine white adipose tissue for peritoneal cancer metastasis. Anal Bioanal Chem 410, 1583–1594 (2018). https://doi.org/10.1007/s00216-017-0813-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0813-9