Abstract
Objective and design
The heterogeneity of response to SARS-CoV-2 infection is directly linked to the individual genetic background. Genetic variants of inflammasome-related genes have been pointed as risk factors for several inflammatory sterile and infectious disease. In the group of inflammasome receptors, NLRP1 stands out as a good novel candidate as severity factor for COVID-19 disease.
Methods
To address this question, we performed an association study of NLRP1, DPP9, CARD8, IL1B, and IL18 single nucleotide variants (SNVs) in a cohort of 945 COVID-19 patients.
Results
The NLRP1 p.Leu155His in the linker region, target of viral protease, was significantly associated to COVID-19 severity, which could contribute to the excessive cytokine release reported in severe cases.
Conclusion
Inflammasome genetic background contributes to individual response to SARS-CoV-2.
Similar content being viewed by others
Introduction
The individual response to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection is considerably variable in the general population, ranging from a flu-like disease to a severe acute respiratory syndrome. High mortality was associated with old age and male sex [1, 2], hypertension and diabetes [3].
SARS-CoV-2 infects airway and lung epithelial cells, and innate receptors may detect it and control its replication or inducing excessive inflammation and broad activation of neighboring cells, including leukocytes, platelets, and endothelium, therefore leading to endotheliitis and diffuse thromboembolism, complications observed in severe patients [3, 4].
Half a dozen genetic loci have been identified as possible risk factors for severe COVID-19 disease, including genes involved in virus-associated molecular patterns detection and in the type 1 interferon pathway (as recently reviewed in [5]).
The cytosolic complex inflammasome comprises an important response to viral infection, as it is responsible for the activation of caspase-1 and the subsequent cleavage and release of the two cytokines IL-1ß and IL-18, which are directly involved in antiviral innate response, and, indirectly, by inducing cell-mediated adaptive response [6]. Increased plasma level of IL-18, but not IL-1ß, was observed in all COVID-19 patients, with the highest titers in severe ones. On the other hand, IL-1ß is upregulated in bronchoalveolar lavage, and both cytokines are highly produced by lung macrophages, suggesting that inflammasome is greatly induced in lungs during SARS-CoV-2 infection [7].
Moreover, gasdermin-D (GSDMD), another substrate of caspase-1, and responsible for the inflammatory lithic cell death known as pyroptosis, has been shown to worsen the prognosis of COVID-19 [8].
How SARS-CoV-2 activates the inflammasome is not yet fully understood. Recent literature supports the hypothesis that the NACHT and LRRs containing receptor with a PYD domain (NLRP) 1 acts as a broad viral proteases’ sensor and decoy factor [9, 10]. NLRP1 has been shown to be the target of some viral proteases, including the picornavirus and enterovirus 3C protease [9, 10], and also the SARS-CoV-2 3CL protease NSP5 [11]. Viral proteases cleave the amino-terminal portion of the NLRP1 within a “tripwire” region localized in the link sequence between the PYD and NACHT domains (Fig. 1). The cleavage leads to proteasomal digestion of the amino-terminal portion of NLRP1 and to the activation of the inflammasome [9, 10, 11]. In a previous activation model, NLRP1 was assumed to suffer self-cleavage in the FIIND domain (at Ser1312), resulting in the dissociation of the endogenous inhibitor dipeptidyl peptidase (DPP)-9 and proteasomal degradation of the amino-terminal portion of the receptor [12]. Whether this mechanism is involved also in the activation of NLRP1 by viral proteases has not been yet elucidated.
Of note, several gain-of-function missense variants and sites of positive selection are present in the NLRP1 linker region, likely related to past events of virus-mediated selective pressure [13].
All this considering, the NLRP1 inflammasome represents a good candidate as a severity factor in the COVID-19 disease. In an attempt to verify whether common variants in the NLRP1 gene may affect host response to the SARS-CoV-2 virus, we performed an association study by analyzing the frequency distribution of eight well-known functional single nucleotide variants (SNVs) in a cohort of Brazilian COVID-19 patients. We have selected the two missense variants in NLRP1 p.Leu155His (rs12150220) and p.Met1184Val (rs11651270) which are known to increase IL-1ß processing in peripheral blood mononuclear cells [14], and their high frequency in the general population is hypothesized to be the consequence of a selective advantage against a viral pandemic [13]. Moreover, the nonsense variant p.Cys10Ter in CARD8 (rs2043211), another recently recognized viral proteases target [15], and whose activation is regulated by the interaction with DPP9 like NLRP1 [16], was also analyzed. Finally, taking in account that the inflammasome is a complex, and that SNV–SNV or gene–gene interaction are expected, we included in the study also the promoter enhancer variant rs16944 in IL1B, two eQTL SNVs in IL18 (rs5744256 and rs1834481), known to affect the cytokines’ level [17, 18, 19], and two SNVs (rs12610495 and rs8101703) with opposite eQTL effect in the DPP9 gene, as a possible upstream regulator of NLRP1 and CARD8 [16].
Domain architecture is specified for the NLRP1 and CARD8 proteins. The common mechanism of endogenous inhibition of the sensors is mediated by the interaction between dipeptidyl peptidase (DPP)-9 and the FIIND domain. DPP-9 hides the auto-cleavage site within the FIIND domain of NLRP1 and CARD8 (Ser1213 and Ser297, respectively). Viral proteases’ digestion of the N-terminal regions of NLRP1 (“tripwire” linker sequence is evidenced) and CARD8 has been shown to be sufficient to induce the mounting and activation of the inflammasome and the consequent cleavage of its substrates pro-IL-1ß, pro-IL-18, and GSDMD, into their biologically active forms. Functional variants selected for this study are indicated in red: p.Leu155His (L155H; rs12150220), p.Met1184Val (M1184V; rs11651270), and p.Cys10ter (C10X; rs2043211).
Patients and methods
Study participants
Nine hundred and forty five patients (n = 945; median age: 47; range: 19–95; males: 51%) positive for polymerase chain reaction (PCR) test for RNA of SARS-CoV-2 in nasopharyngeal swabs were recruited in three Brazilian hospitals: “Hospital das Clinicas/Faculdade de Medicina da Universidade de São Paulo” (HC/FMUSP) and “Hospital Albert Einstein” in São Paulo (state of São Paulo) (n = 249 and 396, respectively), and “Hospital Universitario/Universidade Federal de Mato Grosso do Sul” (HU/UFMS) in Campo Grande (state of Mato Grosso do Sul) (n = 300), between 2020 and 2021, before the availability of the vaccine. Two hundred and eighty subjects (n = 280; 30%) are patients with severe COVID-19 (cases), which is defined as hospitalization with respiratory failure and in need of intensive care and mechanical ventilation. Six hundred and sixty-five participants (n = 665; 70%) with a mild flu-like disease were classified as paucisymptomatic patients (controls). Hypertension and diabetes are present in 43 and 34% of the recruited subjects, respectively (Table 1). The study was approved by the Ethics Committee of each of the hospitals (protocols CAAE: 30,800,520.7.0000.0068 2020; 41,492,620.1.0000.0071; 32,466,020.8.0000.0021). Written informed consent was obtained, sometimes in a delayed fashion, from the patients at each center when possible.
Sample processing and genotyping
The extraction of DNA from peripheral blood was carried out using the “Salting Out” technique [20]. Eight functional SNVs were selected in the NLRP1 (rs12150220, rs11651270), CARD8 (rs2043211), IL1B (rs16944), IL18 (rs1834481, rs5744256), and DPP9 (rs12610495, rs8101703) genes. Their minor allele frequency (MAF) in the general population is greater than 10%. (Table 2) The genotyping of the selected SNVs was performed by the use of allele-specific Taqman® assays (Applied Biosystems, Thermo Fisher Scientific) and qPCR in a QuantStudio real-time PCR equipment (Applied Biosystems, Thermo Fisher Scientific).
Statistical analysis
First, a raw analysis comparing the allelic distribution of the SNVs in the case/control groups was performed, and then a multivariate analysis for the genotypic distribution including variables previously described to affect the disease outcome [1, 2, 21]: age and sex (main analysis), hypertension and diabetes (corrected analysis) was performed. SNPassoc, an R package (http://www.r-project.org), was used for the analysis. Haploview software was used to derive the haplotypes. The commonly accepted threshold of 0.05/number of independent SNVs (Bonferroni correction for multiple comparisons) was implied to determine statistical significant differences (p < 0.01).
Results
The aim of this study was to evaluate whether common variants in NLRP1 inflammasome may contribute to COVID-19 severity. To this end, we recruited patients proceeding from one public and one private hospitals in São Paulo (capital, state of São Paulo), and one public hospital in Campo Grande (capital, state of Mato Grosso do Sul) between 2020 and 2021, before the availability of the vaccine. Demographic and main clinical data of recruited individuals are resumed in Table 1. The proportion of severe patients in each cohort is different and it does not represent the proportion of severe COVID-19 patients in the total number of SARS-CoV-2 infections. Our cohort is enriched of severe patients due to the statistical power calculation and to ensure enough strength of the case/control association analysis. We selected known functional SNVs in the NLRP1 gene (rs12150220 and rs11651270), the main focus of this study, but also in the genes of the two cytokines, IL1B (rs16944) and IL18 (rs5744256 and rs1834481), downstream effectors of the inflammasome activation, and in the DPP9 gene (rs12610495 and rs8101703), as a possible upstream regulator of NLRP1. The nonsense SNV rs2043211 in the CARD8 gene, was also included in the study due to the similarity of the proposed activation mechanism for CARD8 and NLRP1, including the viral protease cleavage and the interaction with DPP-9 [16].
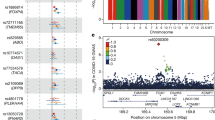
In the raw analysis, we identified three SNVs significantly associated with COVID-19 severity. The minor alleles of NLRP1 rs12150220 and CARD8 rs2043211 variants were more frequent in severe than in paucisymptomatic patients (odds ratio: 1.36 for the minor allele; 95% confidence intervals: 1.12–1.65 and 1.11–1.69, respectively). Moreover, the IL18 rs5744256 variant was significantly associated with the protection of SARS-CoV-2 infected individuals to develop severe disease (O.R.: 0.66 for the minor allele: 95% C.I.: 0.51–0.87) (Table 3).
The SNVs in the IL18 and DPP9 genes formed two haplo-blocks with a strong linkage disequilibrium (LD) (D’ > 90); while SNVs in the NLRP1 gene resulted in medium LD (D’ = 65) in our cohort. (Supplementary File 1). Raw haplotypes’ analysis revealed that IL18 G-G haplotype was more frequent in paucisymptomatic than in severe COVID-19 patients (p = 0.006) (Supplementary File 2), corroborating the single allele association result.
Main analysis included origin, sex, and age as covariates and it was performed according to dominant and recessive models of inheritance for all the SNVs (Table 4). The NLRP1 rs12150220 T/T genotype resulted more frequent in severe than in mild patients according to a recessive model of inheritance for the minor allele (O.R.: 2.42; 95% C.I.: 1.44–4.08).
When hypertension and diabetes were added as covariates in our analysis, the association of the NLRP1 rs12150220 T/T genotype with COVID-19 severity was maintained (O.R.: 2.35; 95% C.I.: 1.39–3.99). Moreover, the IL18 rs5744256 G-containing genotypes resulted significantly less frequently in severe than in mild patients according to a dominant model of inheritance for the minor allele (O.R.: 0.60; 95% C.I.: 0.41–0.89). (Table 4).
Given that the selected genes belong to the same biologic pathway, the NLRP1 inflammasome, we then realized an interaction analysis between SNVs localized in distinct genes (NLRP1 and IL1B, or IL18, or DPP9). Possibly due to the relatively low frequency of rs5744256 variant (16%), the combination between rs12150220 and rs5744256 genotypes resulted in a not significant association after Bonferroni correction (p = 0.040).
Discussion
Using a candidate SNV approach, based on known common functional variants, and the cooperation of teams of three Brazilian hospitals, we performed an association study to evaluate the hypothesis that the newly identified cytosolic receptor of SARS-CoV-2, NLRP1 could represent a susceptibility gene for COVID-19 severity, in terms of respiratory failure and ICU internalization.
We detected a significant association of NLRP1 p.Leu155His missense variant (rs12150220) and severe COVID-19. This SNV is localized in the linker region between amino-terminal PYD and central NACHT domain, very close to the so-called “tripwire” region (a.a. 127–134), which is the target of viral proteases [10], including the Coronavirus 3CL protease NSP5 [11]. The known functional effect of p.Leu155His regards the increased activation of NLRP1 inflammasome and IL-1ß release in peripheral blood mononuclear cells [14]. Far from recent discoveries, Vasseur and colleagues have hypothesized that the high frequency of rs12150220 in the European population (about 50%) may suggest a rapid evolution in response to a viral epidemic [13], reinforcing the idea that this specific SNV could be related to the host antiviral response.
Intriguingly, the other missense variant in NLRP1, the p.Met1184Val substitution (rs11651270), which also had been identified in the study of Vasseur et al. [13] as target of a rapid positive selective pressure together with rs12150220, did not associate with the severity of COVID-19, at least in our cohort. It is interesting to emphasize, however, that p.Met1184Val is localized within the FIIND domain, not a direct target of SARS-CoV-2 protease, supporting a specific effect of rs12150220 in COVID-19.
We have included in the study also the nonsense variant CARD8 rs2043211, due to the structural similarity of CARD8 and NLRP1, and the recent findings about the role of CARD8 as cytosolic target of HIV-1 protease [15]. However, this variant in the CARD8 gene did not result associated to COVID-19 disease. Of note, the two variants in the DPP9 gene did not associate with the disease, once more supporting the idea that, in this specific context of SARS-CoV-2 infection, the “tripwire” region is more determinant for NLRP1 activation than the DPP-9/FIIND mechanism.
As such, it is reasonable to conclude that NLRP1 p.Leu155His variant can be specifically involved in COVID-19 severity. We hypothesized that this polymorphism could affect the secondary structure of the “tripwire” sequence leading to increased accessibility to viral proteases and consequently to augment the activation of NLRP1 inflammasome. When we performed an “in silico” analysis using the “PredictProtein” online software (https://predictprotein.org/), we observed that the amino acid substitution Leu- > His modifies the predicted secondary structure of the N-terminal portion of NLRP1. A helix structure is lost in the “tripwire” region (a.a. 127–134) and the accessibility of the sequence (exposed versus buried residues) also is affected (Supplementary File 3), suggesting that the rs12150220 variant can result in a NLRP1 N-terminal region prone to proteases’ digestion.
It is important to note that the association resulted significant in all the three steps of analysis, even when well-known risk variables, age, sex, hypertension, and diabetes, have been included in the multivariate analysis. As NLRP1 has been previously associated to Addison disease [22], and type-1 diabetes [23, 24], we performed an association analysis using diabetes as main variable, instead of a covariate, however, at least in our cohort, none of the studied SNVs resulted associated to diabetes. The same result was obtained when hypertension was used as main variable, finally excluding a possible effect of rs12150220 on these two conditions in our cohort, and emphasizing the robustness of the main association analysis. The meaning of the association of NLRP1 p.Leu155His with COVID-19 severity may be related to excessive cytokines release, and literature have yet showed that both IL-1ß and IL-18 are dysregulated during SARS-CoV-2 response and in severe patients [8]. Although we expected to find a relationship between the SNVs in NLRP1 and IL1B and/or IL18 genes, we did not observe any association between rs12150220 and the well-known variant -511 C>T (rs16944) in the promoter of the IL1B gene, which is related to the increase in expression and release of the cytokine [17]. On the other hand, the variants in the IL18 gene, and in particular, the rs5744256 SNV, were inversely associated with the severity of the COVID-19. This intronic variant has been described as an eQTL for reduced plasma IL-18 concentration [18], suggesting here that lower cytokine levels might be beneficial in severe disease. This result appears to be in contrast with the common view of IL-18 as an anti-viral factor [6] and with the elevated levels of IL-18 observed in SARS-CoV-2 infection [8]. However we can hypothesize that the role of the cytokine may or not be protective depending on the time of the disease. As genetic background has been pointed out as a risk factor for the development of severe COVID-19 disease [6], our findings added another piece of the complex puzzle of the individual response to SARS-CoV-2, and at the same time confirm the key role of the NLRP1 sensor in antiviral response. Fundings: This study was funded by “Fundação de Apoio a Pesquisa do Estado de Sao Paulo” (FAPESP) process n.21/05420–4 (A.P.) and “Coordination for the Improvement of Higher Education Personnel” (CAPES) process number (n.) 88,887.503842/2020–00 (A.J.D.S.; M.N.S; A.P.), “Brazilian Ministry of Education, TED SESU/MEC nº 9233/2020” (J.V). Authors are supported by fellowship programs: Excellence in Research Fellowship from “National Council for Scientific and Technological Development” (CNPq) (A.P., M.N.S, A.J.S.D.); Postdoctoral Fellowship from São Paulo Research Foundation (FAPESP) n. 2021/13049–4 (V.N.C.L), PhD Fellowships n.2021/09161–3 (R.W.A), n. 2019/22448–0 (S.C.G.S), n. 2018/18230–6 (F.M.E.T); CAPES PhD Fellowships (n. 88,887.469122/2019–00; R.A.G.C); and Undergraduate Scholarship (SMY: PIBIC n.158842/2021–9).
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper or in the Supplementary Section.
References
Li X, Xu S, Yu M, Wang K, Tao Y, Zhou Y, Shi J, Zhou M, Wu B, Yang Z, Zhang C, Yue J, Zhang Z, Renz H, Liu X, Xie J, Xie M, Zhao J. Risk factors for severity and mortality in adult COVID-19 inpatients in Wuhan. J Allergy Clin Immunol. 2020;146:110–8.
Zhou F, Yu T, Du R, Fan G, Liu Y, Liu Z, Xiang J, Wang Y, Song B, Gu X, Guan L, Wei Y, Li H, Wu X, Xu J, Tu S, Zhang Y, Chen H, Cao B. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet (London, England). 2020;395:1054–62.
Liu J, Li S, Liu J, Liang B, Wang X, Wang H, Li W, Tong Q, Yi J, Zhao L, Xiong L, Guo C, Tian J, Luo J, Yao J, Pang R, Shen H, Peng C, Liu T, Zhang Q, Wu J, Xu L, Lu S, Wang B, Weng Z, Han C, Zhu H, Zhou R, Zhou H, Chen X, Ye P, Zhu B, Wang L, Zhou W, He S, He Y, Jie S, Wei P, Zhang J, Lu Y, Wang W, Zhang L, Li L, Zhou F, Wang J, Dittmer U, Lu M, Hu Y, Yang D, Zheng X. Longitudinal characteristics of lymphocyte responses and cytokine profiles in the peripheral blood of SARS-CoV-2 infected patients. EBioMedicine. 2020;55: 102763.
Ruhl L, Pink I, Kühne JF, Beushausen K, Keil J, Christoph S, Sauer A, Boblitz L, Schmidt J, David S, Jäck HM, Roth E, Cornberg M, Schulz TF, Welte T, Höper MM, Falk CS. Endothelial dysfunction contributes to severe COVID-19 in combination with dysregulated lymphocyte responses and cytokine networks. Signal Transduct Target Ther. 2021;6:418.
Zhang Q, Bastard P, Karbuz A, Gervais A, Tayoun AA, Aiuti A, Belot A, Bolze A, Gaudet A, Bondarenko A, Liu Z, Spaan AN, Guennoun A, Arias AA, Planas AM, Sediva A, Shcherbina A, Neehus A-L, Puel A, Froidure A, Novelli A, Parlakay AÖ, Pujol A, Yahşi A, Gülhan B, Bigio B, Boisson B, Drolet BA, Franco CAA, Flores C, Rodríguez-Gallego C, Prando C, Biggs CM, Luyt C-E, Dalgard CL, O’Farrelly C, Matuozzo D, Dalmau D, Perlin DS, Mansouri D, van de Beek D, Vinh DC, Dominguez-Garrido E, Hsieh EWY, Erdeniz EH, Jouanguy E, Şevketoglu E, Talouarn E, Quiros-Roldan E, Andreakos E, Husebye E, Alsohime F, Haerynck F, Casari G, Novelli G, Aytekin G, Morelle G, Alkan G, Bayhan GI, Feldman HB, Su HC, von Bernuth H, Resnick I, Bustos I, Meyts I, Migeotte I, Tancevski I, Bustamante J, Fellay J, El Baghdadi J, Martinez-Picado J, Casanova J-L, Rosain J, Manry J, Chen J, Christodoulou J, Bohlen J, Franco JL, Li J, Anaya JM, Rojas J, Ye J, Uddin KMF, Yasar KK, Kisand K, Okamoto K, Chaïbi K, Mironska K, Maródi L, Abel L, Renia L, Lorenzo L, Hammarström L, Ng LFP, Quintana-Murci L, Erazo LV, Notarangelo LD, Reyes LF, Allende LM, Imberti L, Renkilaraj MRLM, Moncada-Velez M, Materna M, Anderson MS, Gut M, Chbihi M, Ogishi M, Emiroglu M, Seppänen MRJ, Uddin MJ, Shahrooei M, Alexander N, Hatipoglu N, Marr N, Akçay N, Boyarchuk O, Slaby O, Akcan OM, Zhang P, Soler-Palacín P, Gregersen PK, Brodin P, Garçon P, Morange P-E, Pan-Hammarström Q, Zhou Q, Philippot Q, Halwani R, de Diego RP, Levy R, Yang R, Öz ŞKT, Muhsen SA, Kanık-Yüksek S, Espinosa-Padilla S, Ramaswamy S, Okada S, Bozdemir SE, Aytekin SE, Karabela ŞN, Keles S, Senoglu S, Zhang S-Y, Duvlis S, Constantinescu SN, Boisson-Dupuis S, Turvey SE, Tangye SG, Asano T, Ozcelik T, Le Voyer T, Maniatis T, Morio T, Mogensen TH, Sancho-Shimizu V, Beziat V, Solanich X, Bryceson Y, Lau Y-L, Itan Y, Cobat A, Casanova J-L, Effort CHG. Human genetic and immunological determinants of critical COVID-19 pneumonia. Nature. 2022;603:587–98.
Mantovani A, Dinarello CA, Molgora M, Garlanda C. Interleukin-1 and related cytokines in the regulation of inflammation and immunity. Immunity. 2019;50:778–95.
Declercq J, De Leeuw E, Lambrecht BN. Inflammasomes and IL-1 family cytokines in SARS-CoV-2 infection: from prognostic marker to therapeutic agent. Cytokine. 2022;157: 155934.
Bittner, Z.A., Schrader, M., George, S.E., Amann, R., (2022) pyroptosis and Its role in SARS-CoV-2 infection. Cells 11
Robinson, K.S., Teo, D.E.T., Tan, K.S., Toh, G.A., Ong, H.H., Lim, C.K., Lay, K., Au, B.V., Lew, T.S., Chu, J.J.H., Chow, V.T.K., Wang, Y., Zhong, F.L., Reversade, B., (2020). Enteroviral 3C protease activates the human NLRP1 inflammasome in airway epithelia. Science (New York, N.Y.) 370
Tsu, B.V., Beierschmitt, C., Ryan, A.P., Agarwal, R., Mitchell, P.S., Daugherty, M.D., 2021. Diverse viral proteases activate the NLRP1 inflammasome. eLife 10 e60609
Planès, R., Pinilla, M., Santoni, K., Hessel, A., Passemar, C., Lay, K., Paillette, P., Valadão, A.C., Robinson, K.S., Bastard, P., Lam, N., Fadrique, R., Rossi, I., Pericat, D., Bagayoko, S., Leon-Icaza, S.A., Rombouts, Y., Perouzel, E., Tiraby, M., Zhang, Q., Cicuta, P., Jouanguy, E., Neyrolles, O., Bryant, C.E., Floto, A.R., Goujon, C., Lei, F.Z., Martin-Blondel, G., Silva, S., Casanova, J.L., Cougoule, C., Reversade, B., Marcoux, J., Ravet, E., Meunier, E., 2022. Human NLRP1 is a sensor of pathogenic coronavirus 3CL proteases in lung epithelial cells. Molecular cell
Zhong FL, Robinson K, Teo DET, Tan KY, Lim C, Harapas CR, Yu CH, Xie WH, Sobota RM, Au VB, Hopkins R, D’Osualdo A, Reed JC, Connolly JE, Masters SL, Reversade B. Human DPP9 represses NLRP1 inflammasome and protects against autoinflammatory diseases via both peptidase activity and FIIND domain binding. J Biol Chem. 2018;293:18864–78.
Vasseur E, Boniotto M, Patin E, Laval G, Quach H, Manry J, Crouau-Roy B, Quintana-Murci L. The evolutionary landscape of cytosolic microbial sensors in humans. Am J Hum Genet. 2012;91:27–37.
Levandowski CB, Mailloux CM, Ferrara TM, Gowan K, Ben S, Jin Y, McFann KK, Holland PJ, Fain PR, Dinarello CA, Spritz RA. NLRP1 haplotypes associated with vitiligo and autoimmunity increase interleukin-1β processing via the NLRP1 inflammasome. Proc Natl Acad Sci USA. 2013;110:2952–6.
Wang, Q., Gao, H., Clark, K.M., Mugisha, C.S., Davis, K., Tang, J.P., Harlan, G.H., DeSelm, C.J., Presti, R.M., Kutluay, S.B., Shan, L., 2021. CARD8 is an inflammasome sensor for HIV-1 protease activity. Science (New York, N.Y.) 371
Taabazuing, C.Y., Griswold, A.R., Bachovchin, D.A., (2020) The NLRP1 and CARD8 inflammasomes. 297, 13–25
Hall SK, Perregaux DG, Gabel CA, Woodworth T, Durham LK, Huizinga TW, Breedveld FC, Seymour AB. Correlation of polymorphic variation in the promoter region of the interleukin-1 beta gene with secretion of interleukin-1 beta protein. Arthritis Rheum. 2004;50:1976–83.
Frayling TM, Rafiq S, Murray A, Hurst AJ, Weedon MN, Henley W, Bandinelli S, Corsi AM, Ferrucci L, Guralnik JM, Wallace RB, Melzer D. An interleukin-18 polymorphism is associated with reduced serum concentrations and better physical functioning in older people. J gerontol Series A, Bio sci med sci. 2007;62:73–8.
He M, Cornelis MC, Kraft P, van Dam RM, Sun Q, Laurie CC, Mirel DB, Chasman DI, Ridker PM, Hunter DJ, Hu FB, Qi L. Genome-wide association study identifies variants at the IL18-BCO2 locus associated with interleukin-18 levels. Arterioscler Thromb Vasc Biol. 2010;30(4):885–90.
Miller SA, Dykes DD, Polesky HF. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988;16:1215–1215.
Richardson S, Hirsch JS, Narasimhan M, Crawford JM, McGinn T, Davidson KW, Barnaby DP, Becker LB, Chelico JD, Cohen SL, Cookingham J, Coppa K, Diefenbach MA, Dominello AJ, Duer-Hefele J, Falzon L, Gitlin J, Hajizadeh N, Harvin TG, Hirschwerk DA, Kim EJ, Kozel ZM, Marrast LM, Mogavero JN, Osorio GA, Qiu M, Zanos TP. Presenting characteristics, comorbidities, and outcomes among 5700 patients hospitalized with COVID-19 in the New York city area. JAMA. 2020;323:2052–9.
Magitta NF, Bøe Wolff AS, Johansson S, Skinningsrud B, Lie BA, Myhr KM, Undlien DE, Joner G, Njølstad PR, Kvien TK, Førre Ø, Knappskog PM, Husebye ES. A coding polymorphism in NALP1 confers risk for autoimmune Addison’s disease and type 1 diabetes. Genes Immun. 2009;10:120–4.
Costa FRC, Leite JA, Rassi DM, da Silva JF, Elias-Oliveira J, Guimarães JB, Foss-Freitas MC, Câmara NOS, Pontillo A, Tostes RC, Silva JS, Carlos D. NLRP1 acts as a negative regulator of Th17 cell programming in mice and humans with autoimmune diabetes. Cell Rep. 2021;35: 109176.
Sun X, Xia Y, Liu Y, Wang Y, Luo S, Lin J, Huang G, Li X, Xie Z, Zhou Z. Polymorphisms in NLRP1 gene are associated with Type 1 diabetes. J Diabetes Res. 2019;2019:7405120.
Author information
Authors and Affiliations
Contributions
Conception and design: VNCL; AP, patients recruitment: LCN; JV; CBB; CAV; AMS; APHY; JMK; LMP; WMSS, acquisition of data: VNCL; MMSA; CAV; FMET; RAGC; SMY; RWA; SCGS; TMY; VFD; CBB; AMS; APHY; JMK; JV; LMP; WMSS. Analysis and interpretation of data: VNCL, AP. Writing of the manuscript: VNCL, AP. Critical reagents and manuscript editing: AP. Funding acquisition: AP, MNS, AJSD; JV.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Responsible Editor: John Di Battista.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Leal, V.N.C., Paulino, L.M., Cambui, R.A.G. et al. A common variant close to the “tripwire” linker region of NLRP1 contributes to severe COVID-19. Inflamm. Res. 72, 1933–1940 (2023). https://doi.org/10.1007/s00011-022-01670-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-022-01670-3