Abstract
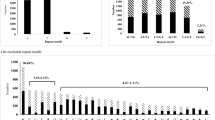
Single nucleotide polymorphisms (SNP) are the most abundant type of DNA polymorphism found in animal and plant genomes. They provide an important new source of molecular markers that are useful in genetic mapping, map-based positional cloning, quantitative trait locus mapping and the assessment of genetic distances between individuals. Very little is known on the frequency of SNPs in cassava. We have exploited the recently-developed collection of cassava expressed sequence tags (ESTs) to detect SNPs in the five cultivars of cassava used to generate the sequences. The frequency of intra-cultivar and inter-cultivar SNPs after analysis of 111 contigs was one polymorphism per 905 and one per 1,032 bp, respectively; totaling 1 each 509 bp. We have obtained further information on the frequency of SNPs in six cassava cultivars by analysis of 33 amplicons obtained from 3′ EST and BAC end sequences. Overall, about 11 kb of DNA sequence was obtained for each cultivar. A total of 186 SNPs (136 and 50 from ESTs and BAC ends, respectively) were identified. Among these, 146 were intra-cultivar polymorphisms, while 80 were inter-cultivar polymorphisms. Thus the total frequency of SNPs was one per 62 bp. This information will help to develop new strategies for EST mapping as well as their association with phenotypic characteristics.
Similar content being viewed by others
References
Batley J, Barker G, O’Sullivan H, Edwards KJ, Edwards D (2003) Mining for single nucleotide polymorphisms and insertions/deletions in maize expressed sequence tag data. Plant Physiol 132:84–91
Brumfield RT, Beerli P, Nickerson DA, Edwards SV (2003) The utility of single nucleotide polymorphisms in inferences of population history. Trends Ecol Evol 18:249–256
Ching A, Rafalski A (2002) Rapid genetic mapping of ESTs using SNP pyrosequencing and indel analysis. Cell Mol Biol Lett 7:803–810
Ching A, Caldwell KS, Jung M, Dolan M, Smith OS, Tingey S, Morgante M, Rafalski AJ (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19
Cho RJ, Mindrinos M, Richards DR, Sapolsky RJ, Anderson M, Drenkard E, Dewdney J, Reuber TL, Stammers M, Federspiel N, Theologis A, Yang WH, Hubbell E, Au M, Chung EY, Lashkari D, Lemieux B, Dean C, Lipshutz RJ, Ausubel FM, Davis RW, Oefner PJ (1999) Genome-wide mapping with biallelic markers in Arabidopsis thaliana. Nat Genet 23:203–207
CIAT (2002) Annual Report, Centro Internacional de Agricultura Tropical, Cali, Colombia
Cooper DN, Smith BA, Cooke HJ, Niemann S, Schmidtke J (1985) An estimate of unique DNA sequence heterozygosity in the human genome. Hum Genet 69:201–205
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: version II. Plant Mol Biol Rep 1:19–21
Ellis JG, Lawrence GJ, Luck JE, Dodds PN (1999) Identification of regions in alleles of the flax rust resistance gene L that determine differences in gene-for-gene specificity. Plant Cell 11:495–506
Ewing B, Hillier L, Wendl MC, Green P (1998) Base-calling of automated sequencer traces using phred I accuracy assessment. Genome Res 8:175–185
Fregene M, Angel F, Gomez R, Rodriguez F, Chavarriaga P, Roca W, Tohme J, Bonierbale M (1997) A molecular genetic map of cassava (Manihot esculenta Crantz). Theor Appl Genet 95:431–441
Garg K, Green P, Nickerson DA (1999) Identification of candidate coding region single nucleotide polymorphisms in 165 human genes using assembled expressed sequence tags. Genome Res 9:1087–1092
Gotoh K, Oishi M (2003) Screening of gene-associated polymorphisms by use of in-gel competitive reassociation and EST (cDNA) array hybridization. Genome Res 13:492–495
Grivet L, Glaszmann JC, Vincentz M, da Silva F, Arruda P (2003) ESTs as a source for sequence polymorphism discovery in sugarcane: example of the Adh genes. Theor Appl Genet 106:190–197
Jander G, Norris SR, Rounsley SD, Bush DF, Levin IM, Last RL (2002) Arabidopsis map-based cloning in the post-genome era. Plant Physiol 129:440–450
Jorge V, Fregene MA, Duque MC, Bonierbale MW, Tohme J, Verdier V (2000) Genetic mapping of resistance to bacterial blight disease in cassava (Manihot esculenta Crantz). Theor Appl Genet 101:865–872
Jorge V, Fregene M, Velez CM, Duque MC, Tohme J, Verdier V (2001) QTL analysis of field resistance to Xanthomonas axonopodis pv . manihotis in cassava. Theor Appl Genet 102:564–571
Kota R, Rudd S, Facius A, Kolesov G, Thiel T, Zhang H, Stein N, Mayer K, Graner A (2003) Snipping polymorphisms from large EST collections in barley (Hordeum vulgare L.). Mol Genet Genomics 270:24–33
Lopez CE, Zuluaga AP, Cooke R, Delseny M, Tohme J, Verdier V (2003) Isolation of resistance gene candidates (RGCs) and characterization of an RGC cluster in cassava. Mol Genet Genomics 269:658–671
Lopez CE, Jorge V, Piegu B, Mba Ch, Cortes D, Restrepo S, Soto M, Laudié M, Berger Ch, Cooke R, Delseny M, Tohme J, Verdier V (2004) A unigene catalogue of 5,700 expressed genes in cassava. Plant Mol Biol (in press)
Marth GT, Korf I, Yandell MD, Yeh RT, Gu Z, Zakeri H, Stitziel NO, Hillier L, Kwok PY, Gish WR (1999) A general approach to single-nucleotide polymorphism discovery. Nat Genet 23:452–456
Mba REC, Stephenson P, Edwards K, Melzer S, Nkumbira J, Gullberg U, Apel K, Gale M, Tohme J, Fregene M (2001) Simple sequence repeat (SSR) markers survey of the cassava (Manihot esculenta Crantz) genome: towards an SSR-based molecular genetic map of cassava. Theor Appl Genet 102:21–31
Miller RT, Christoffels AG, Gopalakrishnan C, Burke J, Ptitsyn AA, Broveak TR, Hide WA (1999) A comprehensive approach to clustering of expressed human gene sequence: the sequence tag alignment and consensus knowledge base. Genome Res 9:1143–1155
Mohan M, Nair S, Bhagwat A, Krishna TG, Yano M, Bhatia CR, Sasaki T (1997) Genome mapping, molecular markers and marker-assisted selection in crop plants. Mol Breed 3:87–103
Okogbenin E, Fregene M (2003) Genetic mapping of QTLs affecting productivity and plant architecture in a full-sib cross from non-inbred parents in Cassava (Manihot esculenta Crantz). Theor Appl Genet 107:1452–1462
Olsen KM, Hernandez M, Schaal BA (1998) A survey of DNA sequence variation in cassava and other Manihot species. In: Cassava biotechnology: proceedings of the IVth international scientific meeting of the cassava biotechnology network. EMBRAPA-Recursos Geneticos e Biotecnologia, Brasilia
Pacey-Miller T, Henry R (2003) Single-nucleotide polymorphism detection in plants using a single-stranded pyrosequencing protocol with a universal biotinylated primer. Anal Biochem 317:166–170
Phillips RL, Vasil IK (2001) DNA-based markers in plants. Kluwer, Dordrecht
Picoult-Newberg L, Ideker TE, Pohl MG, Taylor SL, Donaldson MA, Nickerson DA, Boyce-Jacino M (1999) Mining SNPs from EST databases. Genome Res 9:167–174
Rafalski A (2002) Applications of single nucleotide polymorphisms in crop genetics. Curr Opin Plant Biol 5:94–100
Roger DJ, Appan SG (1973) Manihot and Manihotoides (Euphorbiaceae): a computer assisted study. Monograph no. 13, flora neotropica. Hafner, New York
Schmid KJ, Sorensen TR, Stracke R, Torjek O, Altmann T, Mitchell-Olds T, Weisshaar B (2003) Large-scale identification and analysis of genome-wide single-nucleotide polymorphisms for mapping in Arabidopsis thaliana. Genome Res 13:1250–1257
Schwarz G, Baumler S, Block A, Felsenstein FG, Wenzel G (2004) Determination of detection and quantification limits for SNP allele frequency estimation in DNA pools using real time PCR. Nucleic Acids Res 32:e24
Useche FJ, Gao G, Harafey M, Rafalski A (2001) High-throughput identification, database storage and analysis of SNPs in EST sequences. Genome Inform Ser Workshop Genome Inform 12:194–203
Wang DG, Fan JB, Siao CJ, Berno A, Young P, Sapolsky R, Ghandour G, Perkins N, Winchester E, Spencer J, Kruglyak L, Stein L, Hsie L, Topaloglou T, Hubbell E, Robinson E, Mittmann M, Morris MS, Shen N, Kilburn D, Rioux J, Nusbaum C, Rozen S, Hudson TJ, Lander ES et al (1998) Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome. Science 280:1077–1082
Zhu YL, Song QJ, Hyten DL, Van Tassell CP, Matukumalli LK, Grimm DR, Hyatt SM, Fickus EW, Young ND, Cregan PB (2003) Single-nucleotide polymorphisms in soybean. Genetics 163:1123–1134
Acknowledgements
We are grateful to Chike Mba and Olivier Panaud for critically reading the manuscript. Thanks to Christel Llauro-Berger and Michèle Laudié for sequencing. All the sequencing was performed by the facilities of the Montpellier Languedoc Rousillon Genopole. Camilo Lopez was supported by a doctoral fellowship awarded by the IRD.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E. Guiderdoni
Rights and permissions
About this article
Cite this article
Lopez, C., Piégu, B., Cooke, R. et al. Using cDNA and genomic sequences as tools to develop SNP strategies in cassava (Manihot esculenta Crantz). Theor Appl Genet 110, 425–431 (2005). https://doi.org/10.1007/s00122-004-1833-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-004-1833-3