Abstract
Halophilic archaeal strain ZS-47-ST was isolated from Zhoushan marine solar saltern, China. Cells were pleomorphic, stained Gram-negative, and formed red-pigmented colonies on agar plate. Strain ZS-47-ST was able to grow at 20–50 °C (optimum 37 °C), at 0.9–4.8 M NaCl (optimum 3.4 M), at 0.005–1.0 M MgCl2 (optimum 0.03 M), and at pH 5.5–9.5 (optimum pH 6.5–7.5). The cells lysed in distilled water and the minimal NaCl concentration to prevent cell-lysis was 5% (w/v). The major polar lipids were C20C20 and C20C25 diether derivatives of phosphatidylglycerol, phosphatidylglycerol phosphate methyl ester, phosphatidylglycerol sulfate, glucosyl mannosyl glucosyl diether, and three unidentified glycolipids. The 16S rRNA gene and rpoB′ gene of strain ZS-47-ST were phylogenetically related to the corresponding genes of Halomarina oriensis JCM 16495T (98.57 and 92.94% similarities, respectively) and Halomarina salina CGMCC 1.12543T (97.96 and 93.65% similarities, respectively). The DNA G + C content of strain ZS-47-ST was 64.6 mol % (T m). The phenotypic, chemotaxonomic, and phylogenetic properties suggested that strain ZS-47-ST (=CGMCC 1.12563T = JCM 30037T) represents a new species of Halomarina, for which the name Halomarina rubra sp. nov. is proposed.

Similar content being viewed by others
References
Cui H-L, Lin Z-Y, Dong Y, Zhou P-J, Liu S-J (2007) Halorubrum litoreum sp. nov., an extremely halophilic archaeon from a solar saltern. Int J Syst Evol Microbiol 57:2204–2206
Cui H-L, Zhou P-J, Oren A, Liu S-J (2009) Intraspecific polymorphism of 16S rRNA genes in two halophilic archaeal genera, Haloarcula and Halomicrobium. Extremophiles 13:31–37
Cui H-L, Gao X, Yang X, Xu X-W (2010) Halorussus rarus gen. nov., sp. nov., a new member of the family Halobacteriaceae isolated from a marine solar saltern. Extremophiles 14:493–499
Dussault HP (1955) An improved technique for staining red halophilic bacteria. J Bacteriol 70:484–485
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Gonzalez C, Gutierrez C, Ramirez C (1978) Halobacterium vallismortis sp. nov. an amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can J Microbiol 24:710–715
Gupta RS, Naushad S, Baker S (2015) Phylogenomic analyses and molecular signatures for the class Halobacteria and its two major clades: a proposal for division of the class Halobacteria into an emended order Halobacteriales and two new orders, Haloferacales ord. nov. and Natrialbales ord. nov., containing the novel families Haloferacaceae fam. nov. and Natrialbaceae fam. nov. Int J Syst Evol Microbiol 65:1050–1069
Gupta RS, Naushad S, Fabros R, Adeolu M (2016) A phylogenomic reappraisal of family-level divisions within the class Halobacteria: proposal to divide the order Halobacteriales into the families Halobacteriaceae, Haloarculaceae fam. nov., and Halococcaceae fam. nov., and the order Haloferacales into the families, Haloferacaceae and Halorubraceae fam nov. Antonie Van Leeuwenhoek 109:565–587
Gutiérrez C, González C (1972) Method for simultaneous detection of proteinase and esterase activities in extremely halophilic bacteria. Appl Microbiol 24:516–517
Inoue K, Itoh T, Ohkuma M, Kogure K (2011) Halomarina oriensis gen. nov., sp. nov., a halophilic archaeon isolated from a seawater aquarium. Int J Syst Evol Microbiol 61:942–946
Ivanov VM (2004) The 125th anniversary of the Griess reagent. J Anal Chem 59:1002–1005
Kim M, Oh H-S, Park S-C, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Marmur J, Doty P (1962) Determination of the base composition of deoxyribonucleic acid from its thermal denaturation temperature. J Mol Biol 5:109–118
McDade JJ, Weaver RH (1959) Rapid methods for the detection of gelatin hydrolysis. J Bacteriol 77:60–64
Minegishi H, Kamekura M, Itoh T, Echigo A, Usami R, Hashimoto T (2010) Further refinement of the phylogeny of the Halobacteriaceae based on the full-length RNA polymerase subunit B′ (rpoB′) gene. Int J Syst Evol Microbiol 60:2398–2408
Oren A (2014) Taxonomy of halophilic Archaea: current status and future challenges. Extremophiles 18:825–834
Oren A, Ventosa A, Grant WD (1997) Proposed minimal standards for description of new taxa in the order Halobacteriales. Int J Syst Bacteriol 47:233–238
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Xu W-M, Xu J-Q, Zhou Y, Li Y, Lü Z-Z, Hou J, Zhu L, Cui H-L (2016) Halomarina salina sp. nov., isolated from a marine solar saltern. Antonie Van Leeuwenhoek 109:1121–1126
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31370054) and the 11th “Six Talents Peak” Project of Jiangsu Province (No. 2014-SWYY-021).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
The Digital Protologue database Taxon Number for strain ZS-47-ST is TA00228. The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene and rpoB ′ gene sequences of strain ZS-47-ST are KC918830 and MF358965, respectively.
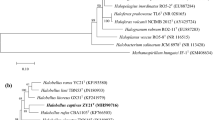
A phase-contrast micrograph of strain ZS-47-ST, thin-layer chromatograms of polar lipids extracted from strain ZS-47-ST, maximum-parsimony, and neighbour-joining phylogenetic tree reconstructions based on 16S rRNA gene and rpoB′ gene sequences showing the relationships between strain ZS-47-ST and related members within the class Halobacteria are available as supplementary materials.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, Y., Li, Y., Lü, ZZ. et al. Halomarina rubra sp. nov., isolated from a marine solar saltern. Arch Microbiol 199, 1431–1435 (2017). https://doi.org/10.1007/s00203-017-1420-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-017-1420-z