Summary
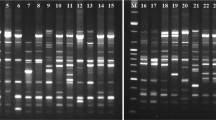
RAPD (Random Amplified Polymorphic DNA) markers generated by 4 arbitrary 10-mer primers, discriminated 14 broccoli and 12 cauliflower cultivars (Brassica oleracea L.) by banding profiles. The size of the amplified DNA fragments ranged from 300 to 2600 base pairs. Twenty-eight percent of the markers were fixed in both broccoli and cauliflower, whereas 12.5% were specific to either crop. The rest were polymorphic in either or both crops. The markers generated by two and three primers were sufficient to distinguish each of the broccoli and cauliflower cultivars, respectively. The average difference in markers was 14.5 between broccoli and cauliflower markers, 5.8 between two broccoli cultivars and 7.9 between two cauliflower cultivars. Larger differences for each crop were found between cultivars from different seed companies than within the same company. RAPD markers provide a quick and reliable alternative to identify broccoli and cauliflower cultivars.
Similar content being viewed by others
References
Arus, P., Tanksley, S. D., Orton, T. J., Jones, R. A. (1982) Electrophoretic variation as tool for determining seed purity and for breeding hybrid varieties of Brassica oleracea. Euphytica 31:417–428
Bassiri, A. (1976) Barley cultivar identification by use of isozyme electrophoretic patterns. Can J Plant Sci 56:1–6
Buehler, R.E., McDonald, M. B., TTV, J. Tori, Martin, S. K. S. (1989) Soybean cultivar identification using high performance liquid chromatography of seed proteins. Crop Sci 29:32–37
Burnouf, T., Bietz, J. A. (1987) Identification of wheat cultivars and prediction of quality by reversed-phase high-performance liquid chromatographic analysis of endosperm storage proteins. Seed Sci & Technol 15:79–99
Dellaporta, S. L., Wood, J., Hicks, J.B. (1983) A plant molecular DNA minipreparation version: II. Plant Mol Biol Rep 1:19–21
Gray, A. R. (1982) Taxonomy and evolution of broccoli (Brassica oleracea var italica). Econ Bot 36:397–410
Gupta, S. K., Robbelen, G. (1986) Identification of rapeseed (Brassica napus) cultivars by electrophoresis. Plant Breeding 96:363–370
Kianian, S.F. (1990) Evolutionary trends in the Brassica oleracea cytodeme: Chromosomal and linkage relationship. Ph.D. Thesis, University of California, Davis
Ladizinsky, G., Hymowitz, T. (1979) Seed protein electrophoresis in taxonomic and evolutionary studies. TAG 54:145–151
Nybom, H., Rogstad, S. H., Schaal, B. A. (1990) Genetic variation detected by use of the M13 DNA fingerprint probed in Malus, Prunus and Rubus (Rosaceae). Theo App Gene 79:153–156
Quiros, C. F., Hu, J., This, P., Chevre, A. M., Delseny, M. (1991) Development and chromosomal localization of genome specific markers by polymerase chain reaction in Brassica. TAG (In press)
Soller, M., Beckmann, J. S. (1983) Genetic polymorphism in varietal identification and genetic improvement. TAG 67:25–33
Stegemann, H. (1984) Retrospect on 25 years of cultivar identification by protein patterns and prospects for the future. Seed Sci & Technol 15:20–31
Vaughan, J. G., Denford, K. E. (1968) An acrylamine gel electrophoretic study of the seed protein of Brassica and Sinapis species, with special reference to their taxonomic nature. J Exp Bot 19:724–732
Welsh, J., McClelland, M. (1990) Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res 18:7213–7218
Williams, J. G. K., Kubelik, A. E., Levak, K. J., Rafalski, J. A., Tingey, S. C. (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Author information
Authors and Affiliations
Additional information
Communicated by R. Shoemaker
Rights and permissions
About this article
Cite this article
Hu, J., Quiros, C.F. Identification of broccoli and cauliflower cultivars with RAPD markers. Plant Cell Reports 10, 505–511 (1991). https://doi.org/10.1007/BF00234583
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF00234583