Abstract
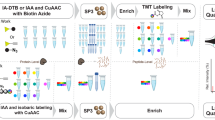
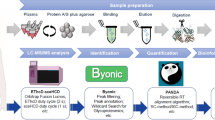
Detailed characterization of glycoprotein structures requires determining both the sites of glycosylation as well as the glycan structures associated with each site. In this work, we developed an analytical strategy for characterization of intact N-glycopeptides in complex proteome samples. In the first step, tryptic glycopeptides were enriched using ZIC-HILIC. Secondly, a portion of the glycopeptides was treated with endoglycosidase H (Endo H) to remove high-mannose (Man) and hybrid N-linked glycans. Thirdly, a fraction of the Endo H-treated glycopeptides was further subjected to PNGase F treatment in 18O water to remove the remaining complex glycans. The intact glycopeptides and deglycosylated peptides were analyzed by nano-RPLC–MS/MS, and the glycan structures and the peptide sequences were identified by using the Byonic or pFind tools. Sequential digestion by endoglycosidase provided candidate glycosites information and indication of the glycoforms on each glycopeptide, thus helping to confine the database search space and improve the confidence regarding intact glycopeptide identification. We demonstrated the effectiveness of this approach using RNase B and IgG and applied this sequential digestion strategy for the identification of glycopeptides from the HepG2 cell line. We identified 4514 intact glycopeptides coming from 947 glycosites and 1011 unique peptide sequences from HepG2 cells. The intensity of different glycoforms at a specific glycosite was obtained to reach the occupancy ratios of site-specific glycoforms. These results indicate that our method can be used for characterizing site-specific protein glycosylation in complex samples.

Through integrating the information of intact glycopeptide, fragment ions filters and endoglycosidase digestion, the reliability of the identification could be significantly improved. We quantified the site-specific glycoforms occupancy ratios through the MS response signaling of each glycopeptide at the same time.





Similar content being viewed by others
References
Mariño K, Bones J, Kattla JJ, Rudd PM. A systematic approach to protein glycosylation analysis: a path through the maze. Nat Chem Biol. 2010;6(10):713–23.
Zhu Z, Desaire H. Carbohydrates on proteins: site-specific glycosylation analysis by mass spectrometry. Annu Rev Anal Chem. 2015;8:463–83.
Kuo C-W, Wu I-L, Hsiao H-H, Khoo K-H. Rapid glycopeptide enrichment and N-glycosylation site mapping strategies based on amine-functionalized magnetic nanoparticles. Anal Bioanal Chem. 2012;402(9):2765–76.
Dalziel M, Crispin M, Scanlan CN, Zitzmann N, Dwek RA. Emerging principles for the therapeutic exploitation of glycosylation. Science. 2014;343(6166):1235681.
Kailemia MJ, Park D, Lebrilla CB. Glycans and glycoproteins as specific biomarkers for cancer. Analytical and bioanalytical chemistry. 2016:1–16.
Hu W, Su X, Zhu Z, Go EP, Desaire H. GlycoPep MassList: software to generate massive inclusion lists for glycopeptide analyses. Analytical and bioanalytical chemistry. 2016:1–10.
Stowell SR, Ju T, Cummings RD. Protein glycosylation in cancer. Annu Rev Pathol. 2015;10:473.
Liu M, Zhang Y, Chen Y, Yan G, Shen C, Cao J, et al. Efficient and accurate glycopeptide identification pipeline for high-throughput site-specific N-glycosylation analysis. J Proteome Res. 2014;13(6):3121–9.
Kolli V, Schumacher KN, Dodds ED. Engaging challenges in glycoproteomics: recent advances in MS-based glycopeptide analysis. Bioanalysis. 2015;7(1):113–31.
Desaire H. Glycopeptide analysis, recent developments and applications. Mol Cell Proteomics. 2013;12(4):893–901.
He L, Xin L, Shan B, Lajoie GA, Ma B. GlycoMaster DB: software to assist the automated identification of N-linked glycopeptides by tandem mass spectrometry. J Proteome Res. 2014;13(9):3881–95.
Toghi Eshghi S, Shah P, Yang W, Li X, Zhang H. GPQuest: a spectral library matching algorithm for site-specific assignment of tandem mass spectra to intact N-glycopeptides. Anal Chem. 2015;87(10):5181–8.
Bern M, Kil YJ, Becker C. Byonic. Advanced peptide and protein identification software. Current Protocols in Bioinformatics. 2012:13.20. 1–13.20. 14.
Becker C, Tang W, Kil YJ, Yin X, Mayr M, Khoo K-H, et al. Search strategies for glycopeptide identification. J Biomol Tech. 2013;24(Suppl):S33.
Zeng W, Liu M, Zhang Y, Wu J, Fang P, Peng C, et al. pGlyco: a pipeline for the identification of intact N-glycopeptides by using HCD-and CID-MS/MS and MS3. Sci Rep. 2016;6:25102.
Lee LY, Moh ES, Parker BL, Bern M, Packer NH, Thaysen-Andersen M. Toward automated N-glycopeptide identification in glycoproteomics. J Proteome Res. 2016;15(10):3904–15.
Hu H, Khatri K, Klein J, Leymarie N, Zaia J. A review of methods for interpretation of glycopeptide tandem mass spectral data. Glycoconj J. 2016;33(3):285–96.
Alley Jr WR, Mann BF, Novotny MV. High-sensitivity analytical approaches for the structural characterization of glycoproteins. Chem Rev. 2013;113(4):2668–732.
Zhang C, Ye Z, Xue P, Shu Q, Zhou Y, Ji Y, et al. Evaluation of different N-glycopeptide enrichment methods for N-glycosylation sites mapping in mouse brain. J Proteome Res. 2016;15(9):2960–8.
Malerod H, Graham RL, Sweredoski MJ, Hess S. Comprehensive profiling of N-linked glycosylation sites in HeLa cells using hydrazide enrichment. J Proteome Res. 2013;12(1):248–59.
Zhang W, Wang H, Zhang L, Yao J, Yang P. Large-scale assignment of N-glycosylation sites using complementary enzymatic deglycosylation. Talanta. 2011;85(1):499–505.
Cao Q, Zhao X, Zhao Q, Lv X, Ma C, Li X, et al. Strategy integrating stepped fragmentation and glycan diagnostic ion-based spectrum refinement for the identification of core fucosylated glycoproteome using mass spectrometry. Anal Chem. 2014;86(14):6804–11.
Wang L, Li DQ, Fu Y, Wang HP, Zhang JF, Yuan ZF, et al. pFind 2.0: a software package for peptide and protein identification via tandem mass spectrometry. Rapid Commun Mass Spectrom. 2007;21(18):2985–91.
Li D, Fu Y, Sun R, Ling CX, Wei Y, Zhou H, et al. pFind: a novel database-searching software system for automated peptide and protein identification via tandem mass spectrometry. Bioinformatics. 2005;21(13):3049–50.
Park G, Kim J, Hwang H, Lee J, Ahn Y, Lee H, et al. Integrated GlycoProteome Analyzer (I-GPA) for automated identification and quantitation of site-specific N-glycosylation. Sci Rep. 2016;6:21175.
Tyanova S, Temu T, Cox J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc. 2016;11(12):2301–19.
Acknowledgements
We are grateful for the financial support from the National Key Program for Basic Research of China (2016YFA0501300, 2014CBA02001), the State Key Laboratory of NBC Protection of Civilian (SKLNBC03010), and the National Natural Science Foundation of China (81530021).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Yu, Z., Zhao, X., Tian, F. et al. Sequential fragment ion filtering and endoglycosidase-assisted identification of intact glycopeptides. Anal Bioanal Chem 409, 3077–3087 (2017). https://doi.org/10.1007/s00216-017-0195-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0195-z