Abstract
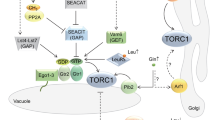
The major signaling pathway that regulates cell growth and metabolism is under the control of the target of rapamycin complex 1 (TORC1). In Saccharomyces cerevisiae the SEA complex is one of the TORC1 upstream regulators involved in amino acid sensing and autophagy. Here, we performed analysis of the expression, interactions and localization of SEA complex proteins under different conditions, varying parameters such as sugar source, nitrogen availability and growth phase. Our results show that the SEA complex promotes mitochondria degradation either by mitophagy or by general autophagy. In addition, the SEACIT subcomplex is involved in the maintenance of the vacuole–mitochondria contact sites. Thus, the SEA complex appears to be an important link between the TORC1 pathway and regulation of mitochondria quality control.







Similar content being viewed by others
Abbreviations
- ROS:
-
Reactive oxygen species
- SEACAT:
-
SEA subcomplex activating TORC1
- SEACIT:
-
SEA subcomplex inhibiting TORC1
- TORC1:
-
Target of rapamycin complex 1
- vCLAMPs:
-
Vacuole–mitochondria contact sites
References
Loewith R, Hall MN (2011) Target of rapamycin (TOR) in nutrient signaling and growth control. Genetics 189(4):1177–1201. https://doi.org/10.1534/genetics.111.133363
Laplante M, Sabatini DM (2012) mTOR signaling in growth control and disease. Cell 149(2):274–293. https://doi.org/10.1016/j.cell.2012.03.017
Huang K, Fingar DC (2014) Growing knowledge of the mTOR signaling network. Semin Cell Dev Biol 36:79–90. https://doi.org/10.1016/j.semcdb.2014.09.011
Ruetenik A, Barrientos A (2015) Dietary restriction, mitochondrial function and aging: from yeast to humans. Biochem Biophys Acta 1847 11:1434–1447. https://doi.org/10.1016/j.bbabio.2015.05.005
Groenewoud MJ, Zwartkruis FJ (2013) Rheb and mammalian target of rapamycin in mitochondrial homoeostasis. Open Biol 3(12):130185. https://doi.org/10.1098/rsob.130185
Li M, Zhao L, Liu J, Liu A, Jia C, Ma D, Jiang Y, Bai X (2010) Multi-mechanisms are involved in reactive oxygen species regulation of mTORC1 signaling. Cell Signal 22(10):1469–1476. https://doi.org/10.1016/j.cellsig.2010.05.015
Kawai S, Urban J, Piccolis M, Panchaud N, De Virgilio C, Loewith R (2011) Mitochondrial genomic dysfunction causes dephosphorylation of Sch9 in the yeast Saccharomyces cerevisiae. Eukaryot Cell 10(10):1367–1369. https://doi.org/10.1128/EC.05157-11
Jazwinski SM, Kriete A (2012) The yeast retrograde response as a model of intracellular signaling of mitochondrial dysfunction. Front Physiol 3:139. https://doi.org/10.3389/fphys.2012.00139
Dokudovskaya S, Waharte F, Schlessinger A, Pieper U, Devos DP, Cristea IM, Williams R, Salamero J, Chait BT, Sali A, Field MC, Rout MP, Dargemont C (2011) A conserved coatomer-related complex containing Sec13 and Seh1 dynamically associates with the vacuole in Saccharomyces cerevisiae. Mol Cell Proteomics 10(6):M110006478. https://doi.org/10.1074/mcp.m110.006478
Dokudovskaya S, Rout MP (2011) A novel coatomer-related SEA complex dynamically associates with the vacuole in yeast and is implicated in the response to nitrogen starvation. Autophagy 7(11):1392–1393. https://doi.org/10.4161/auto.7.11.17347
Bar-Peled L, Chantranupong L, Cherniack AD, Chen WW, Ottina KA, Grabiner BC, Spear ED, Carter SL, Meyerson M, Sabatini DM (2013) A Tumor suppressor complex with GAP activity for the Rag GTPases that signal amino acid sufficiency to mTORC1. Science 340(6136):1100–1106. https://doi.org/10.1126/science.1232044
Panchaud N, Peli-Gulli MP, De Virgilio C (2013) SEACing the GAP that nEGOCiates TORC1 activation: evolutionary conservation of Rag GTPase regulation. Cell Cycle 12(18):1–5. https://doi.org/10.4161/cc.26000
Dokudovskaya S, Rout MP (2015) SEA you later alli-GATOR—a dynamic regulator of the TORC1 stress response pathway. J Cell Sci 128(12):2219–2228. https://doi.org/10.1242/jcs.168922
Bar-Peled L, Sabatini DM (2014) Regulation of mTORC1 by amino acids. Trends Cell Biol 24(7):400–406. https://doi.org/10.1016/j.tcb.2014.03.003
Powis K, De Virgilio C (2016) Conserved regulators of Rag GTPases orchestrate amino acid-dependent TORC1 signaling. Cell Discov 2:15049. https://doi.org/10.1038/celldisc.2015.49
Costanzo M, Baryshnikova A, Bellay J, Kim Y, Spear ED, Sevier CS, Ding H, Koh JL, Toufighi K, Mostafavi S, Prinz J, St Onge RP, VanderSluis B, Makhnevych T, Vizeacoumar FJ, Alizadeh S, Bahr S, Brost RL, Chen Y, Cokol M, Deshpande R, Li Z, Lin ZY, Liang W, Marback M, Paw J, San Luis BJ, Shuteriqi E, Tong AH, van Dyk N, Wallace IM, Whitney JA, Weirauch MT, Zhong G, Zhu H, Houry WA, Brudno M, Ragibizadeh S, Papp B, Pal C, Roth FP, Giaever G, Nislow C, Troyanskaya OG, Bussey H, Bader GD, Gingras AC, Morris QD, Kim PM, Kaiser CA, Myers CL, Andrews BJ, Boone C (2010) The genetic landscape of a cell. Science 327(5964):425–431. https://doi.org/10.1126/science.1180823
Costanzo M, VanderSluis B, Koch EN, Baryshnikova A, Pons C, Tan G, Wang W, Usaj M, Hanchard J, Lee SD, Pelechano V, Styles EB, Billmann M, van Leeuwen J, van Dyk N, Lin ZY, Kuzmin E, Nelson J, Piotrowski JS, Srikumar T, Bahr S, Chen Y, Deshpande R, Kurat CF, Li SC, Li Z, Usaj MM, Okada H, Pascoe N, San Luis BJ, Sharifpoor S, Shuteriqi E, Simpkins SW, Snider J, Suresh HG, Tan Y, Zhu H, Malod-Dognin N, Janjic V, Przulj N, Troyanskaya OG, Stagljar I, Xia T, Ohya Y, Gingras AC, Raught B, Boutros M, Steinmetz LM, Moore CL, Rosebrock AP, Caudy AA, Myers CL, Andrews B, Boone C (2016) A global genetic interaction network maps a wiring diagram of cellular function. Science. https://doi.org/10.1126/science.aaf1420
Algret R, Fernandez-Martinez J, Shi Y, Kim SJ, Pellarin R, Cimermancic P, Cochet E, Sali A, Chait BT, Rout MP, Dokudovskaya S (2014) Molecular architecture and function of the SEA complex, a modulator of the TORC1 pathway. Mol Cell Proteomics 13(11):2855–2870. https://doi.org/10.1074/mcp.M114.039388
Elbaz-Alon Y, Rosenfeld-Gur E, Shinder V, Futerman AH, Geiger T, Schuldiner M (2014) A dynamic interface between vacuoles and mitochondria in yeast. Dev Cell 30(1):95–102. https://doi.org/10.1016/j.devcel.2014.06.007
Merz S, Westermann B (2009) Genome-wide deletion mutant analysis reveals genes required for respiratory growth, mitochondrial genome maintenance and mitochondrial protein synthesis in Saccharomyces cerevisiae. Genome Biol 10(9):R95. https://doi.org/10.1186/gb-2009-10-9-r95
Honscher C, Mari M, Auffarth K, Bohnert M, Griffith J, Geerts W, van der Laan M, Cabrera M, Reggiori F, Ungermann C (2014) Cellular metabolism regulates contact sites between vacuoles and mitochondria. Dev Cell 30(1):86–94. https://doi.org/10.1016/j.devcel.2014.06.006
Alber F, Dokudovskaya S, Veenhoff LM, Zhang W, Kipper J, Devos D, Suprapto A, Karni-Schmidt O, Williams R, Chait BT, Rout MP, Sali A (2007) Determining the architectures of macromolecular assemblies. Nature 450(7170):683–694. https://doi.org/10.1038/nature06404
Neklesa TK, Davis RW (2009) A genome-wide screen for regulators of TORC1 in response to amino acid starvation reveals a conserved Npr2/3 complex. PLoS Genet 5(6):e1000515. https://doi.org/10.1371/journal.pgen.1000515
LaCava J, Molloy KR, Taylor MS, Domanski M, Chait BT, Rout MP (2015) Affinity proteomics to study endogenous protein complexes: pointers, pitfalls, preferences and perspectives. Biotechniques 58(3):103–119. https://doi.org/10.2144/000114262
Brosch M, Yu L, Hubbard T, Choudhary J (2009) Accurate and sensitive peptide identification with Mascot Percolator. J Proteome Res 8(6):3176–3181. https://doi.org/10.1021/pr800982s
Kurihara Y, Kanki T, Aoki Y, Hirota Y, Saigusa T, Uchiumi T, Kang D (2012) Mitophagy plays an essential role in reducing mitochondrial production of reactive oxygen species and mutation of mitochondrial DNA by maintaining mitochondrial quantity and quality in yeast. J Biol Chem 287(5):3265–3272. https://doi.org/10.1074/jbc.M111.280156
Wu X, Tu BP (2011) Selective regulation of autophagy by the Iml1-Npr2-Npr3 complex in the absence of nitrogen starvation. Mol Biol Cell 22(21):4124–4133. https://doi.org/10.1091/mbc.E11-06-0525
Chong YT, Koh JL, Friesen H, Duffy SK, Cox MJ, Moses A, Moffat J, Boone C, Andrews BJ (2015) Yeast proteome dynamics from single cell imaging and automated analysis. Cell 161(6):1413–1424. https://doi.org/10.1016/j.cell.2015.04.051
Huh WK, Falvo JV, Gerke LC, Carroll AS, Howson RW, Weissman JS, O’Shea EK (2003) Global analysis of protein localization in budding yeast. Nature 425(6959):686–691. https://doi.org/10.1038/nature02026nature02026
Gonzalez Montoro A, Auffarth K, Honscher C, Bohnert M, Becker T, Warscheid B, Reggiori F, van der Laan M, Frohlich F, Ungermann C (2018) Vps39 interacts with Tom40 to establish one of two functionally distinct vacuole-mitochondria contact sites. Dev cell 45(5):621-636 e627. https://doi.org/10.1016/j.devcel.2018.05.011
Lang AB, John Peter AT, Walter P, Kornmann B (2015) ER-mitochondrial junctions can be bypassed by dominant mutations in the endosomal protein Vps13. J Cell Biol 210(6):883–890. https://doi.org/10.1083/jcb.201502105
John Peter AT, Herrmann B, Antunes D, Rapaport D, Dimmer KS, Kornmann B (2017) Vps13-Mcp1 interact at vacuole-mitochondria interfaces and bypass ER-mitochondria contact sites. J Cell Biol. https://doi.org/10.1083/jcb.201610055
Binda M, Peli-Gulli MP, Bonfils G, Panchaud N, Urban J, Sturgill TW, Loewith R, De Virgilio C (2009) The Vam6 GEF controls TORC1 by activating the EGO complex. Mol Cell 35(5):563–573. https://doi.org/10.1016/j.molcel.2009.06.033
Kuzmin E, VanderSluis B, Wang W, Tan G, Deshpande R, Chen Y, Usaj M, Balint A, Mattiazzi Usaj M, van Leeuwen J, Koch EN, Pons C, Dagilis AJ, Pryszlak M, Wang JZY, Hanchard J, Riggi M, Xu K, Heydari H, San Luis BJ, Shuteriqi E, Zhu H, Van Dyk N, Sharifpoor S, Costanzo M, Loewith R, Caudy A, Bolnick D, Brown GW, Andrews BJ, Boone C, Myers CL (2018) Systematic analysis of complex genetic interactions. Science. https://doi.org/10.1126/science.aao1729
van Leeuwen J, Pons C, Mellor JC, Yamaguchi TN, Friesen H, Koschwanez J, Usaj MM, Pechlaner M, Takar M, Usaj M, VanderSluis B, Andrusiak K, Bansal P, Baryshnikova A, Boone CE, Cao J, Cote A, Gebbia M, Horecka G, Horecka I, Kuzmin E, Legro N, Liang W, van Lieshout N, McNee M, San Luis BJ, Shaeri F, Shuteriqi E, Sun S, Yang L, Youn JY, Yuen M, Costanzo M, Gingras AC, Aloy P, Oostenbrink C, Murray A, Graham TR, Myers CL, Andrews BJ, Roth FP, Boone C (2016) Exploring genetic suppression interactions on a global scale. Science. https://doi.org/10.1126/science.aag0839
Usaj M, Tan Y, Wang W, VanderSluis B, Zou A, Myers CL, Costanzo M, Andrews B, Boone C (2017) TheCellMap.org: a web-accessible database for visualizing and mining the global yeast genetic interaction network. G3 (Bethesda) 7(5):1539–1549. https://doi.org/10.1534/g3.117.040220
Panchaud N, Peli-Gulli MP, De Virgilio C (2013) Amino acid deprivation inhibits TORC1 through a GTPase-activating protein complex for the Rag family GTPase Gtr1. Sci Signal 6(277):ra42. https://doi.org/10.1126/scisignal.2004112
Deffieu M, Bhatia-Kissova I, Salin B, Klionsky DJ, Pinson B, Manon S, Camougrand N (2013) Increased levels of reduced cytochrome b and mitophagy components are required to trigger nonspecific autophagy following induced mitochondrial dysfunction. J Cell Sci 126(Pt 2):415–426. https://doi.org/10.1242/jcs.103713
Lee J, Giordano S, Zhang J (2012) Autophagy, mitochondria and oxidative stress: cross-talk and redox signalling. Biochem J 441(2):523–540. https://doi.org/10.1042/BJ20111451
Pan Y, Schroeder EA, Ocampo A, Barrientos A, Shadel GS (2011) Regulation of yeast chronological life span by TORC1 via adaptive mitochondrial ROS signaling. Cell Metab 13(6):668–678. https://doi.org/10.1016/j.cmet.2011.03.018
Graef M, Nunnari J (2011) Mitochondria regulate autophagy by conserved signalling pathways. EMBO J 30(11):2101–2114. https://doi.org/10.1038/emboj.2011.104
Sutter BM, Wu X, Laxman S, Tu BP (2013) Methionine inhibits autophagy and promotes growth by inducing the SAM-responsive methylation of PP2A. Cell 154(2):403–415. https://doi.org/10.1016/j.cell.2013.06.041
Kira S, Tabata K, Shirahama-Noda K, Nozoe A, Yoshimori T, Noda T (2014) Reciprocal conversion of Gtr1 and Gtr2 nucleotide-binding states by Npr2-Npr3 inactivates TORC1 and induces autophagy. Autophagy 10(9):1565–1578. https://doi.org/10.4161/auto.29397
Laxman S, Sutter BM, Shi L, Tu BP (2014) Npr2 inhibits TORC1 to prevent inappropriate utilization of glutamine for biosynthesis of nitrogen-containing metabolites. Sci Signal 7(356):ra120. https://doi.org/10.1126/scisignal.2005948
Laxman S, Sutter BM, Wu X, Kumar S, Guo X, Trudgian DC, Mirzaei H, Tu BP (2013) Sulfur amino acids regulate translational capacity and metabolic homeostasis through modulation of tRNA thiolation. Cell 154(2):416–429. https://doi.org/10.1016/j.cell.2013.06.043
Klionsky DJ (2016) Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12(1):1–222. https://doi.org/10.1080/15548627.2015.1100356
Abeliovich H, Zarei M, Rigbolt KT, Youle RJ, Dengjel J (2013) Involvement of mitochondrial dynamics in the segregation of mitochondrial matrix proteins during stationary phase mitophagy. Nat Commun 4:2789. https://doi.org/10.1038/ncomms3789
Sugiura A, McLelland GL, Fon EA, McBride HM (2014) A new pathway for mitochondrial quality control: mitochondrial-derived vesicles. EMBO J 33(19):2142–2156. https://doi.org/10.15252/embj.201488104
Kissova I, Salin B, Schaeffer J, Bhatia S, Manon S, Camougrand N (2007) Selective and non-selective autophagic degradation of mitochondria in yeast. Autophagy 3(4):329–336
Wei Y, Chiang WC, Sumpter R Jr, Mishra P, Levine B (2017) Prohibitin 2 is an inner mitochondrial membrane mitophagy receptor. Cell 168(1–2):224-238 e210. https://doi.org/10.1016/j.cell.2016.11.042
Liu Y, Okamoto K (2018) The TORC1 signaling pathway regulates respiration-induced mitophagy in yeast. Biochem Biophys Res Commun 502(1):76–83. https://doi.org/10.1016/j.bbrc.2018.05.123
Okamoto K, Kondo-Okamoto N, Ohsumi Y (2009) Mitochondria- anchored receptor Atg32 mediates degradation of mitochondria via selective autophagy. Dev Cell 17(1):87–97. https://doi.org/10.1016/j.devcel.2009.06.013
Kanki T, Wang K, Baba M, Bartholomew CR, Lynch-Day MA, Du Z, Geng J, Mao K, Yang Z, Yen WL, Klionsky DJ (2009) A genomic screen for yeast mutants defective in selective mitochondria autophagy. Mol Biol Cell 20(22):4730–4738. https://doi.org/10.1091/mbc.E09-03-0225
Muller M, Kotter P, Behrendt C, Walter E, Scheckhuber CQ, Entian KD, Reichert AS (2015) Synthetic quantitative array technology identifies the Ubp3-Bre5 deubiquitinase complex as a negative regulator of mitophagy. Cell Rep 10(7):1215–1225. https://doi.org/10.1016/j.celrep.2015.01.044
Mao K, Wang K, Zhao M, Xu T, Klionsky DJ (2011) Two MAPK-signaling pathways are required for mitophagy in Saccharomyces cerevisiae. J Cell Biol 193(4):755–767. https://doi.org/10.1083/jcb.201102092
Algret R, Dokudovskaya S (2012) The SEA complex—the beginning. Biopolym Cell 28(N4):281–284
Li F, Wang F, Haraldson K, Protopopov A, Duh FM, Geil L, Kuzmin I, Minna JD, Stanbridge E, Braga E, Kashuba VI, Klein G, Lerman MI, Zabarovsky ER (2004) Functional characterization of the candidate tumor suppressor gene NPRL2/G21 located in 3p21.3C. Cancer Res 64(18):6438–6443
Luc R, Tortorella SM, Ververis K, Karagiannis TC (2015) Lactate as an insidious metabolite due to the Warburg effect. Mol Biol Rep 42(4):835–840. https://doi.org/10.1007/s11033-015-3859-9
Acknowledgements
We are grateful to Jaclyn Tetenbaum-Novatt and Renaud Legouis for critical reading of manuscript. We thank Sebastien Leon, Christian Ungermann, Benedikt Westermann and Claudio de Virgilio for sharing plasmids and antibodies.
Funding
SD acknowledge financial support from La Ligue contre le Cancer (Comité de Paris/Ile-de-France and Comité d’Oise), PICS USA (CNRS); YM is grateful to the support from Chinese Scholarship Council (CSC). We thank Proteomic Core Facility (support from grants TA2013 and TA2016), Imaging and Cytometry Facility (PFIC, UMS CNRS 3655—INSERM US23, support from grant TA2017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ma, Y., Moors, A., Camougrand, N. et al. The SEACIT complex is involved in the maintenance of vacuole–mitochondria contact sites and controls mitophagy. Cell. Mol. Life Sci. 76, 1623–1640 (2019). https://doi.org/10.1007/s00018-019-03015-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-019-03015-6